Clear Sky Science · en
NvashA function reveals temporal differences in neural subtype generation in cnidarians
How Simple Nerve Nets Tell an Ancient Story
The starlet sea anemone may look simple, but its nerve net holds clues to how all animal brains first evolved. This study asks a surprisingly modern question in a very ancient creature: not just where nerve cells form, but when different types of neurons are born, and whether timing helps diversify the nervous system. By tracking waves of newborn nerve cells over several days of development, the authors show that this humble sea animal uses time, as well as space, to build a rich cast of neural cell types—hinting that this strategy may date back to the common ancestor of most animals with nerves.
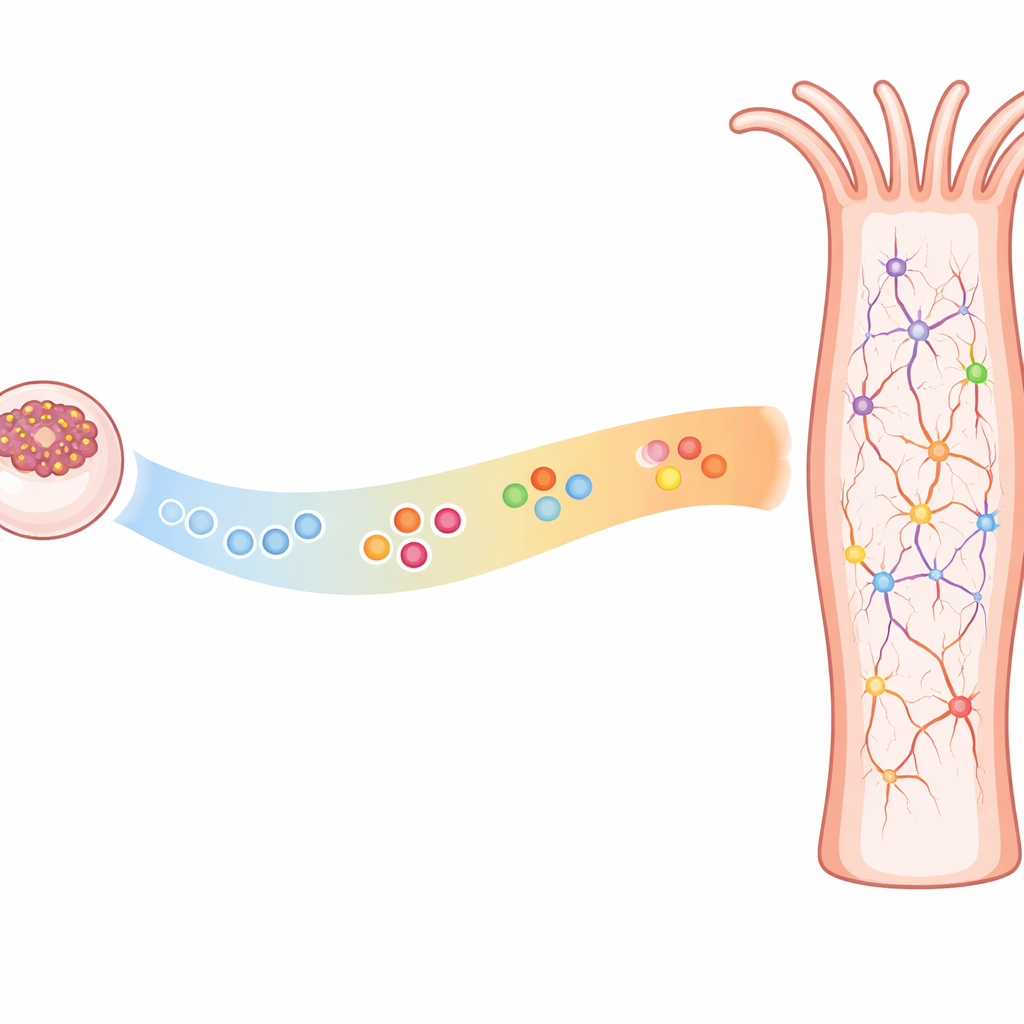
Building a Nerve Net Over Time
Cnidarians like anemones and jellyfish have a diffuse nerve net rather than a central brain, yet they share many of the same genetic tools that shape nervous systems in more complex animals. Earlier work showed that sea anemones and bilaterally symmetric animals (such as flies and mammals) use similar gene networks to turn on neural identity, choose progenitor cells, and map broad body regions where certain neurons arise. What was not clear was whether timing also matters in these simple nerve nets—whether specific neuron subtypes appear in a set sequence as development unfolds, as they do in brains and spinal cords of more complex animals.
