Clear Sky Science · en
Population genetic variation characterised through serial independent pool-seq: the Cyprus Genome Project
Why the DNA of a small island matters
Imagine trying to diagnose and treat disease using maps that mostly describe other people, from other places. That is the situation many countries face in modern genetics. This study tackles that gap for Cyprus by building the first large-scale genetic reference map made specifically from people living on the island. By cleverly pooling DNA from 10,000 volunteers, the researchers created a cost‑effective, privacy‑preserving way to chart which genetic changes are common or rare in Cypriots—and how those patterns differ from global averages that doctors usually rely on.
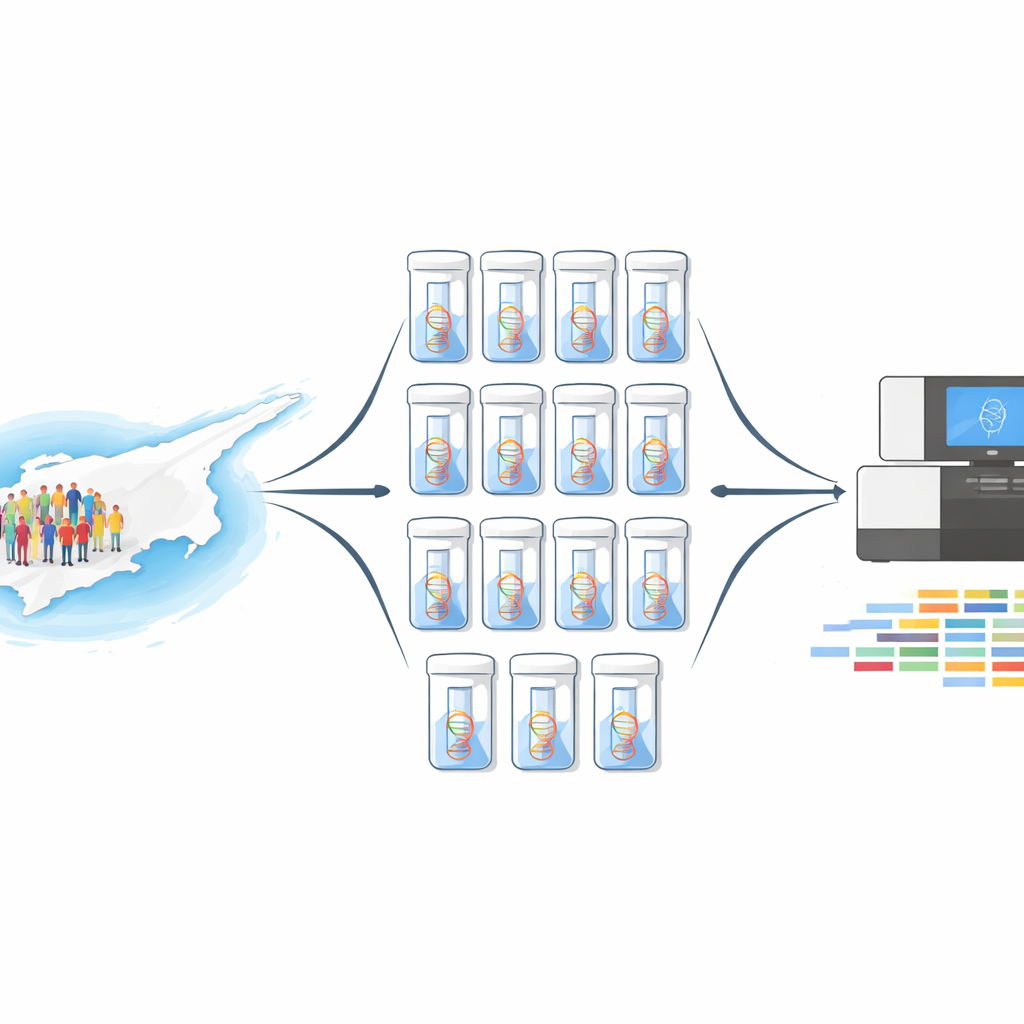
A new way to look at thousands of people at once
Sequencing the full genetic code of 10,000 individuals one by one would be hugely expensive and time‑consuming. Instead, the team used a method called DNA pooling. They combined equal amounts of DNA from 1,000 volunteer bone marrow donors into a single mixture, and repeated this ten times, creating ten independent pools. Each pool was then sequenced twice: once using whole‑exome sequencing, which scans all protein‑coding regions of the genome, and once using a focused panel of 813 medically important genes. This design delivered both breadth across the genome and very deep coverage in genes that matter most for disease.
Separating real signals from noise
Pooling DNA has trade‑offs. While it hides the identity of any single person and cuts costs, it also makes it harder to spot very rare changes and to be sure that a faint signal is not just a machine error. To address this, the authors built strong safeguards into their study. They required that any reported genetic variant be supported by at least 20 sequencing reads and show up in at least three of the ten independent pools. For a high‑confidence subset, they demanded thousands of total reads across the pools. They also checked that the pattern of variant frequencies in each pool matched the others and compared their results against global databases and smaller Cypriot studies. Strong statistical agreement showed that the pooled approach could measure how frequent each variant is with high precision.
