Clear Sky Science · en
Method comparison of microscopy, metabarcoding, and multispectral imaging flow cytometry for identification and relative abundance analysis of insect-dispersed pollen
Why counting pollen matters
Pollen may look like simple yellow dust, but for wildflowers, crops, and the insects that visit them, it is a lifeline. Understanding which plants insects visit, and how much pollen they carry, helps scientists track the health of ecosystems, food production, and pollinator nutrition. Yet actually identifying and counting these microscopic grains is painstaking work. This paper asks a practical question with big implications: which modern tools do the best job of telling us what pollen is present and in what amounts on pollinating insects?
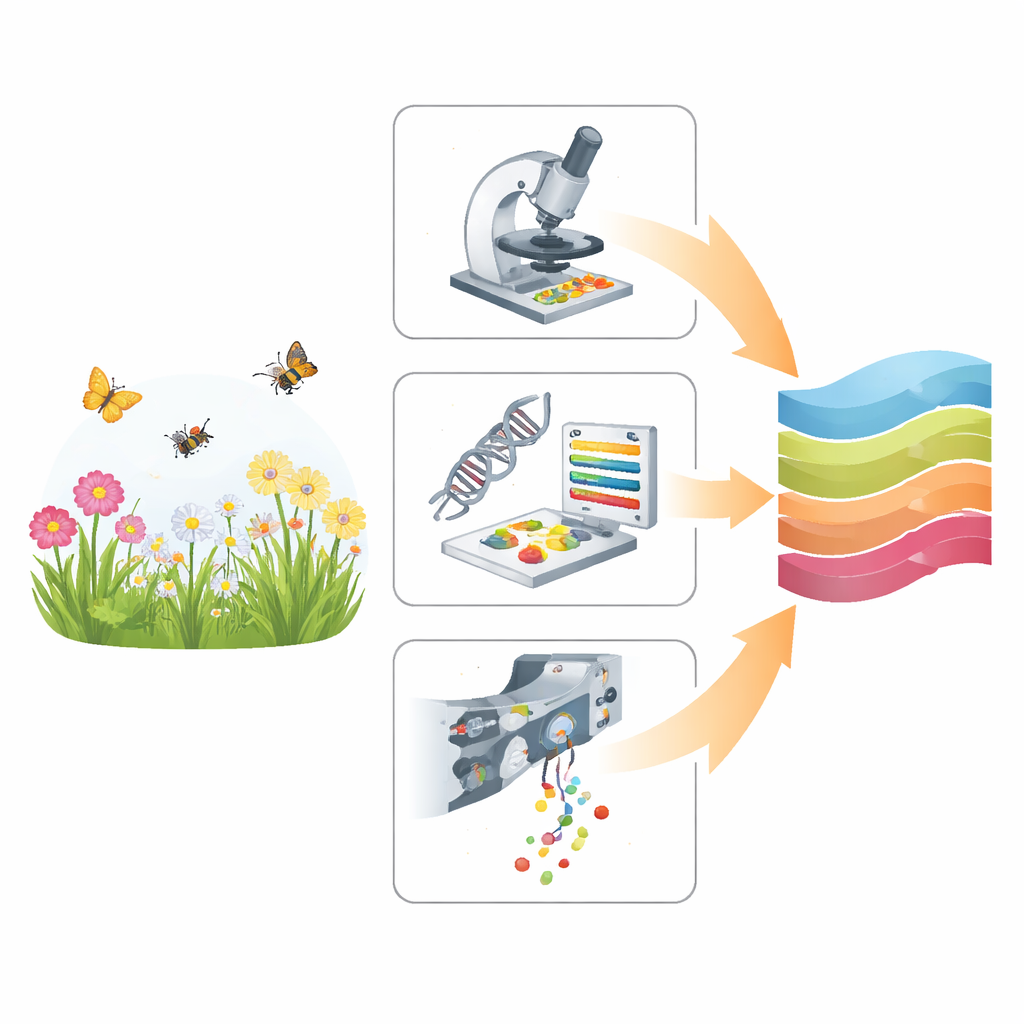
Three different ways to read pollen
The study compares three approaches that all start from the same basic material—mixed pollen collected from flowers and from insect bodies—but extract information in very different ways. Traditional light microscopy relies on a trained expert who looks at magnified grains and recognizes their shapes and surface patterns. Metabarcoding skips the shapes and reads short pieces of DNA from the pollen, matching them to large genetic reference libraries. A newer technique, multispectral imaging flow cytometry (MIFC), streams thousands of grains past cameras and sensors, capturing images and light signals that a computer model uses to sort them into species. Together, these methods span the spectrum from slow, hands-on observation to highly automated, high‑throughput analysis.
