Clear Sky Science · en
Whole-protein screening and multi-modal profiling of antigen-specific CD4+ T cells at single-cell resolution
Why this matters for health and cancer care
Our immune system relies on helper cells that coordinate attacks on viruses and tumors, but scientists have struggled to track these cells in detail. This study introduces a powerful lab method that can spot rare helper T cells in blood, figure out exactly what they recognize, and read out their behavior one cell at a time. The work could sharpen vaccine design and support new cell therapies for infections like COVID-19 and cancers linked to human papillomavirus (HPV).
Finding the immune conductors
Helper T cells, also called CD4+ T cells, act like conductors of the immune orchestra. They respond when other cells show them short pieces of viruses or cancer proteins, and in turn they help antibody-producing B cells, killer T cells, and other defenders do their jobs. Studying these conductors has been hard because each one recognizes a slightly different protein piece, and the tools to find them in blood either disturb their natural state or can only test a small number of targets at once. The authors set out to build a method that could search entire proteins, catch the relevant helper cells directly from blood, and read both their genetic programs and their unique T cell receptors in a single sweep.

Building a protein library for helper cells
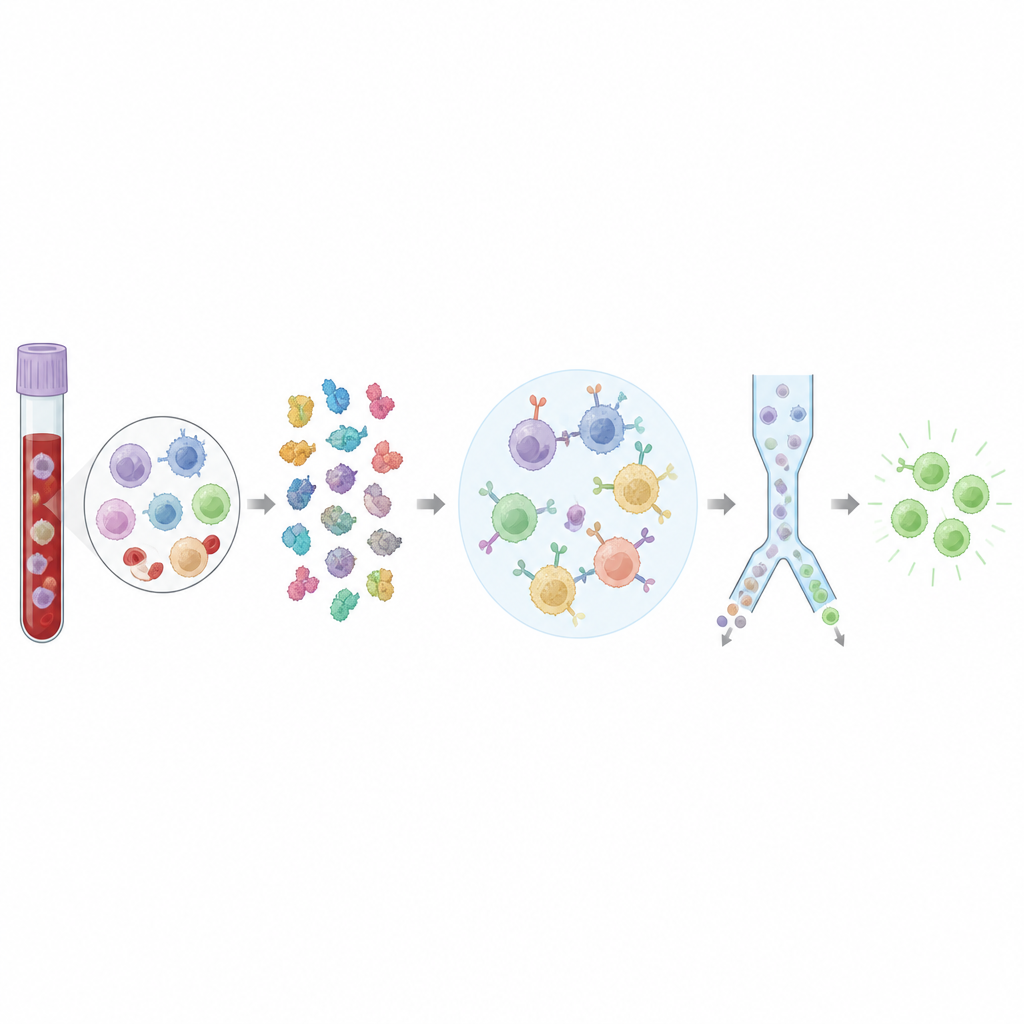
The team engineered a large library of artificial protein complexes that mimic what immune cells show to helper T cells. Each complex holds a specific protein fragment from a virus or cancer protein in a fixed shape, attached to a human display molecule. By linking the parts into a single chain, they made it easy to mass-produce hundreds of distinct complexes in human cell cultures without having to make each peptide separately. They showed that these complexes bind only to matching helper T cells, work across several common immune gene types, and can detect even rare cells in blood at very low frequencies. Importantly, binding to these complexes also predicted that the same T cells would respond when targets were presented on living cells, not just in artificial tests.
Tracking COVID-19 helper cells over time
With this platform in hand, the researchers scanned the entire receptor binding domain of the SARS-CoV-2 spike protein, along with other viral regions, in blood samples from 22 people who had COVID-19. Using barcodes and single-cell sequencing, they captured 2,188 virus-specific helper T cells and learned, for each cell, which viral piece it recognized, which HLA gene presented it, what its T cell receptor looked like, and what genes it was expressing. They saw a rich mix of cell states, including