Clear Sky Science · en
CLWD: a Chinese histopathology dataset for lung adenocarcinoma subtype classification
Why a New Lung Cancer Image Collection Matters
Lung cancer remains one of the deadliest cancers worldwide, and in China it affects more people than anywhere else. Doctors now know that one common form, lung adenocarcinoma, is not a single disease but a patchwork of different growth patterns that carry very different risks for the patient. Distinguishing these patterns under the microscope is difficult, even for experts, and it takes time. This article introduces a new open dataset of high‑quality lung tissue images from Chinese patients, designed to help researchers build computer tools that can recognize these subtle patterns more consistently and, ultimately, support more accurate diagnosis and treatment.
The Challenge Inside the Lung
When a patient’s lung tumor is removed, pathologists slice the tissue into thin sections, stain them, and examine the slides under a microscope. In lung adenocarcinoma, these slides reveal several distinct ways that tumor cells grow and invade: some patterns are relatively gentle and associated with better outcomes, while others are aggressive and linked to higher chances of the cancer coming back. Current international guidelines group these patterns into categories such as in situ, acinar, papillary, lepidic, micropapillary, solid, and cribriform. Correctly identifying which pattern dominates in a tumor helps doctors estimate risk and decide how closely to monitor or treat a patient. However, this process is labor‑intensive and subject to disagreement between specialists.
Turning Glass Slides into Digital Data
Advances in digital scanners now allow entire microscope slides to be captured as enormous, detailed images that computers can analyze. Building reliable artificial‑intelligence tools, however, requires large, carefully labeled datasets that reflect real clinical practice. The authors created the Chinese Lung Adenocarcinoma WSI Dataset (CLWD) by collecting 408 stained slides from 210 patients treated at a major hospital in Yunnan Province between 2020 and 2023. Each slide was scanned at very high magnification, giving a level of detail comparable to what a pathologist sees through a microscope. Experienced lung cancer pathologists selected representative sections, verified staining quality and tissue integrity, and excluded any slides that were ambiguous or could be misinterpreted. Alongside the images, the team compiled de‑identified information such as patient age, sex, diagnostic category, and detailed growth pattern labels that are compatible with both 2015 and 2021 World Health Organization classifications.
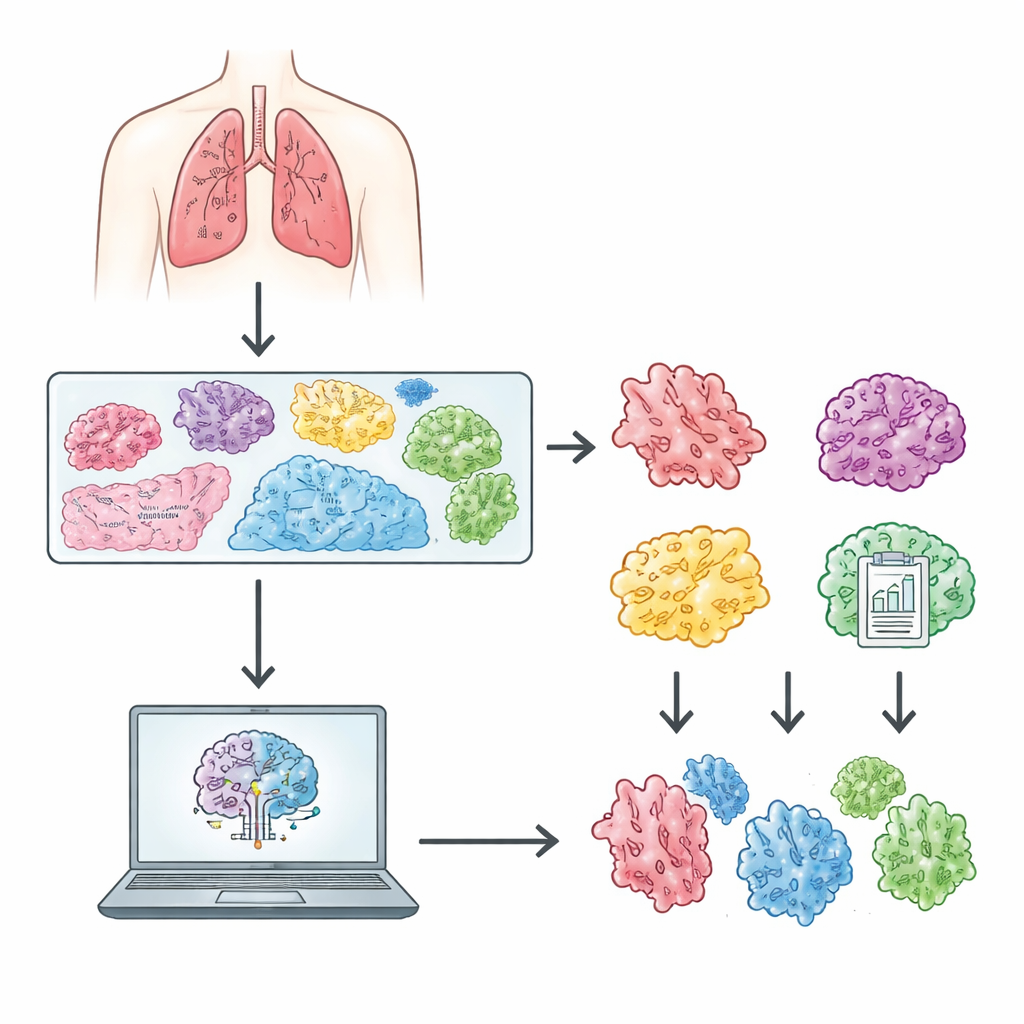
How Computers Learn from the Slides
The images in CLWD are so large that they cannot be fed into a neural network all at once. Instead, each whole‑slide image is automatically broken into many small square patches that contain only tissue, filtering out empty background and scanning artefacts. The study uses an approach known as multiple‑instance learning, in which all the patches from a slide are treated as a group. A pre‑trained neural network first extracts visual features from each patch, and then specialized models learn how to combine these features to decide which subtype label best fits the entire slide. The authors evaluated three modern attention‑based methods—CLAM, TransMIL, and a Graph Transformer—each designed to focus on the most informative regions and the relationships among patches. This framework mirrors how a human expert visually scans different regions of a slide before forming an overall judgment.