Putting the methods to a fair test
To compare performance, the researchers first built “artificial” pollen mixtures in the lab from nine common wildflower species in Romanian hay meadows. For these samples, the exact species and their true proportions were known, allowing a direct test of accuracy. They created three mixture types that differed mainly in how common small‑grained pollen was, then split identical portions of each mixture for all three methods. In a second step, they analyzed pollen that had naturally fallen off wild bees, bumblebees, and flies caught in the field, where the real composition was unknown but closely mirrored real‑world ecological studies.
Who is best at naming species?
When the challenge was simply to detect which plant kinds were present in the artificial mixtures—without any hints—DNA metabarcoding was the clear winner. It correctly picked up a higher share of the target genera than either microscopy or MIFC and produced fewer spurious detections. Because it relies on differences in DNA rather than in shape, metabarcoding can separate look‑alike grains and even distinguish closely related species, something that often defeats the human eye and automated image classifiers. However, all methods occasionally missed certain taxa, and the morphology‑based tools were especially sensitive to pollen that had shrunk or changed during storage, or that differed in subtle ways from the reference images used to train the computer model.
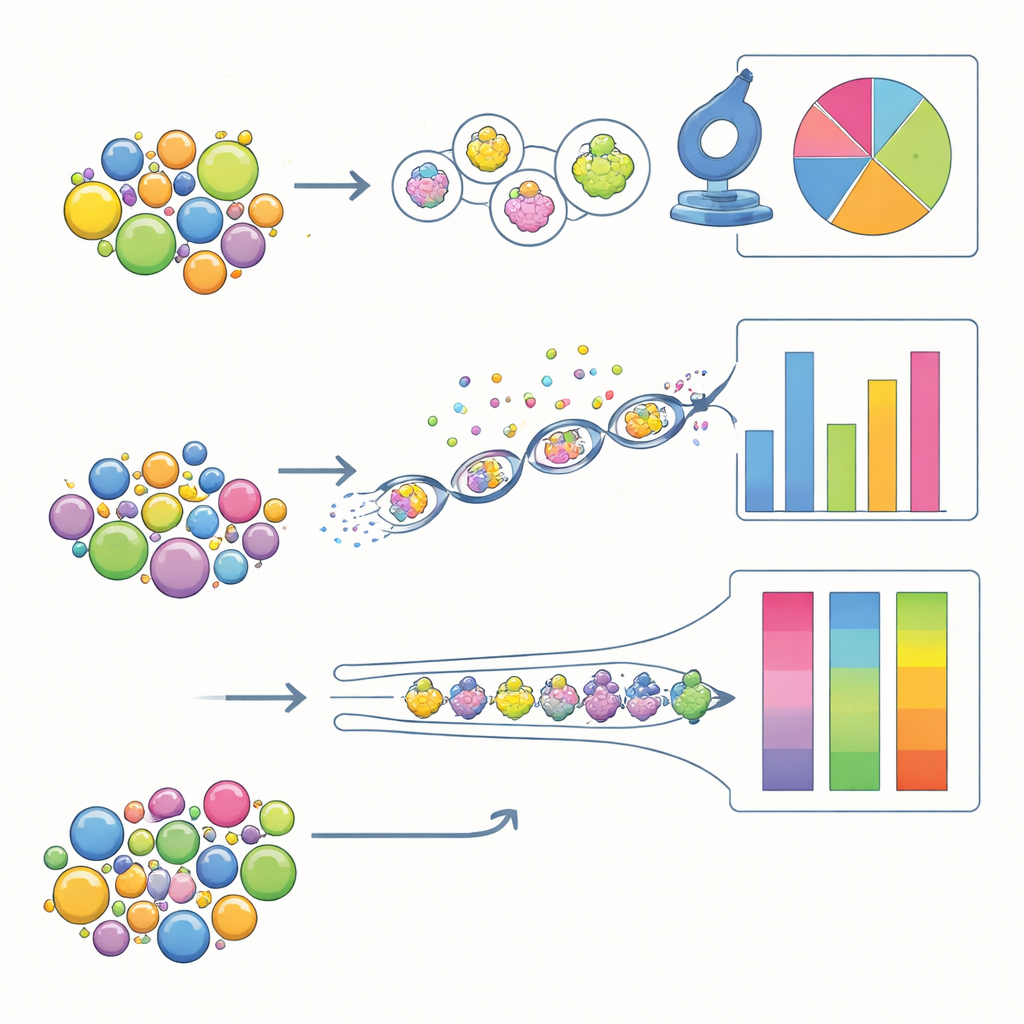
Who is best at counting fairly?
Accuracy looked different when the task was to estimate how much of each pollen type was present. After correcting for misidentifications and assuming that the set of species in the sample was known, traditional microscopy came closest to the true proportions in the artificial mixtures, followed by MIFC. Metabarcoding did the worst on this front: some species were consistently over‑represented in DNA read counts, while others were under‑represented. These biases likely arise from unequal amounts of DNA per grain, differences in how easily DNA is extracted, and quirks of the amplification process. MIFC processed far more grains than microscopy and showed generally good precision, but its accuracy still depended heavily on how well the image library reflected real‑world variation in pollen.