What makes Cypriot DNA stand out
From the pooled data, the project catalogued more than four million genetic variants, including over 100,000 that do not appear in major international resources such as gnomAD and ClinVar. Many of these differences matter clinically. Some gene changes that are linked to blood disorders or bleeding problems, for example, are several dozen times more common in Cyprus than in global datasets, suggesting they may deserve a place in national screening programs. Conversely, the team also found variants that look extremely rare worldwide but turn out to be relatively frequent in healthy Cypriot donors. Without local data, such changes could be wrongly suspected of causing disease simply because they are so uncommon elsewhere.
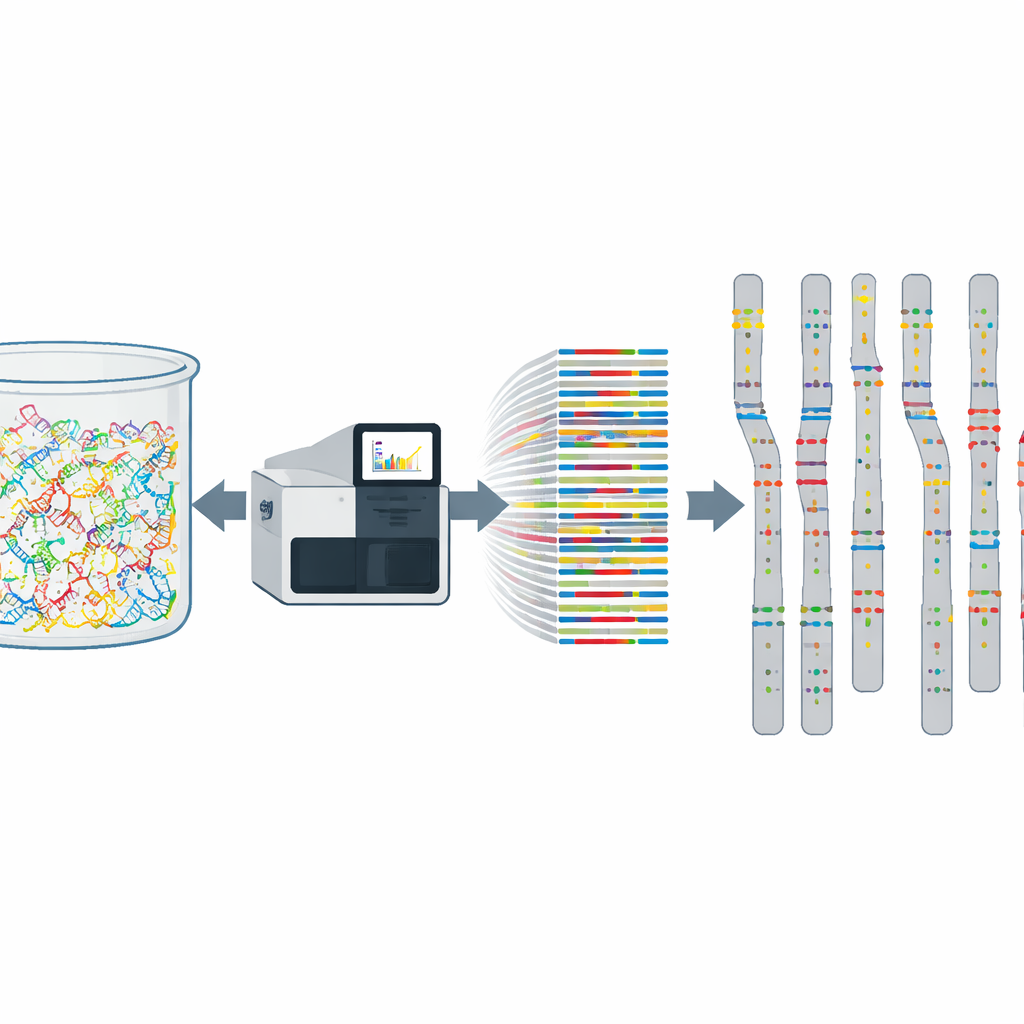
Rethinking risk scores and genetic tests
The new map also shows that some risk‑raising variants that are common globally are much rarer in Cyprus, and vice versa. That mismatch has practical consequences. Many modern tools, such as polygenic risk scores that estimate a person’s chance of developing conditions like cancer or heart disease, assume that the underlying variant frequencies in the target population resemble those in the reference population used to build the tool. The Cyprus data reveal that this assumption often fails. As a result, risk scores and diagnostic panels built on foreign data may over‑ or underestimate risk for Cypriot patients unless they are recalibrated using this new local reference.
How this resource can shape future healthcare
For non‑specialists, the bottom line is that this project turns the genetic background of Cyprus from a blind spot into a detailed map. The database at cyprusgenome.org lets clinicians and researchers see whether a genetic variant is truly rare and suspicious locally, or simply a harmless feature of Cypriot ancestry that global studies have missed. That clarity can prevent misdiagnoses, guide smarter national screening strategies, and lay the groundwork for more accurate risk prediction tools tailored to the island’s population. As additional sequencing and future layers of data, such as immune gene profiles and gene activity measurements, are added, this resource is poised to become a cornerstone for precision medicine and public health planning in Cyprus.
Citation: Antoniades, A., Chi, J., Brown, C. et al. Population genetic variation characterised through serial independent pool-seq: the Cyprus Genome Project. Sci Rep 16, 14215 (2026). https://doi.org/10.1038/s41598-026-44707-x
Keywords: Cyprus genome, population genetics, pooled DNA sequencing, rare variants, precision medicine