Clear Sky Science · en
An information-theoretic argument for the restriction of the current biological alphabets to 4 nucleotides and 20 amino acids
Why life’s tiny alphabets matter
All life on Earth writes its genetic and protein "texts" using surprisingly small alphabets: just four chemical letters for DNA and RNA, and twenty for proteins. Chemists can imagine many more building blocks, so why does biology stick to these restricted sets? This paper argues that the answer lies in how easily these molecules can fold into useful shapes and how efficiently evolution can search through all possible sequences. By connecting ideas from physics and information theory, the authors show that the familiar alphabets of life are just large enough for molecules to fold reliably and still be evolvable.
From tangled strings to useful shapes
Proteins and RNA start as flexible chains that jiggle through countless possible shapes. Only a tiny fraction of those shapes are stable and functional. For life to work, a chain must quickly find its correct folded form without trying every possibility at random. The authors use an information-theory lens: when a chain folds, it "gains information" by selecting one native shape out of many alternatives. This gain can be measured as how much the range of possible shapes shrinks per position along the chain. They compare this with the information gained when evolution winnows random sequences down to those that actually fold, showing that both processes must be balanced for folding to be fast and reliable.
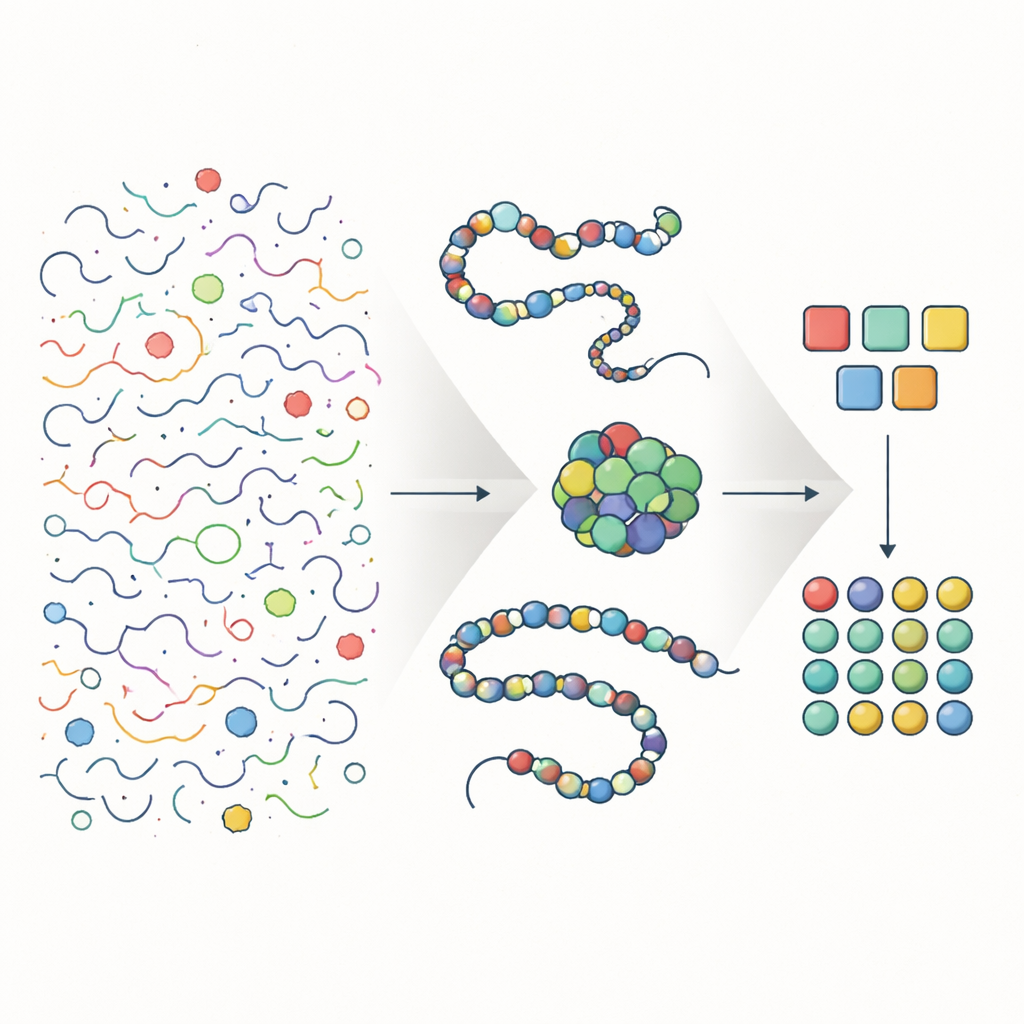
Matching digital letters to physical motions
The key insight is a simple mathematical link between three things: the size of the chemical alphabet, how many shapes each position in an unfolded chain can adopt, and how many different building blocks effectively appear at that position in real, evolved molecules. For a polymer that folds to a well-defined structure, the theory predicts that the number of accessible unfolded shapes per position, and the effective diversity of letters used there, should both be roughly equal to the square root of the total alphabet size. When the authors plug in measurements from real proteins and RNA, they find that the average number of unfolded shapes per position and the effective variety of letters per position line up closely with this prediction, for both biopolymer types.
