Clear Sky Science · en
Validation and application of a standardized quantitative PCR assay for the assessment of antimicrobial resistance genes in surface water
Why tiny genes in wild waters matter
When you hike through a national park and see clear, rushing streams, they look untouched by people. But even in remote places, microscopic traces of our antibiotics can show up as genes that make bacteria harder to kill. This study asked a simple but important question: can we reliably measure these antibiotic resistance genes in surface waters, and use that information to understand how much human activity is affecting even our most protected landscapes?
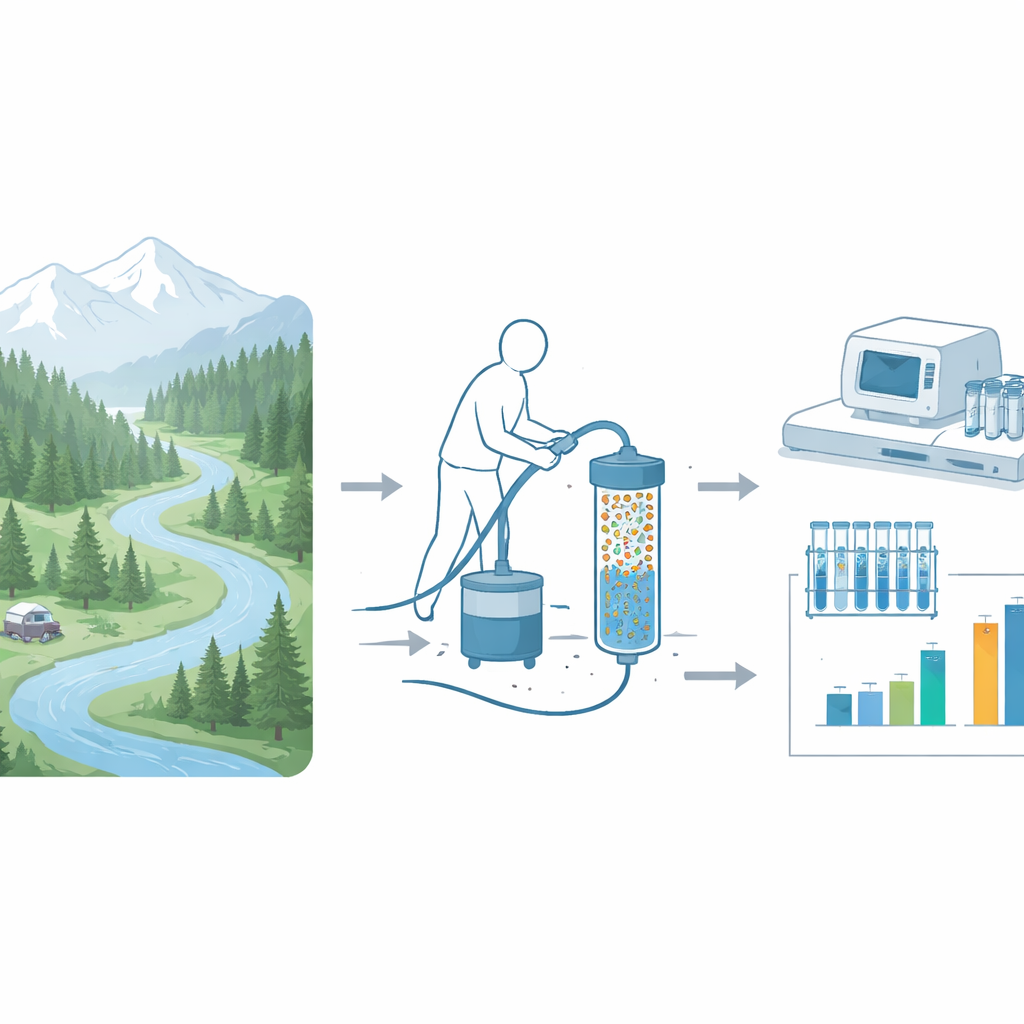
Looking for warning signs in flowing water
Antibiotics save lives, but their heavy use in medicine and agriculture has helped bacteria evolve defenses. Those defense instructions are written in DNA as antibiotic resistance genes. When people or animals shed bacteria, or when wastewater escapes into rivers, these genes can spread through the environment, sometimes ending up in places that seem pristine. Scientists and public land managers need ways to monitor these genetic warning signs in lakes and streams so they can spot changes early and judge whether visitor use, historical pollution, or natural background processes are at work.