Mixed signals from real insects
When the team turned to pollen from wild insects, agreement among methods dropped markedly. For some insects carrying mostly one pollen type, all three approaches converged on similar pictures. For more diverse samples, however, the methods often disagreed, even when the morphology‑based tools were “guided” by the species list from metabarcoding. Interestingly, the two image‑based methods—light microscopy and MIFC—matched each other least well, while both aligned somewhat better with the DNA results. These discrepancies highlight how sample handling, clumping of grains, storage effects, and blind spots in each reference library can all shape the final picture of what an insect has been visiting.
A practical recipe for future pollen work
The authors conclude that no single technique can yet deliver both perfect identification and perfect counting. For studies that mainly need to know which plants are being visited, DNA metabarcoding is recommended as the most reliable option. When the priority is to quantify how much pollen is carried, traditional microscopy or MIFC perform better, with MIFC offering a major time advantage for large sample sets. To obtain both identity and relative abundance—the situation in many ecological and conservation projects—the study advocates a two‑step strategy: first use metabarcoding to build a trustworthy plant list for each sample, then use that information to guide high‑throughput image‑based counting, especially with MIFC. This combined approach, the authors argue, is well suited to tracking pollination and pollinator diets across the broad spatial and temporal scales demanded by today’s biodiversity and climate challenges.
Citation: Motivans Švara, E., Rakosy, D., Knight, T.M. et al. Method comparison of microscopy, metabarcoding, and multispectral imaging flow cytometry for identification and relative abundance analysis of insect-dispersed pollen. Sci Rep 16, 12578 (2026). https://doi.org/10.1038/s41598-026-47800-3
Keywords: pollen analysis, pollinators, DNA metabarcoding, microscopy, imaging flow cytometry