Why four nucleotides and about twenty amino acids
For RNA, experimental studies of backbone flexibility and base-pair usage suggest that each nucleotide has around two and a half relevant unfolded shapes. Squaring this value gives an alphabet size very close to four, precisely what life uses. For proteins, estimates of backbone freedom and sequence variation imply about four to five effective shapes and effective letters per position, which points to an optimal alphabet in the range of roughly twenty or a few dozen amino acids. The fact that modern biology uses twenty chemically distinct amino acids sits comfortably at the lower end of this range, consistent with additional practical limits such as how complex the protein-making machinery can be and how many different side-chain types can be kept reliably distinct.
Clues to life’s early, floppy proteins
The authors then turn this framework into a window on early evolution. They combine their formulas with previous reconstructions of when different amino acids entered the genetic code. In the earliest stages, the alphabet appears to have been too small to support stable, neatly folded proteins at all. Instead, the theory predicts chains that remained highly flexible and disordered, yet could still clump into droplets or loose networks thought to be important for primitive, membraneless cell-like structures. As more amino acids were added, the alphabet crossed a threshold where folded proteins became possible, first favoring intrinsically disordered but functional chains, and only later allowing sharply defined three-dimensional structures and efficient catalysts.
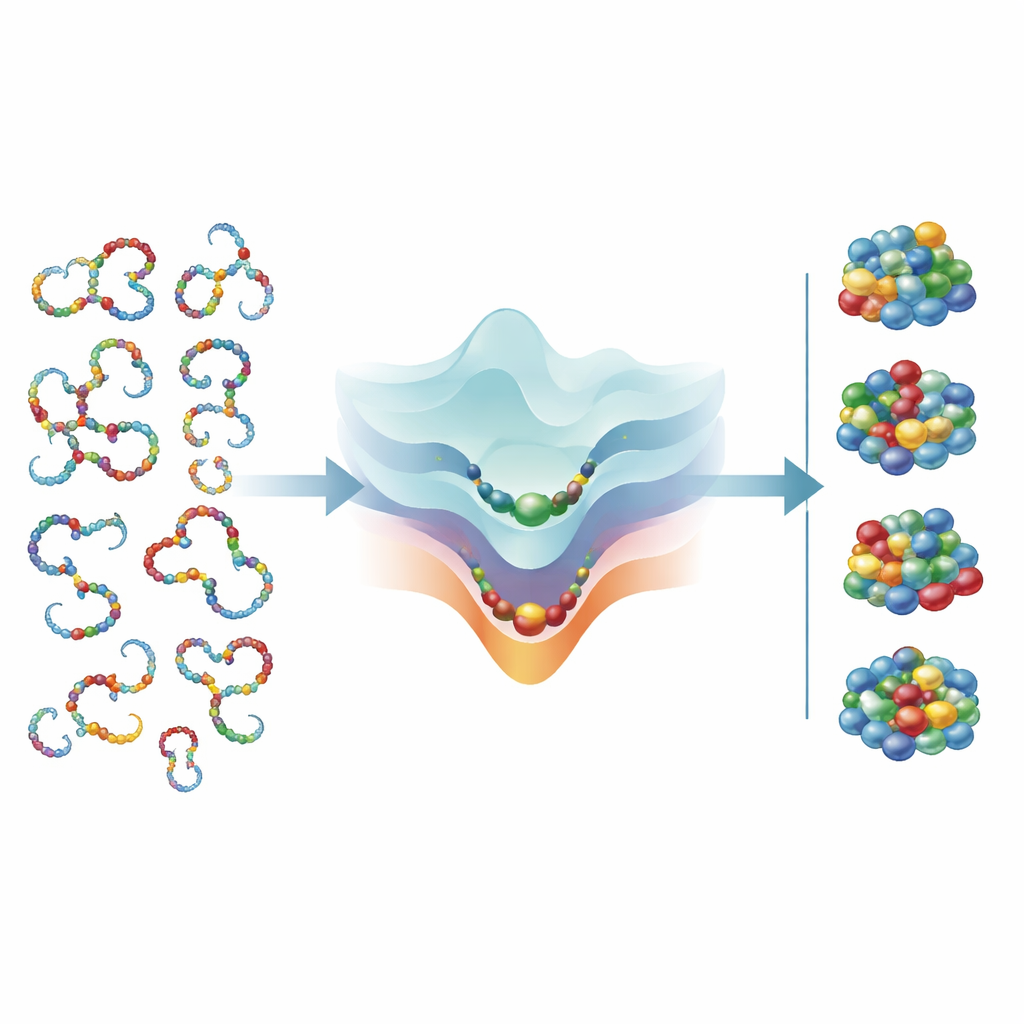
What this means for life’s limits
In everyday terms, the study suggests that there is a sweet spot between having too few chemical letters, which makes it hard to encode specific shapes, and too many, which makes the search for workable molecules impossibly slow. Earth’s four nucleotides and twenty amino acids sit very near that sweet spot, given how floppy these chains naturally are in water. Below those alphabet sizes, evolution would struggle to find well-folded molecules; above them, added letters bring little advantage because a single stable structure can already be encoded. In this view, life’s alphabets are not arbitrary: they are near-minimal solutions that let information-rich molecules both fold quickly and evolve efficiently.
Citation: Galpern, E.A., Ferreiro, D.U. & Sánchez, I.E. An information-theoretic argument for the restriction of the current biological alphabets to 4 nucleotides and 20 amino acids. Sci Rep 16, 10751 (2026). https://doi.org/10.1038/s41598-026-46009-8
Keywords: genetic code, protein folding, RNA structure, molecular evolution, biopolymer alphabets