Clear Sky Science · en
High-resolution, high-throughput detection of hidden antibiotic resistance with the dilution-and-delay (DnD) susceptibility assay
Why hidden germs matter for your health
Antibiotics are cornerstones of modern medicine, but they do not always work as expected. Even when lab tests say a bacterial strain is “susceptible,” treatment can still fail because a tiny fraction of cells quietly survives. This paper introduces a new lab method, called the dilution‑and‑delay (DnD) assay, that can reveal those rare survivors at levels as low as one in one hundred million cells. By exposing this hidden fraction, the method helps explain puzzling treatment failures and offers a more nuanced way to think about antibiotic resistance.
Beyond yes-or-no antibiotic tests
Hospitals currently rely on simple tests that ask a blunt question: does a bacterial population grow or not at a given antibiotic dose? The result is summarized as a single number, the minimum inhibitory concentration (MIC), and then turned into a yes-or-no label: susceptible or resistant. These tests are fast and practical but average over millions of cells, hiding differences within the population. In many infections, only a small minority of cells is tougher than the rest—able to tolerate or survive high drug levels through mechanisms such as heteroresistance (a resistant minority), persistence (dormant survivors), or adaptive resistance (drug‑induced hardening). Standard tests are largely blind to these rare cells, even though they can seed treatment failure and fuel the future spread of full-blown resistance.
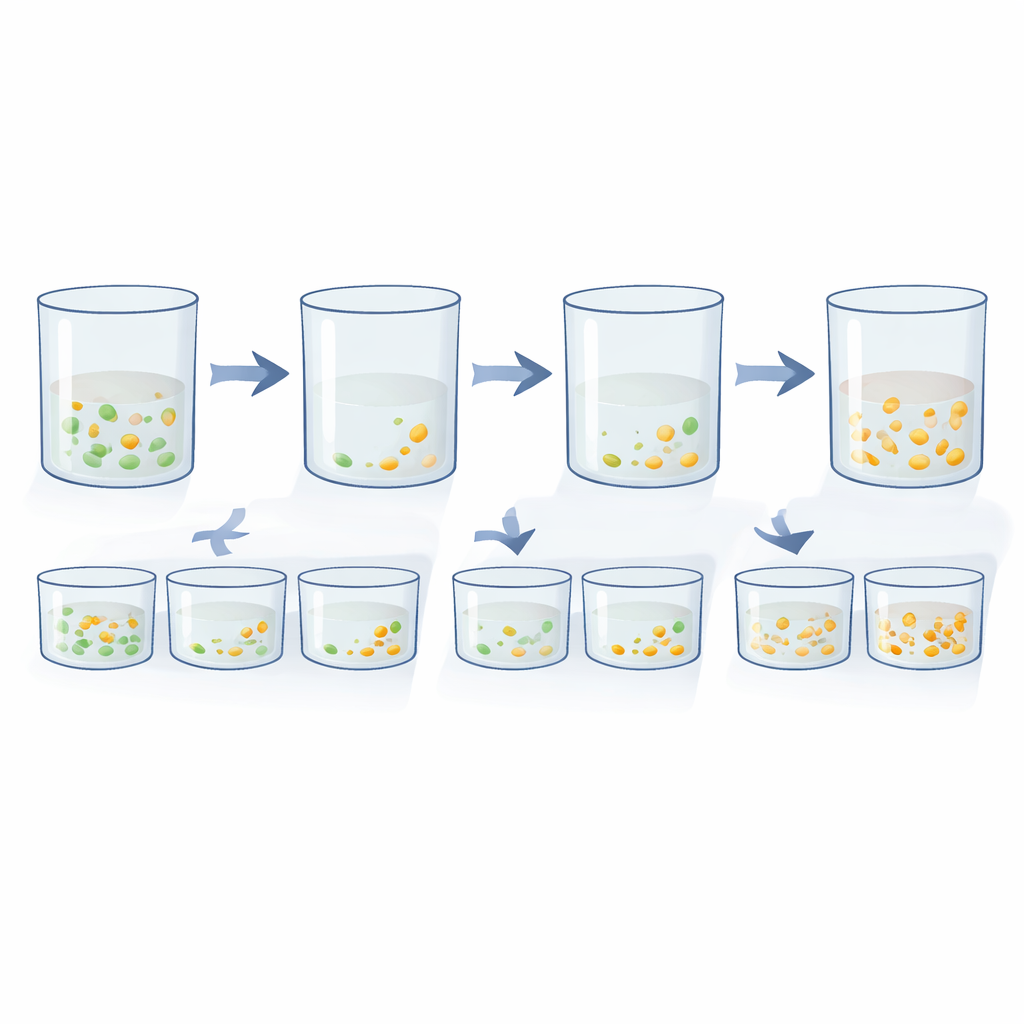
Two simple ideas: fewer cells and more time
The DnD assay builds on two intuitive principles of bacterial growth. The first is “dilution‑to‑extinction.” If you have a mixture of mostly sensitive cells plus very rare resistant ones, the chance that a test tube contains at least one resistant cell depends on how many cells you put in. By preparing a series of ten‑fold dilutions and checking which tubes become cloudy in the presence of antibiotics, the researchers can pinpoint the dilution where growth disappears. Comparing this extinction point with and without antibiotics reveals how many resistant cells were hiding in the original culture. The second principle is “delay‑to‑growth.” When a culture is exposed to an antibiotic, sensitive cells die off, and the total cloudiness drops. If a few resistant cells survive, they will eventually regrow—but only after a noticeable delay. By tracking cloudiness over time with a plate reader and mathematically rewinding the growth curve back to the starting point, the team can estimate how many survivors were present at the beginning.
