Clear Sky Science · en
Miller–Ross-functions coefficients and kernel-based CT-scan enhancement technique
Sharper Brain Scans for Clearer Answers
Computed tomography (CT) scans are central to diagnosing brain disorders, but the raw images can be noisy, blurry, or low in contrast—especially when doctors try to reduce radiation dose. This paper explores a mathematically driven way to sharpen brain CT images before they are analyzed by radiologists or computer algorithms. Instead of relying on large neural networks, the authors use a carefully designed 3 × 3 image filter built from a special family of mathematical functions, aiming for clearer structure and better visibility of fine details in medical scans.
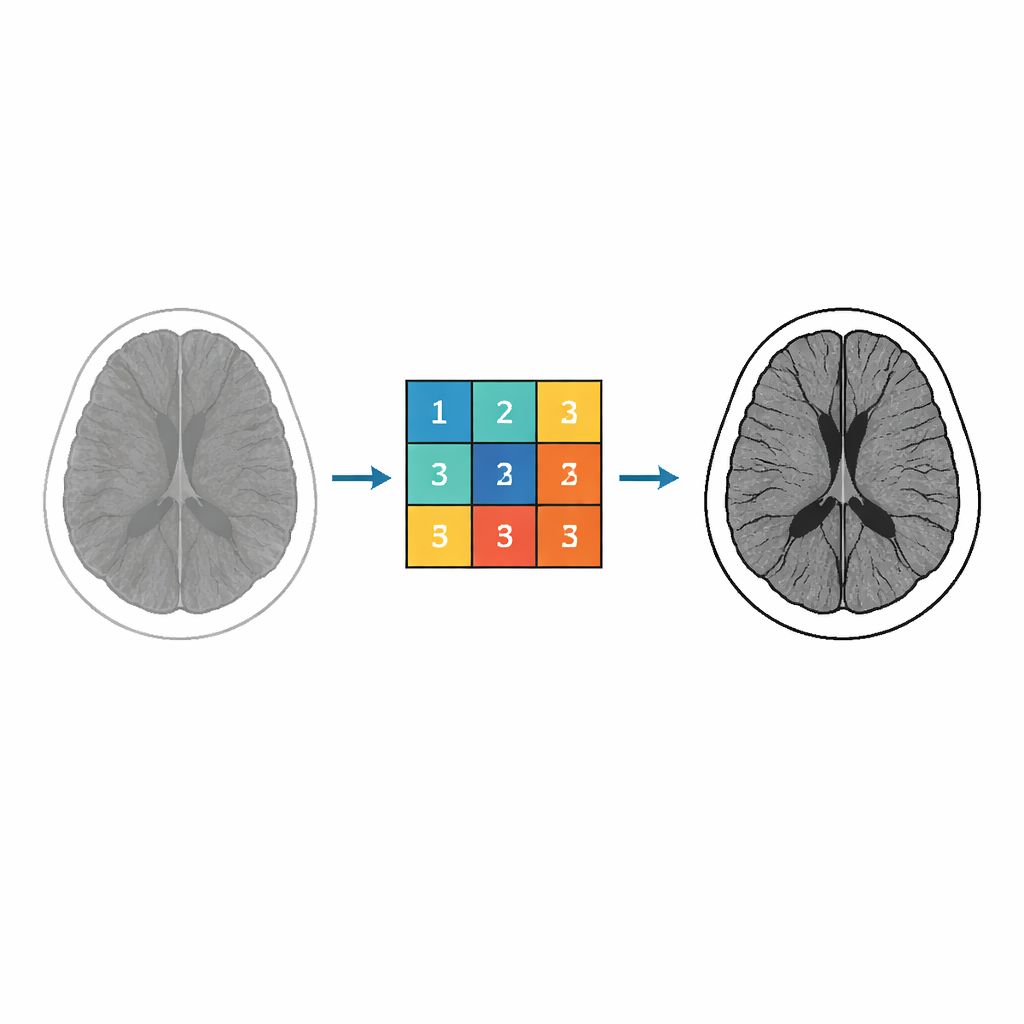
From Abstract Math to Image Filters
At the heart of the work lies a class of complex-valued functions known as Miller–Ross functions, originally developed to solve fractional differential equations. Mathematicians study these functions for their geometric properties, such as how they map one region of the complex plane to another without overlaps. The authors harness these properties to control how pixel intensities in an image are transformed. By analyzing and bounding the coefficients in the series expansion of these functions, they derive explicit numerical values that can be used as weights in an image filter. These weights are then assembled into a 3 × 3 kernel—a tiny grid that slides across the image, combining nearby pixel values to enhance local structure.
Building a Tunable Enhancement Kernel
The key idea is to turn several abstract parameters appearing in the Miller–Ross framework into knobs that determine how the filter behaves. Different parameters influence how strongly edges are emphasized, how much noise is suppressed, and how fine details are treated. Using inequalities they prove for the function coefficients, the authors generate nine numbers that populate the 3 × 3 kernel. They focus in particular on a fractional parameter that strongly affects how subtle structures in the brain—such as tissue boundaries and small lesions—are brought out. By adjusting this parameter and others, they can move between gentler and more aggressive enhancement while staying within mathematically controlled limits.

Putting the Method to the Test on Real CT Data
To assess their approach, the researchers apply these Miller–Ross–based kernels to 155 infected brain CT images drawn from a public dataset. They design three main experimental scenarios, each fixing most parameters while varying one key quantity: in the first, they tune a contrast-like parameter; in the second and third, they vary another fractional parameter under two different settings. For each CT image, they search for the parameter value that gives the best improvement. The quality of the enhanced images is measured using two standard metrics: Peak Signal-to-Noise Ratio (PSNR), which gauges how much the image differs from a reference, and Structural Similarity Index (SSIM), which reflects how well important patterns and structures are preserved. In the best cases, the method produces remarkably high PSNR and SSIM values, with some images reaching SSIM close to 0.99, indicating very strong structural fidelity.
How It Compares to Modern AI Methods
The authors then place their results alongside a range of established CT enhancement and denoising techniques, including popular deep learning models such as convolutional neural networks, generative adversarial networks, and residual networks, as well as classic approaches like non-local means and BM3D filtering. On average, these deep models still achieve higher and more stable PSNR and SSIM scores, reflecting their strength when trained on large datasets. However, the new Miller–Ross–based approach shows a wide performance range: while some images are less impressive, others exceed the best reported scores from traditional filters and rival those of more complex systems. This variability underscores the method’s sensitivity to parameter choices, but also its potential when those choices are well tuned.
Promise and Practical Limits
The study concludes that the proposed kernel-based enhancement technique does not yet outperform top-tier deep learning methods across the board, but it offers important advantages. Because the filter is built directly from transparent mathematical formulas, clinicians and engineers can understand and control how each parameter affects the outcome—something that is often difficult with black-box neural networks. The method is especially good at highlighting fine structural details that may aid diagnosis, even when global quality scores do not always rise. The authors note limitations, including dependence on a small dataset, reliance on numerical metrics alone, and the need for careful parameter tuning, and they call for future work that adds clinical feedback and larger studies. Still, the approach illustrates how ideas from pure mathematics can be translated into practical tools for making life-saving images clearer and more informative.
Citation: Murugusundaramoorthy, G., Nalliah, M. Miller–Ross-functions coefficients and kernel-based CT-scan enhancement technique. Sci Rep 16, 13527 (2026). https://doi.org/10.1038/s41598-026-41609-w
Keywords: CT image enhancement, brain imaging, image filtering, fractional calculus, medical image processing