Clear Sky Science · en
Expression of nano-engineered RNA organelles in bacteria
Tiny factories inside simple cells
Even the humblest bacteria carry out a staggering number of chemical reactions. Scientists are now learning to add new, man‑made “workstations” inside these cells—tiny droplet‑like compartments that can concentrate chosen molecules. This paper shows how to build such compartments from RNA, the same type of molecule that normally carries genetic messages, and to make them appear, disappear, and capture proteins on command inside living bacteria.
Droplets without walls
Many cells organize their chemistry using membraneless organelles—droplets of proteins and nucleic acids that form by phase separation, a bit like oil droplets in water. Bacteria were once thought too simple for such internal structure, but they also use similar droplets to control DNA copying, stress responses, and more. Researchers hope to design artificial versions of these compartments to steer metabolism, assemble useful products, or study how natural droplets work. Existing synthetic systems often rely on disordered proteins, whose sticky interactions are hard to predict and engineer.
Building with programmable RNA shapes
The authors turn instead to RNA nanotechnology, which exploits the predictable base‑pairing rules of nucleic acids. They designed “nanostars”: four‑armed RNA junctions whose tips carry short loops that can pair with matching loops on other nanostars. Two designs, called A and B, each have self‑complementary loops, so nanostars of the same type attract and condense into droplets. Scrambled versions lack this complementarity and should stay dissolved. The team inserted genetic blueprints for these nanostars into E. coli, using fluorescent RNA tags so the structures could be seen under the microscope.
Designer droplets inside living bacteria
When nanostars A or B were expressed, bright droplets appeared inside the bacterial cells, mostly at the poles. Analysis of fluorescence patterns and time‑lapse movies showed that most cells contained two or three droplets that could merge and re‑grow, behaving like tiny liquids. By contrast, the scrambled designs produced diffuse glow with only occasional weak clustering, confirming that specific base pairing drives droplet formation. Despite natural RNA‑cutting enzymes trimming some of the nanostar arms, enough four‑ and three‑armed structures remained to sustain robust condensation, highlighting the resilience of the design.
Multiple droplet types and protein capture
Because A and B nanostars recognize only their own kind, cells engineered to make both produced two non‑mixing droplet types in the same bacterium. These stayed largely separate in space, often at opposite ends of the cell. The authors then added a new feature: a short RNA aptamer within nanostar A that binds green fluorescent protein (GFP). When bacteria produced both the modified nanostars and GFP, the protein became strongly concentrated inside the droplets, while in control cells without the aptamer it stayed evenly spread. Measurements of fluorescence recovery after photobleaching revealed that GFP and nanostars both continued to exchange with the surrounding cytoplasm, again consistent with a liquid‑like state.
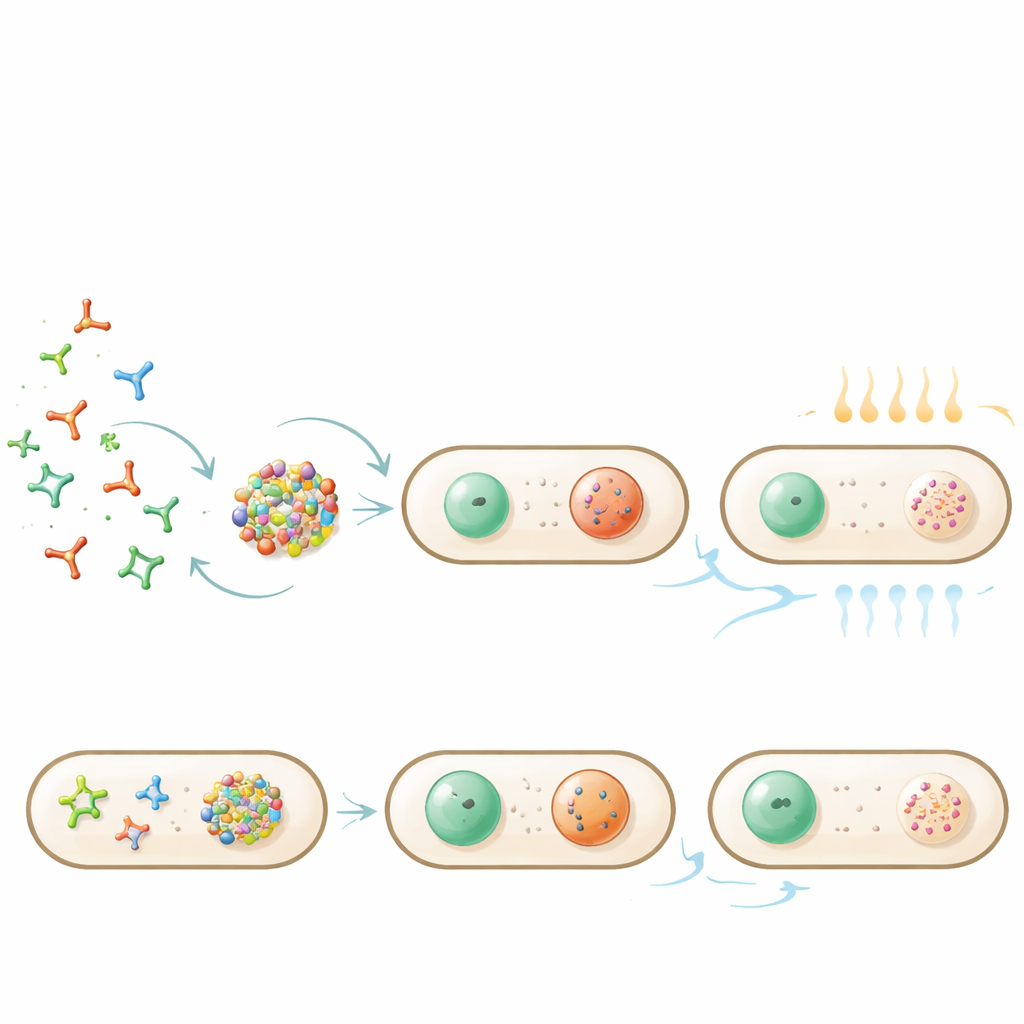
Switching droplets on and off with heat
Because the nanostars stick together through base pairing, temperature provides a simple control knob. Gently heating the bacteria caused the droplets to dissolve as the base pairs melted; cooling made them reappear in their usual polar positions. The precise “melting” temperature depended on the nanostar design and on how much enzymatic trimming had occurred, but the process remained reversible for many cells. When droplets containing captured GFP were heated more strongly, both RNA and protein dispersed throughout the cell; after cooling, droplets re‑formed and again concentrated the protein, showing that cargo can be released and re‑sequestered by a simple thermal cycle.
What this means for future cell engineering
In everyday terms, the researchers have built programmable, RNA‑based “pop‑up rooms” inside bacteria—rooms that can be duplicated, kept separate, filled with selected tools, and folded away on demand. Because the underlying interactions are simple and designable, these synthetic organelles offer a flexible way to route metabolic pathways, isolate toxic intermediates, or probe how natural droplets control life’s chemistry. The work suggests that nano‑engineered RNA condensates could become core components of next‑generation microbial factories and powerful tools for reshaping cellular behavior.
Citation: Ng, B., Fan, C., Dordevic, M. et al. Expression of nano-engineered RNA organelles in bacteria. Nat Commun 17, 2752 (2026). https://doi.org/10.1038/s41467-026-69336-w
Keywords: synthetic organelles, RNA nanotechnology, biomolecular condensates, bacterial cell engineering, metabolic engineering