Clear Sky Science · en
Host preference and specialization in the genus Aphanomyces (Oomycetes) from molecular and interaction network insights
Invisible attackers in water and soil
Many of the world’s crops, fish, and freshwater crustaceans are quietly threatened by microscopic organisms that look like fungi but are actually closer relatives of algae. Among them, the genus Aphanomyces stands out for causing devastating diseases such as crayfish plague and major root rots in peas, beans, and other crops. This study asks a simple but crucial question: which hosts do these organisms really prefer, and how narrowly are they specialized? By combining modern DNA tools with network analysis, the authors map out who infects whom across plants and animals, revealing hidden diversity and sharp divisions in host use.
Tracking hidden diversity with DNA
To understand how many distinct kinds of Aphanomyces there are, the researchers first built an extensive genetic catalogue. They gathered 261 DNA sequences (from a standard marker region) representing known species of Aphanomyces and its close relatives, plus many unnamed strains. Using several complementary algorithms, they asked where natural “breaks” occur in genetic similarity that likely correspond to separate species. Most of the methods agreed on 34 candidate species: 20 matching already described names and 14 representing putative newcomers. In some cases, what had long been treated as a single species, such as Aphanomyces stellatus, actually resolved into three distinct genetic lineages, hinting at cryptic species that had gone unrecognized under the microscope.
Family tree of plant and animal invaders
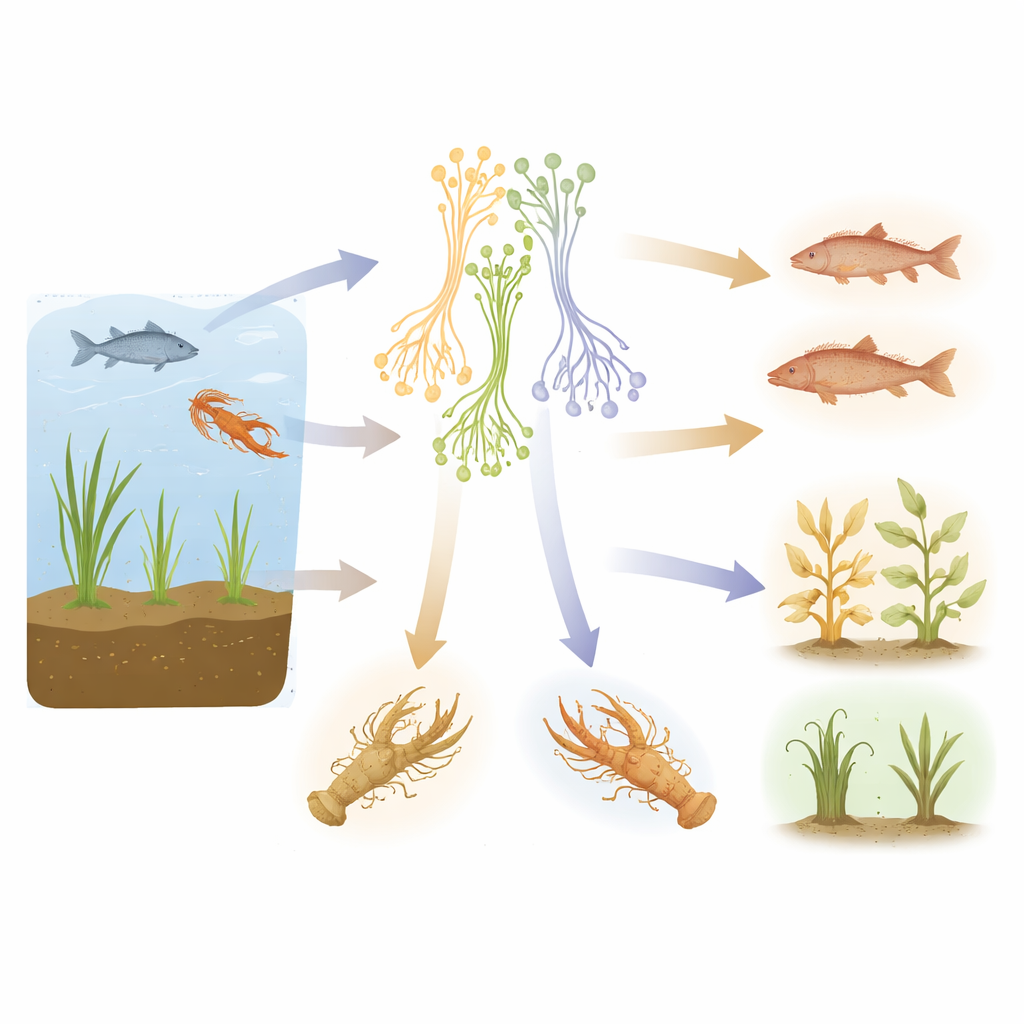
Next, the team reconstructed a family tree for these organisms. The resulting evolutionary portrait showed three major groups. One group includes species that mainly live in water and often infect freshwater animals such as crayfish and fish. A second group is dominated by species that live in wet soils and attack plant roots, including several notorious crop pathogens. A third group contains related genera that are also linked to plants and damp soils. Together, these patterns indicate that the history of Aphanomyces has been strongly shaped by shifts between life in water and life around plant roots, and by repeated tightening of host preferences. The placement of some close relatives within the Aphanomyces branches also suggests the current genus boundaries may need to be revised.
