Clear Sky Science · en
Memory efficient training for 3D brain image registration networks using PatchMorph
Making Sense of Many Brain Scans
Modern brain research depends on comparing 3D scans from many people and animals, but each brain image is slightly different in size, shape, and resolution. Lining them up accurately, without needing enormous computers, is a major challenge. This paper introduces PatchMorph, a new way to align large 3D brain images that dramatically cuts memory use while keeping or even improving accuracy, opening the door to studying bigger and richer brain datasets.
Why Lining Up Brains Is So Hard
To compare brains, scientists first “register” one 3D image to another, warping one volume so that matching structures overlap. Classical tools do this by slowly adjusting a deformation field until a similarity score between the images is maximized. Newer deep learning methods such as VoxelMorph learn to predict this warp in one shot, making registration far faster. However, most of these networks expect both input images to share the same grid size and resolution, and they often need to process full 3D volumes at once. As image sizes and model complexity grow, the required graphics memory quickly explodes, forcing researchers to shrink, crop, or heavily pre-process their data.
Looking at Brains Piece by Piece
PatchMorph tackles this bottleneck by never loading the full 3D volumes into the network at once. Instead, it works on small 3D “patches” sampled throughout a shared reference space that covers the fixed image. At a coarse level, the method places relatively large patches over the whole brain, extracts matching patches from the moving image using precise coordinate transforms, and feeds them to a compact registration network. The network predicts how each patch must shift and warp so that its local content matches. These many local predictions are then scattered back into a global deformation field, using smooth interpolation so that the whole brain warp remains continuous.
From Big Patches to Fine Details
PatchMorph proceeds in multiple scales, from coarse to fine. At the first scale, a patch covers a large brain region at low resolution, capturing broad misalignments like overall size or position. At the next scales, new patches “zoom in” on subregions of previously sampled patches, but always keep the same number of voxels per patch in memory. This means finer scales increase physical detail without increasing memory demand. Patch positions at finer levels are chosen stochastically within their parent patches, so the method does not just focus on patch centers but gradually learns to align peripheral structures as well. Across scales, the estimated deformations are accumulated into a single global field, refined step by step yet always spatially consistent.
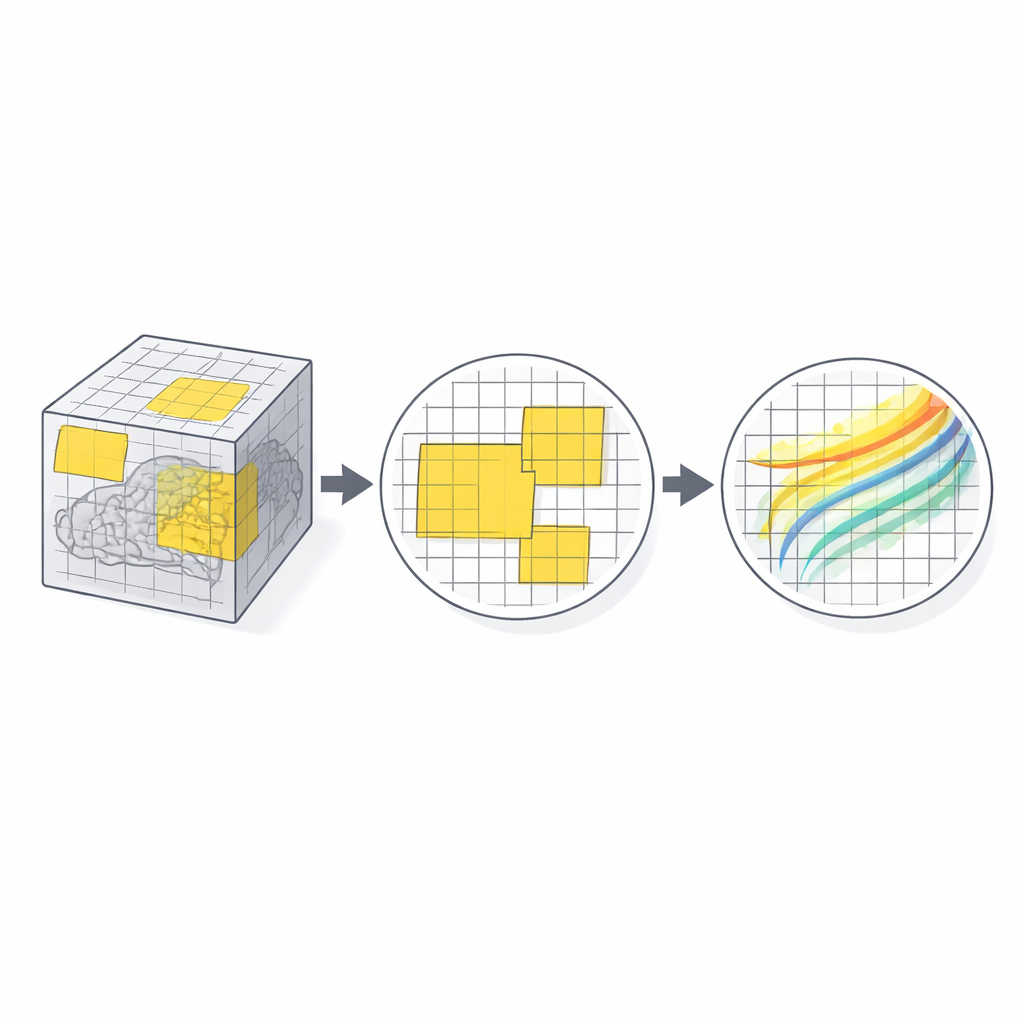
Working With Real-World Brain Data
The authors tested PatchMorph on two demanding datasets: human T1 MRI scans from the MindBoggle project and high-resolution marmoset brain images from serial two-photon microscopy. These images differ in array size, orientation, and sometimes voxel spacing—conditions that strain conventional networks. PatchMorph matched or surpassed state-of-the-art deep learning registration methods in terms of overlap between labeled brain regions, while maintaining very low rates of topological errors (unrealistic foldings in the deformation field). Crucially, when training with transformer-based backbones, PatchMorph reduced the memory needed for 256³ voxel images from about 40 GB to under 10 GB, without sacrificing accuracy.
Speed, Trade-Offs, and Future Uses
Because PatchMorph must extract and recombine many patches, it is somewhat slower at inference than single-shot full-volume networks, though still much faster than traditional iterative tools. In return, it can handle large, high-resolution, and unevenly sampled images directly in their native space, and it can reuse a variety of backbone architectures, from simple convolutional networks to advanced transformers. This makes it easier to bring powerful deep learning models to real-world brain imaging studies that previously did not fit into memory. In practical terms, PatchMorph shows that by thinking in patches instead of whole volumes, researchers can align complex 3D brain data accurately, efficiently, and at scales that were previously out of reach.
Citation: Skibbe, H., Byra, M., Watakabe, A. et al. Memory efficient training for 3D brain image registration networks using PatchMorph. Sci Rep 16, 14386 (2026). https://doi.org/10.1038/s41598-026-44858-x
Keywords: brain image registration, deep learning, medical imaging, 3D MRI, PatchMorph