memory-like, effector, regulatory, and exhausted-like cells. Over time, responses tended to shift toward effector memory cells, but in a way that differed from previously studied killer T cells. The team created an “immunogenicity score” combining how many people and T cell clones responded to a given viral piece, how much those clones expanded, and how long they persisted. Higher scores for spike receptor binding domain fragments were linked with stronger antibody levels later on, suggesting that these helper cells may support better antibody quality.
Mining HPV helper cells for cancer therapy
The researchers then turned to cancer, focusing on HPV-16, a major cause of cervical and other cancers. They built a library that tiled across two HPV proteins, E6 and E7, using fragments of different lengths to probe how “flanking” regions around a central core affect helper T cell recognition. They applied this library to patients with precancerous lesions who had received therapeutic HPV vaccines, and they identified dozens of HPV-specific helper T cells. From these, they selected a panel of T cell receptors for deeper testing. Some receptors showed strong and selective responses when transferred into human T cells, producing multiple helpful signals and killing target cells loaded with HPV fragments or full-length E6 protein, while showing little to no reaction to similar human proteins or to a broad panel of other immune gene types. One receptor in particular, called H2 in the study, emerged as a promising candidate for future engineered T cell therapies.
What this means for future vaccines and therapies
For non-specialists, the key takeaway is that the authors have built a kind of “immune search engine” for helper T cells. It can look across whole proteins from viruses or cancers, pull out the rare cells that recognize specific pieces, and simultaneously reveal how those cells behave and how they might be harnessed. In COVID-19, this approach linked certain spike fragments with both strong helper cell responses and higher antibody levels, offering clues for refining vaccines. In HPV-related disease, it uncovered helper T cell receptors that look both powerful and safe enough to consider for targeted cancer immunotherapy. While more work is needed to extend the method to additional immune gene types and larger patient groups, this platform opens a practical route to map and eventually direct the helper arm of the immune system in many diseases.
Citation: Zhang, R., Qi, J., McKasson, M. et al. Whole-protein screening and multi-modal profiling of antigen-specific CD4+ T cells at single-cell resolution. Nat Commun 17, 3979 (2026). https://doi.org/10.1038/s41467-026-72396-7
Keywords: CD4 T cells, single-cell profiling, SARS-CoV-2 immunity, HPV immunotherapy, T cell receptors