Reconstructing a web of infections
Genetics alone cannot reveal how these organisms behave in nature, so the authors turned to the literature. They compiled 1,221 records from more than a century of studies, each documenting a particular Aphanomyces-like organism found on a given host or substrate, from crayfish gills and fish skin to crop roots and lake sediments. Treating this as a network, they drew links between pathogen species on one side and host families or environmental substrates on the other. They then quantified how densely connected the network is, how much it falls into separate clusters, and how specialized each pathogen appears compared with a random expectation.
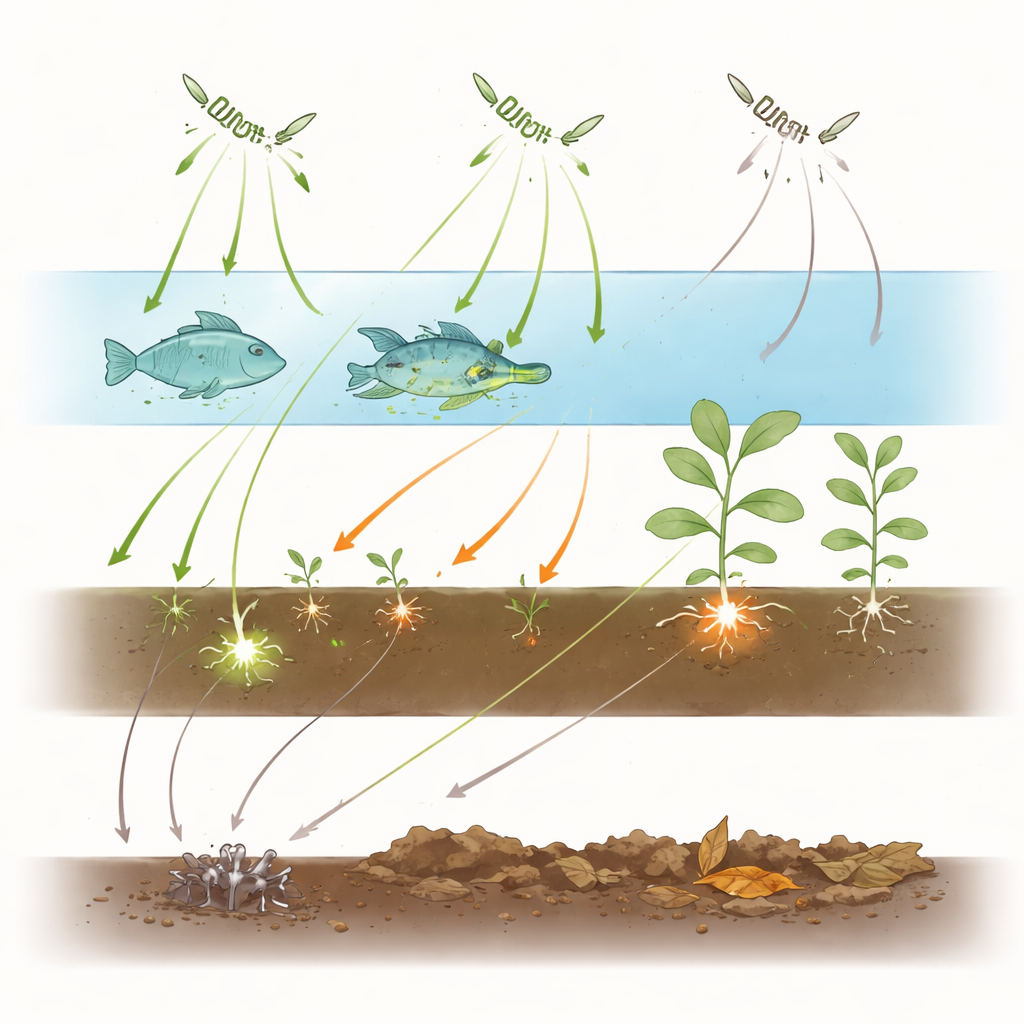
Strong preferences and tight clusters
The infection network turned out to be sparse and strongly clustered, patterns typical of close, intimate relationships like those between parasites and their hosts. Rather than forming a single, highly interconnected web, the network breaks into modules that correspond to broad ecological niches: one dominated by plant pathogens in wet soils, another by parasites of freshwater crayfish and related crustaceans, a separate one centered on fish, and yet another around a soft-shelled turtle host. When the authors calculated a specialization score for each species, generalist, free-living forms in water or soil scored low, while classic disease agents such as Aphanomyces euteiches in legumes, A. cochlioides in sugar beet, and A. astaci and A. invadans in crayfish and fish scored very high, indicating a narrow focus on particular host groups.
Why this matters for food and ecosystems
For non-specialists, the key message is that these fungus-like organisms are not vague, opportunistic threats—they tend to be sharply tuned to particular hosts and habitats. The study uncovers many likely new species and shows that dangerous pathogens of fish, crayfish, and crops sit within a larger, highly structured web of mostly specialized lineages. This more resolved picture of who infects whom can help scientists predict which hosts might be at risk when an Aphanomyces species moves into a new region, improve early-warning systems for aquaculture and agriculture, and guide future work on how these microscopic attackers evolve to exploit their plant and animal targets.
Citation: Casabella-Herrero, G., Martín-Torrijos, L., Pérez-Ortega, S. et al. Host preference and specialization in the genus Aphanomyces (Oomycetes) from molecular and interaction network insights. Sci Rep 16, 14262 (2026). https://doi.org/10.1038/s41598-026-44513-5
Keywords: Aphanomyces, oomycete pathogens, host specialization, plant and fish diseases, interaction networks