Building a standardized genetic test
The research team developed a single genetic test plate based on quantitative PCR, a method that can count specific DNA sequences. Their panel targeted 47 different bacterial gene markers, including many known resistance genes and a few that help trace where contamination comes from, such as humans, birds, or grazing animals. They carefully designed or adapted DNA primers—short sequences that tell the PCR machine what to copy—and then checked that each one truly matched the intended gene by comparing to major reference databases. To see how trustworthy the test was, they ran it on DNA from 41 clinical bacterial isolates and compared the results to full genome sequencing. Agreement between the quick test and the much more detailed sequencing was extremely high, with both sensitivity and specificity near 98 percent.
Concentrating rare signals from large volumes
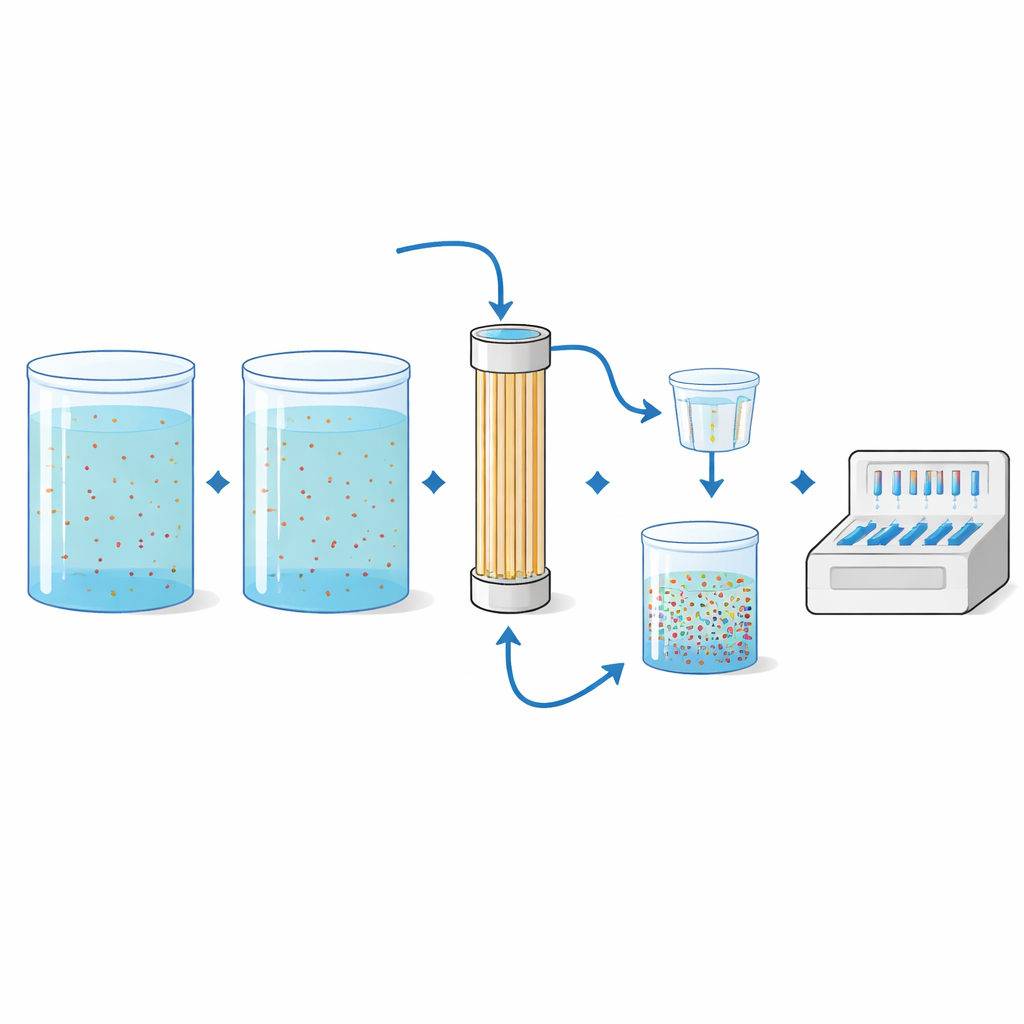
Resistance genes in wild streams can be vanishingly rare, so the team needed a way to “zoom in” on tiny traces in a lot of water. They used hollow-fiber ultrafiltration, a technology borrowed from dialysis and drinking water treatment. Large volumes—up to 50 liters in this study—were pumped through a cartridge filled with fine hollow fibers that trap microbes. The trapped material was then rinsed off the filter, further concentrated on smaller membranes, and finally analyzed with the PCR panel. In controlled lab trials, the scientists spiked sterile water with known bacteria carrying eight different resistance genes at several concentrations and volumes. The method reliably detected many of the spiked genes, especially when larger water volumes were filtered, and the measured ratios of resistance genes to total bacterial DNA closely matched expectations.
What Alaskan parks can reveal
Armed with this approach, the team turned to three national parks in Alaska that differ in how many people visit them: Denali, Kenai Fjords, and Wrangell–St. Elias. At each park they sampled two creeks, one near campgrounds, roads, or old mining sites and one in a more remote location. From these streams, they filtered multiple 20- and 50-liter samples, cultured some bacteria to test which could grow in the presence of antibiotics, and used the PCR panel plus DNA sequencing of general bacterial markers to see how resistance genes and broader microbial communities were distributed. Across all six sites they detected 19 distinct resistance genes with high confidence. Some, like genes linked to common antibiotics such as sulfonamides and tetracyclines, were fairly widespread, while others associated with powerful hospital drugs appeared in only a few samples.
