Clear Sky Science · en
Stereological reconstructions of 3D cellular microstructures by combining adversarial learning and Voronoi tessellations
Why the hidden inner structure of materials matters
Everyday materials, from metal bike frames to plant leaves and foam cushions, are built from tiny three dimensional “cells” packed together. The shapes, sizes, and connections of these cells quietly control how a material bends, breaks, insulates heat, or conducts electricity. Yet seeing these 3D cell networks usually demands expensive or destructive imaging. This paper introduces a new way to rebuild realistic 3D cellular structures from much simpler two dimensional images, helping scientists explore and design materials without cutting them apart.
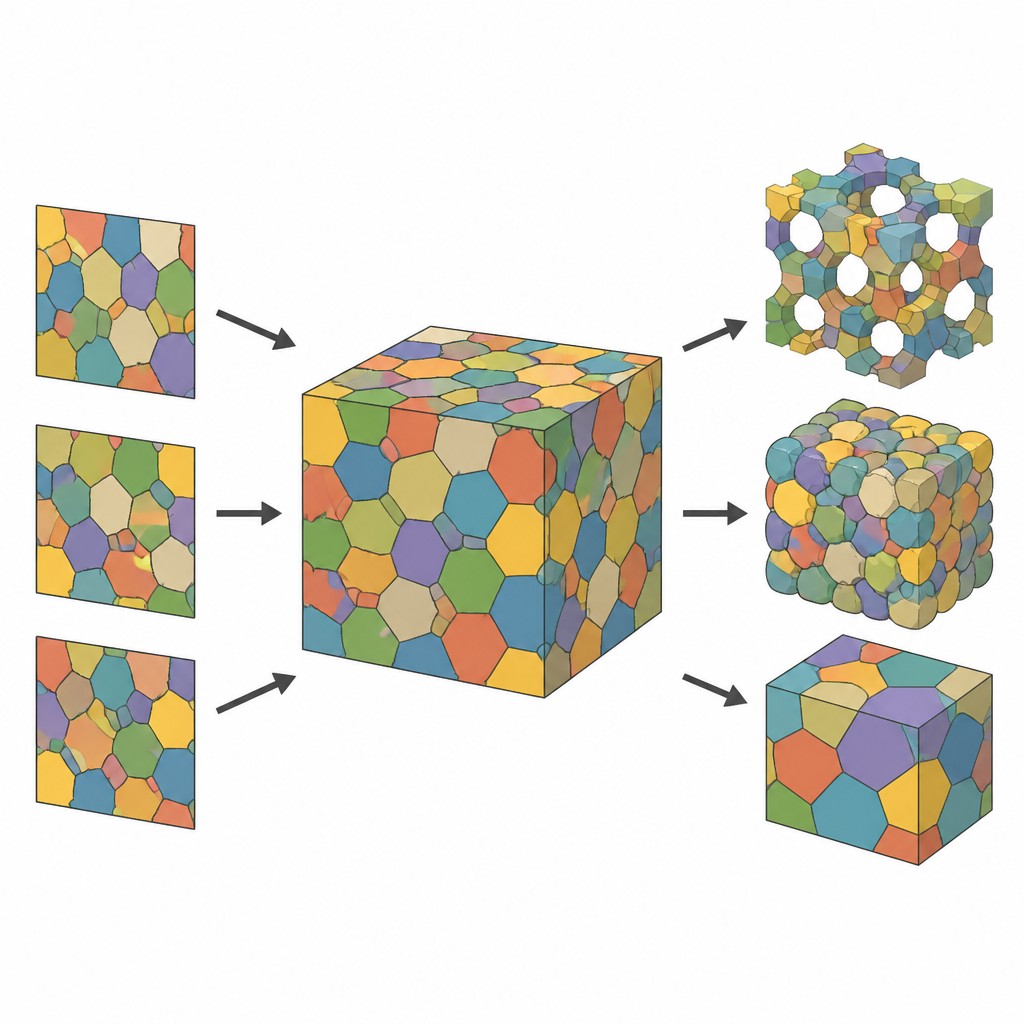
From flat slices to full 3D cell maps
Most experiments capture only thin slices of a material: microscope images that show how cells look in a flat cross section. While useful, such 2D views miss how cells extend and connect in depth. The authors tackle this by treating the unseen 3D structure as a collection of cells that partition space, much like a mosaic of touching polyhedral “bubbles.” By starting from a cloud of points in space and assigning each point the region closest to it, they obtain a Voronoi tessellation, a geometric framework that naturally mimics many cellular materials. Each cell is then colored and represented as a smooth, computer friendly image so the model can be adjusted and compared to measured slices.
Teaching a neural network to judge realism
To make the synthetic 3D structure resemble real materials, the authors train a discriminator network, similar to those used in image generating artificial intelligence. This network learns to tell apart 2D images cut from the current 3D model and real 2D slices measured from experiments. At first, the model’s slices look unrealistic, so the discriminator easily spots them as fake. The algorithm then slightly shifts the seed points that define the cells, using gradient based optimization, to make the model’s slices harder to distinguish from real ones. Over many rounds, the 3D cellular pattern evolves until the discriminator rates simulated and measured slices as statistically similar.