Following a Key Neural Switch
To explore this, the researchers focused on a gene called NvashA, known to switch on as immature neurons emerge from their parent cells and then fade as those neurons fully mature. Because NvashA spans this window from “just born” to “nearly finished,” its presence marks cells in the act of becoming neurons. The team used single-cell RNA sequencing to profile thousands of NvashA-positive cells collected from embryos and free-swimming larvae at four time points, then grouped these cells into clusters based on their gene expression signatures. They also cross-referenced these clusters against existing cell atlases to identify neuron types, secretory cells, and stinging cells.
Early and Late Waves of Neuron Types
The analysis revealed two broad groups of NvashA-marked neurons. One group contained cells present from early embryonic stages through later larval stages, including immature neurons and several previously described neuron subtypes that appear early and then continue to mature. The second group, in contrast, consisted almost entirely of neurons found only in larval stages. Within this later group, the team identified immature cells and several distinct neuron subtypes that seem to arise exclusively after the animal has already formed a basic body plan. Pseudotime and trajectory analyses—a way of ordering cells along developmental paths—showed branches radiating from these immature states into multiple specialized neuron types, with later-stage neurons clearly separated from the earlier-born set.
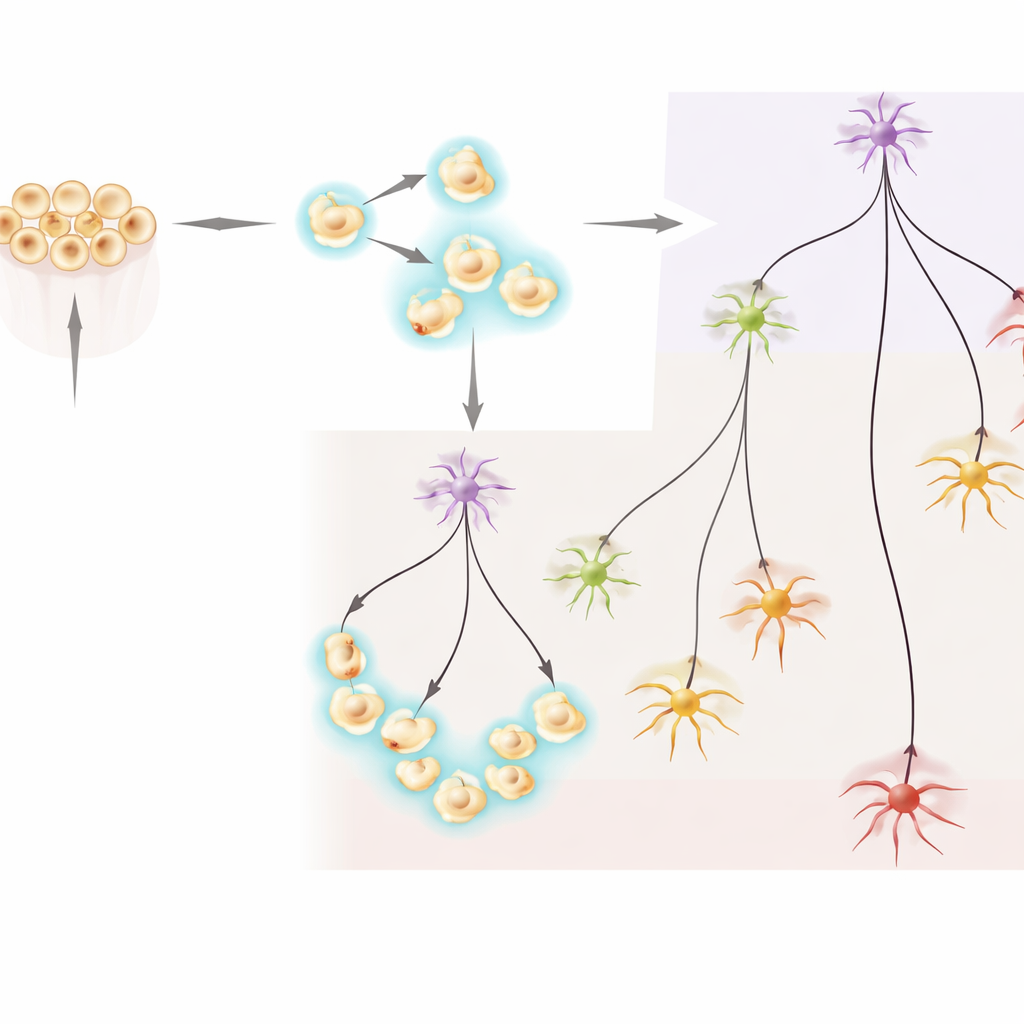
Where New Neurons Appear in the Body
The authors then asked where these late-forming neuron types appear along the animal’s main oral–aboral axis (from mouth to opposite end). Using in situ hybridization, they visualized marker genes that were enriched in specific late-stage neuronal clusters. These markers showed scattered patterns in larvae, but their positions lined up with body regions—like trunk and aboral territories—that are known to be set up much earlier in development. Some late neuron markers were confined mostly to the aboral end, while others were restricted to the trunk, echoing previously mapped spatial “stripes.” This indicates that the same spatial domains can generate different neuron types at different times, implying that timing information overlays pre-existing body patterning to expand neural diversity.
