Clear Sky Science · en
A multi-modal dataset for insect biodiversity with imagery and DNA at the trap and individual level
Why tiny insects and big data matter
All over the world, insect populations are changing rapidly, with some groups declining before scientists have even had a chance to describe them. Traditional methods of sorting, naming, and counting insects rely on scarce expert time and painstaking work with microscopes. This article introduces a new kind of resource that brings together detailed photographs and DNA information for tens of thousands of tiny creatures caught in real field traps. By pairing biology with modern computer vision, the authors aim to speed up how we measure and monitor insect life on a changing planet.
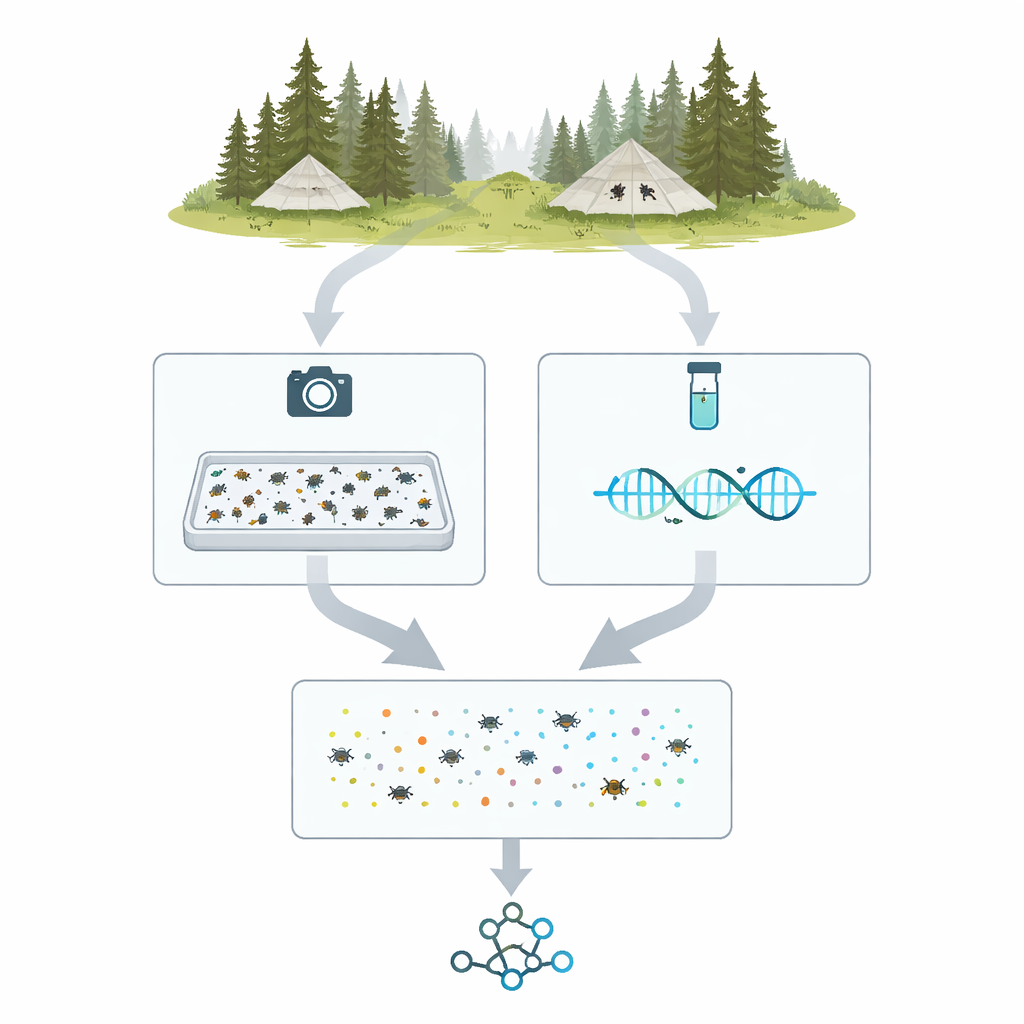
From field tents to digital specimens
The project, called MassID45, starts in northern forests and wetlands of Sweden and Finland, where special tent-like Malaise traps funnel flying insects into collecting bottles. Over the 2021 season, 45 weekly samples from 19 locations were chosen for deep analysis. In the lab, each mixed catch was weighed, gently processed to release DNA, and poured into a shallow tray with a thin layer of alcohol. The insects were spread out and photographed from above with a high‑resolution camera under carefully controlled lighting, creating a single "bulk image" in which thousands of individuals appear as pin‑sized shapes.
Seeing the same insects two ways
After taking bulk images, the team split the samples into individual insects for more detailed work. Each specimen was placed in its own tiny well or pinned and photographed close‑up. At the same time, a short, standardized stretch of DNA—often called a barcode—was read for each insect using modern high‑throughput sequencing machines. This yielded more than 35,000 individual barcode sequences. Comparing those sequences to large reference databases allowed the researchers to place most specimens into well‑known groups, such as flies, beetles, and moth families, providing a DNA‑anchored list of which types of arthropods occurred in each trap sample.
