Clear Sky Science · en
Human brain and organoid transcriptomes reveal key receptor tyrosine kinase pathways and genetic signatures in Alzheimer's disease
Why this matters for understanding Alzheimer’s
Alzheimer’s disease is usually described in terms of sticky plaques and tangled proteins in the brain. But behind those visible changes lies a quieter story written in genes and cell-to-cell signals. This study looks under the hood of the human brain and lab-grown “mini-brains” to find which signaling switches on brain cells are most disturbed in Alzheimer’s, and whether they could help us detect or even treat the disease earlier.
Looking for hidden signals in a vulnerable brain region
The researchers focused on the middle temporal gyrus, a brain region important for language and memory that is hit early in Alzheimer’s. They analyzed gene activity from hundreds of thousands of genes in brain tissue donated by people who had Alzheimer’s and people who did not. By comparing two large public data sets from the United States and Europe, they homed in on 145 genes that were consistently altered in Alzheimer’s brains. Many of these genes clustered around a family of surface molecules called receptors, which act like antennae that let brain cells sense and respond to their environment.
Receptor “antennae” and a short list of suspect genes
Digging deeper, the team used several online biology databases to ask which pathways these 145 genes belonged to. A strong signal emerged from receptor tyrosine kinases, a group of receptors that help control cell survival, growth, and communication. From this group, they narrowed the list to 18 genes tied to these receptor pathways, and then built an interaction map that also included well-known Alzheimer’s genes linked to amyloid and tau. This network analysis highlighted six genes that sat at key crossroads: AXL and ITGB1 (both involved in how cells sense their surroundings), GFAP (an astrocyte marker), CAV1 and RHOA (which help shape cell structure and signaling), and NRG1 (important for brain development). In Alzheimer’s brains, five of these genes were turned up, while NRG1 was turned down.
Testing the findings in mini-brains and neurons
To move beyond computer analyses, the researchers turned to human stem cell–derived brain organoids—three-dimensional “mini-brains” that contain neurons and support cells—and to primary rat neurons grown in dishes. When they exposed these organoids to amyloid-beta, the toxic protein linked to plaques, levels of several of the key genes, including AXL and ITGB1, rose. Protein tests showed that AXL increased inside the organoids, and ITGB1 increased specifically at the cell membrane, where it can influence how cells stick and signal to one another. At the same time, a major internal signaling route called the PI3K–AKT pathway became more active, suggesting that amyloid may be driving a specific branch of receptor signaling rather than just causing general stress.
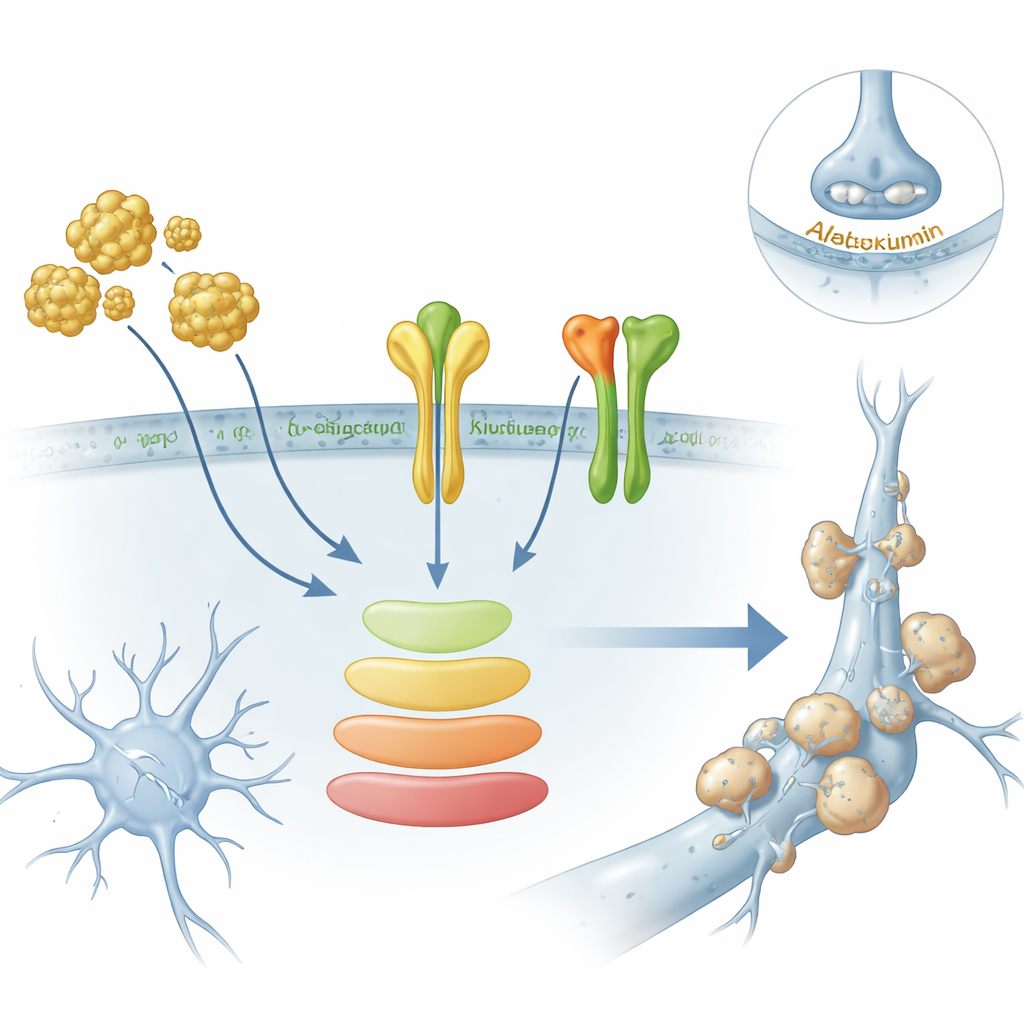
From gene patterns to possible biomarkers
The team then asked whether the combined activity of the six highlighted genes could help tell apart Alzheimer’s brains from cognitively normal ones. Using statistical models, they found that a score built from these six genes separated the two groups with high accuracy in both brain cohorts. Higher scores also tracked with more severe clinical dementia ratings. This suggests that patterns of receptor-related gene activity, especially involving AXL and ITGB1, may serve as molecular fingerprints of Alzheimer’s-related changes in the middle temporal gyrus and perhaps other vulnerable regions such as the hippocampus.
What this could mean for future diagnosis and treatment
Overall, the study proposes that certain receptor pathways, centered on the AXL receptor and the integrin ITGB1, become overactive in Alzheimer’s-like conditions and feed into a signaling loop that may worsen disease. While much work remains—especially experiments that directly switch these genes on or off in living systems—the findings point to a new layer of Alzheimer’s biology beyond plaques and tangles. In the long run, measuring these gene and protein changes, or safely dialing down overactive receptor pathways, could complement existing approaches to diagnosing and treating Alzheimer’s disease.
Citation: Shin, S., Zhu, X., Amartumur, S. et al. Human brain and organoid transcriptomes reveal key receptor tyrosine kinase pathways and genetic signatures in Alzheimer's disease. Exp Mol Med 58, 1230–1241 (2026). https://doi.org/10.1038/s12276-026-01684-5
Keywords: Alzheimer’s disease, brain signaling, receptor pathways, brain organoids, biomarkers