Testing What NvashA Actually Does
To determine how essential NvashA is for building these neurons, the researchers reduced or removed its activity using two approaches: short-hairpin RNA knockdown and a precise gene knockout created with CRISPR. In early embryos, lowering NvashA sharply reduced the expression of several known neural target genes, confirming its central role in early neurogenesis. At later larval stages, many of these same genes partially recovered, but more detailed time-course experiments showed that their activation was delayed when NvashA was impaired. Late-stage neuron subtype markers were also weakened, though not completely lost, in mutant animals. Together, these findings suggest that NvashA is not an on–off switch for whole neuron classes but instead helps control the timing and proper maturation of both early-born and late-born neural subtypes.
What This Means for the Origin of Brains
In plain terms, this work shows that a sea anemone’s nerve net is not a static grid of identical cells; it is built in waves, with different neuron types emerging at distinct times within specific body regions. That “time axis” of nervous system patterning—long known in organisms with complex brains—also operates in a simple, radially symmetric animal. This supports the idea that using both spatial cues (where in the body) and temporal cues (when during development) to generate a variety of neurons is an ancient strategy, likely present in the common ancestor of cnidarians and bilaterian animals. While the exact molecules that encode temporal information may differ across species, the underlying logic—reusing the same body regions at different times to make new neuron types—appears to be a deeply conserved solution for building diverse nervous systems.
Citation: Havrilak, J.A., Cheng, M., Al-Shaer, L. et al. NvashA function reveals temporal differences in neural subtype generation in cnidarians. Sci Rep 16, 12151 (2026). https://doi.org/10.1038/s41598-026-41460-z
Keywords: neurogenesis, sea anemone, nerve net, evolution of nervous systems, single-cell RNA sequencing