Teaching computers to find tiny creatures
To make the bulk tray photos useful for automation, the authors had to teach computers where each insect is and what broad group it belongs to. They used a two‑step annotation process. First, an algorithm roughly outlined every dark object in a tray image, then human annotators refined these outlines using an AI‑assisted web tool, ensuring that each insect—often only a handful of pixels wide—received its own clean mask. Second, an expert examined each masked insect and assigned it to the finest taxonomic level they could see from the photograph, guided by a custom list of expected groups derived from the matching DNA barcodes. This strategy concentrated expert effort on recognition rather than tedious drawing, and resulted in over 17,000 arthropods in the bulk images being linked to robust group names.
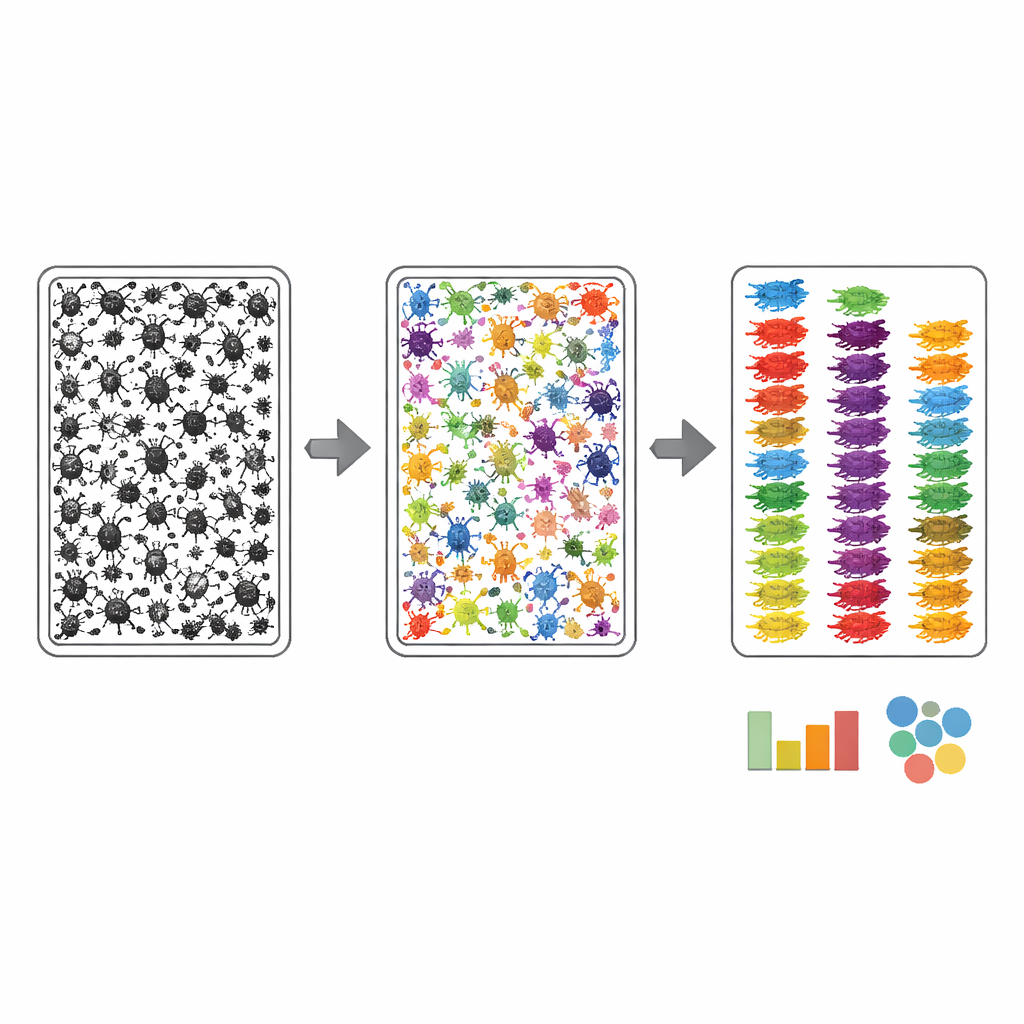
How well does the system work?
The team then treated MassID45 as a stress test for modern computer vision. Bulk images were split into overlapping tiles so that tiny insects remained sharp enough for analysis, and several state‑of‑the‑art image‑segmentation models were evaluated. General "zero‑shot" systems, which had never seen these data before, struggled: they tended to miss the smallest insects and confuse them with bits of debris. In contrast, models that were retrained on the carefully labeled MassID45 images did much better at finding and outlining individuals, especially common groups like flies and wasps. Even so, the very tiniest springtails and other pale, speck‑like forms often remained hard to distinguish from background material, highlighting an inherent visual limit.
What this means for tracking life on Earth
MassID45 is not a single new algorithm but a rich reference dataset that other researchers can download and build upon. By tying together tray‑level photos, individual specimen images, DNA sequences, and expert group labels from real field samples, it provides a realistic training ground for computers to learn how to count and characterize swarms of tiny arthropods. While the images rarely allow species‑level identification, they reliably capture broader groups, which are often enough to reveal shifts in insect communities over time and space. In practice, this means that future monitoring programs could combine simple trap photography with DNA sampling and machine learning to deliver faster, more detailed, and more scalable views of insect biodiversity than would ever be possible by human experts alone.
Citation: Orsholm, J., Quinto, J., Autto, H. et al. A multi-modal dataset for insect biodiversity with imagery and DNA at the trap and individual level. Sci Data 13, 630 (2026). https://doi.org/10.1038/s41597-026-07251-x
Keywords: insect biodiversity, DNA barcoding, computer vision, ecological monitoring, machine learning dataset