Balancing detail, efficiency, and interpretability
The approach is designed to use very few parameters per cell while keeping a clear physical meaning. Each cell is defined mainly by the position of a single point, rather than thousands of image pixels. This compact description cuts memory use by roughly a factor of hundreds compared with full 3D images, yet still allows direct control over important features like cell size, shape, and neighborhood relations. Periodic boundary conditions avoid artificial edge effects in small samples, and a “soft” version of the tessellation, based on smooth distance based weights, keeps the optimization stable and differentiable while preserving the ability to recover crisp cell boundaries afterward.
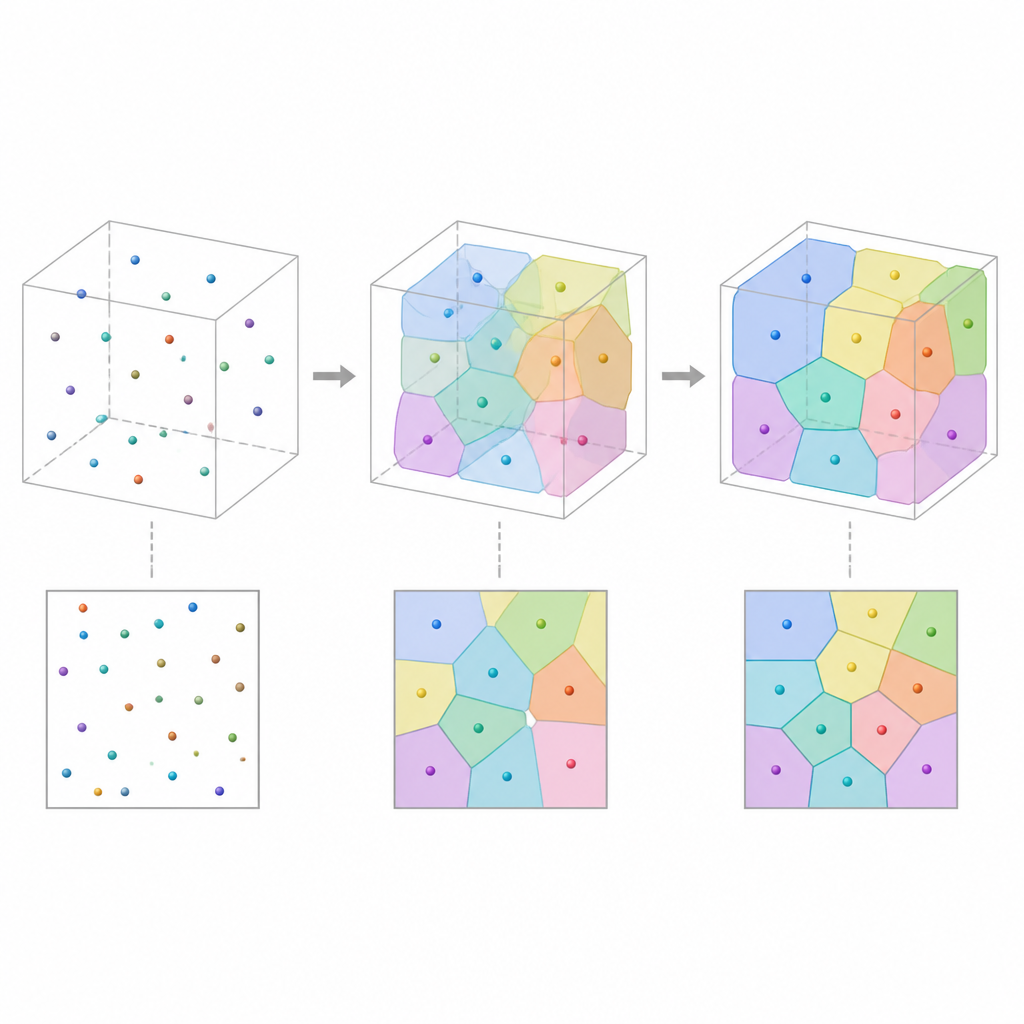
Testing on synthetic and real materials
The team first checks their framework on artificial microstructures that are already known to be well described by Voronoi cells. By reconstructing 3D patterns from many 2D slices and comparing statistics such as cell volume, surface area, elongation, and number of neighbors, they find that the generated structures closely match the originals. They then apply the method to real measured data: polymer foams, nearly spherical biological cells, and polycrystalline metals with more elongated and twinned grains. In all three cases, the reconstructed 3D tessellations reproduce key geometric trends seen in experiments, and often do so more accurately in terms of cell sizes and surfaces than a purely voxel based competing method.
What this means for future material design
The study shows that it is possible to infer rich 3D cellular architectures from 2D images using a blend of geometric tessellations and adversarial learning. While the present version is limited to cells with straight, flat faces and isotropic statistics, it already offers a fast, low parameter, and physically interpretable route to build virtual 3D samples that can feed directly into simulation tools. For non experts, the key message is that scientists can now “reinflate” flat material slices into realistic 3D models, reducing the reliance on costly 3D imaging while still capturing the essential internal structure that governs a material’s behavior.
Citation: Fuchs, L., Wilhelm, T., Furat, O. et al. Stereological reconstructions of 3D cellular microstructures by combining adversarial learning and Voronoi tessellations. Sci Rep 16, 15058 (2026). https://doi.org/10.1038/s41598-026-52851-7
Keywords: 3D microstructure, cellular materials, Voronoi model, adversarial learning, stereology