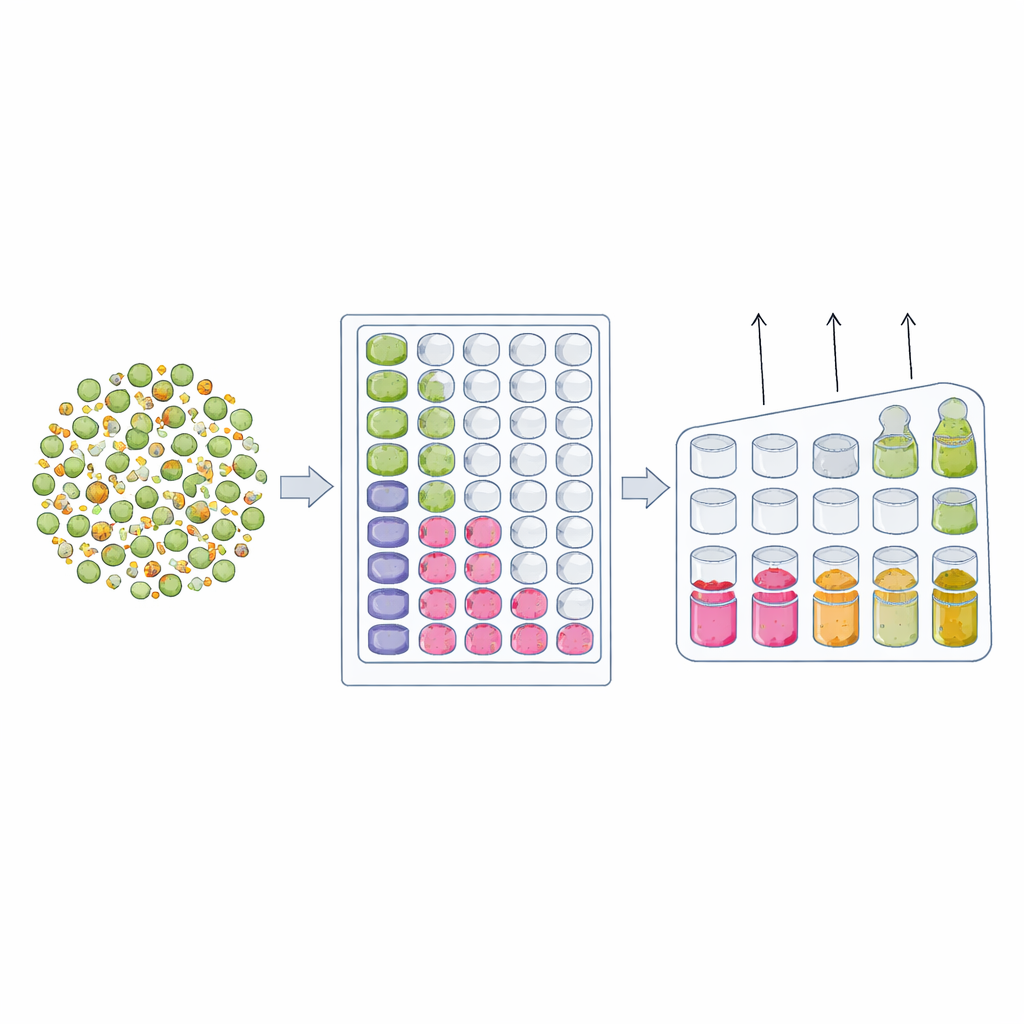
Combining signals to see rare survivors clearly
Because both dilution‑to‑extinction and delay‑to‑growth use the same basic setup—a broth culture in multi‑well plates—the authors combined them into one integrated DnD framework. Each plate contains a serial dilution of the bacterial sample, and a standard instrument automatically measures how cloudy each well becomes over many hours. Wells that never turn cloudy define where resistant cells run out, while wells that show late regrowth reveal how long rare survivors took to bounce back. By averaging information across the dilution series, the method provides a precise estimate of the frequency of antibiotic‑insensitive cells. Tests with controlled mixtures of fully sensitive and fully resistant strains showed that DnD faithfully recovered known fractions over many orders of magnitude, matching or surpassing older, labor‑intensive plate‑count methods.
Revealing a spectrum, not a switch
Once validated, the DnD assay was turned on real clinical isolates from several common hospital pathogens and five important antibiotics. The results were striking: nearly one in five strain–drug combinations fell between clearly susceptible and clearly resistant, forming a broad plateau of “in‑between” behavior. Many isolates also carried minority subpopulations that could persist after intense antibiotic exposure, with their frequencies spread across five orders of magnitude. These patterns showed that what we call “resistance” is not a sharp on/off state but a continuum of survival probabilities. The study further demonstrated that the widely used MIC threshold mainly reflects the detection limits of conventional assays: it is the point where resistant minorities become so rare that a standard‑sized inoculum is unlikely to contain even one survivor.
What this means for patients and medicine
For a non‑specialist, the key message is that infections can harbor a small but dangerous reserve army of bacteria that standard tests fail to see. The DnD assay offers a practical, scalable way to measure how many such hidden survivors exist, long before they take over and push a strain into obvious resistance. While more clinical studies are needed to tie these detailed measurements to patient outcomes, the work lays the groundwork for more informed antibiotic choices, better tracking of early resistance evolution, and updated guidelines that treat antibiotic susceptibility as a spectrum rather than a simple yes‑or‑no label.
Citation: Ma, M., Kim, M. High-resolution, high-throughput detection of hidden antibiotic resistance with the dilution-and-delay (DnD) susceptibility assay. Nat Commun 17, 3641 (2026). https://doi.org/10.1038/s41467-026-70174-z
Keywords: antibiotic resistance, heteroresistance, bacterial persistence, susceptibility testing, clinical microbiology