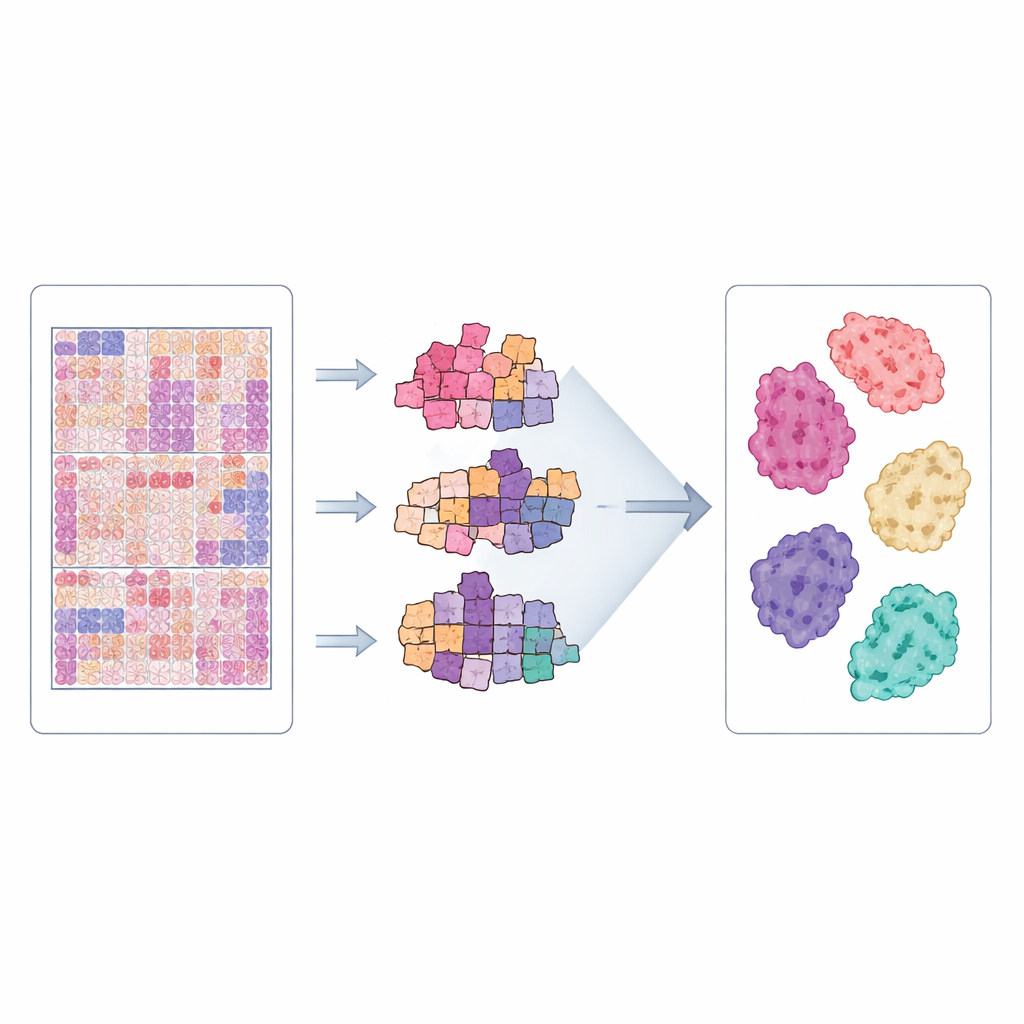
Putting the Dataset to the Test
To check whether CLWD is truly useful for computer‑aided diagnosis, the team ran extensive experiments. They split patients into separate groups for training and testing so that images from the same person never appeared in both sets, and used repeated cross‑validation to reduce random fluctuations. The three models were trained to distinguish seven growth patterns and related diagnostic groupings. Performance was measured using standard metrics that assess how well the models separate one subtype from the others. Across many runs, the models achieved high discrimination, especially for clearly defined patterns such as in situ and several invasive forms, showing that the dataset contains consistent and learnable visual signals. When the same methods were applied to an existing U.S. dataset from Dartmouth, CLWD often produced equal or better results, suggesting it is a strong benchmark and a valuable complement for cross‑country comparisons.
What This Means for Patients and Researchers
The CLWD collection offers an open, well‑curated set of lung cancer images from Chinese patients, bridging a gap in current resources that have largely been built on Western cohorts. By pairing rich clinical information with carefully checked slide labels, it gives researchers a solid foundation to develop and compare artificial‑intelligence systems for early detection and refined subtyping of lung adenocarcinoma. While the dataset has limitations—it comes from a single hospital, some subtypes are less common, and only standard staining is included—it still represents an important step toward more inclusive, data‑driven pathology. As future tools trained on CLWD and similar datasets mature, they could help pathologists spot high‑risk patterns more reliably, guide follow‑up care, and ultimately improve outcomes for people facing lung cancer.
Citation: Chen, Y., Zhao, H., Wang, L. et al. CLWD: a Chinese histopathology dataset for lung adenocarcinoma subtype classification. Sci Data 13, 599 (2026). https://doi.org/10.1038/s41597-026-06906-z
Keywords: lung adenocarcinoma, digital pathology, histopathology images, deep learning, cancer subtypes