Human footprints in cold, clear streams
The park with the highest visitor use, Denali, showed the greatest variety of resistance genes overall. Yet the creek that stood out most for sheer abundance of resistance DNA was Exit Creek in Kenai Fjords National Park, which flows below a busy visitor center and trailhead. There, the proportion of resistance genes relative to total bacterial DNA was significantly higher than at every other sampled site. By contrast, Wrangell–St. Elias, with far fewer visitors, showed fewer types of resistance genes, and some patterns hinted at other influences such as historical copper mining. Most of the bacterial groups identified by broad DNA sequencing were not classic human pathogens, and only a single cultured isolate from all sites showed clear resistance in traditional lab tests, underscoring that many resistance genes may reside in environmental microbes rather than in obvious disease-causing bacteria.
What this means for protecting people and parks
This work shows that it is possible to use a standardized, field-ready combination of large-volume filtration and targeted genetic testing to monitor antibiotic resistance genes in surface waters, even where contamination is low. By comparing both how often resistance genes appear and how abundant they are, managers can begin to distinguish between natural background levels and likely human influence from visitors, wastewater, or legacy pollution. While the presence of resistance genes alone does not prove an immediate health threat, tracking them over time offers an early-warning system and a way to evaluate how well parks are protecting both ecosystems and downstream communities in a world where antibiotic resistance is a growing concern.
Citation: Scott, L.C., Ahlstrom, C.A., Woksepp, H. et al. Validation and application of a standardized quantitative PCR assay for the assessment of antimicrobial resistance genes in surface water. Sci Rep 16, 11597 (2026). https://doi.org/10.1038/s41598-026-35635-x
Keywords: antimicrobial resistance, surface water, national parks, qPCR monitoring, water quality