Clear Sky Science · en
Supercoiled DNA recognition and cleavage control in topoisomerase VI
Untangling Life’s Twisted Code
Inside every cell, DNA is not a neat straight ladder but a long, flexible thread that often twists into coils and loops. These twists are essential for packing meters of genetic material into microscopic spaces, yet they also create knots and tangles that can stall vital processes like copying and reading genes. This study reveals how one molecular machine, topoisomerase VI, specifically recognizes and safely cuts highly twisted DNA, offering a window into how cells manage the mechanical stress in their genetic code.
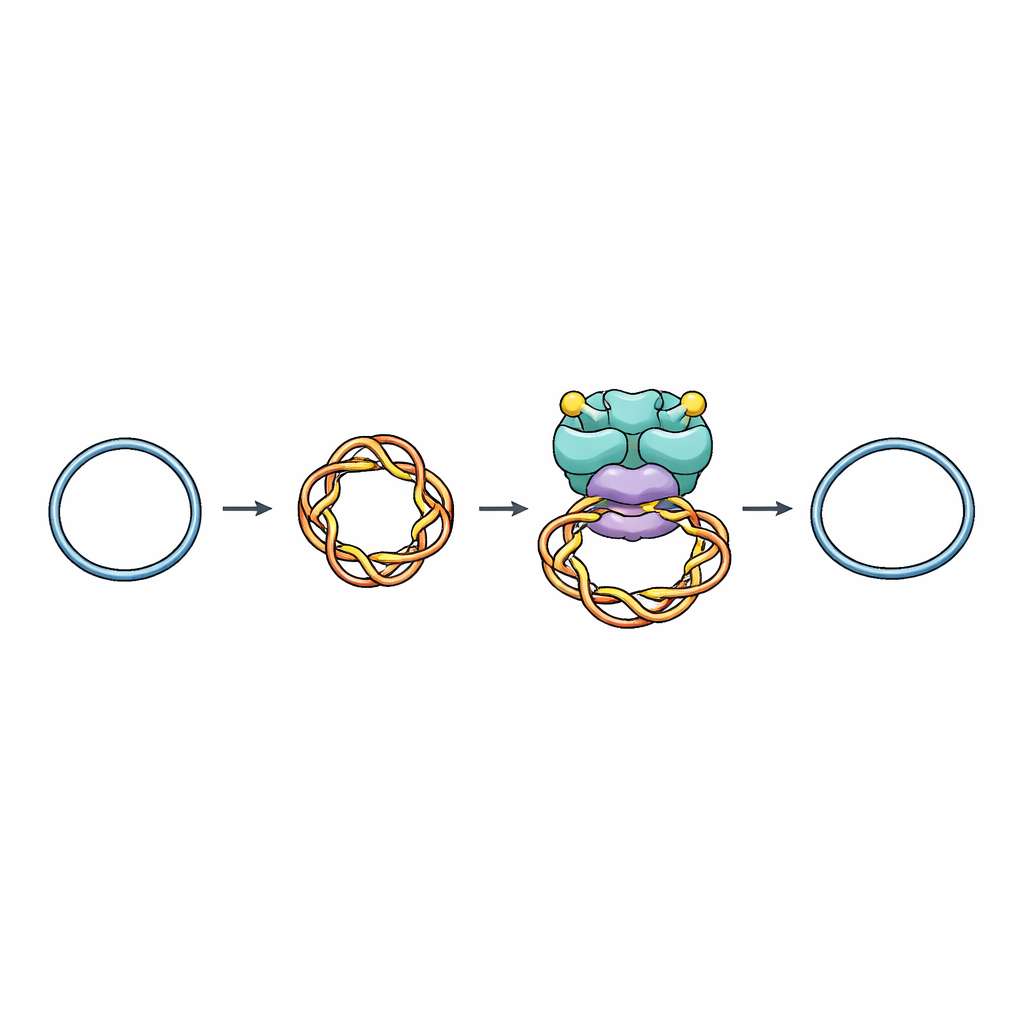
A Molecular Tool for Managing DNA Knots
Cells rely on enzymes called topoisomerases to add or remove twists in DNA, preventing dangerous snarls during replication, transcription, and chromosome separation. Topoisomerase VI (Top6), found in archaea and plants and related to proteins that start meiosis and defend bacteria from viruses, belongs to a family that cuts both strands of the DNA helix, passes another piece of DNA through the gap, and then reseals the break. Until now, scientists lacked a detailed picture of how Top6 recognizes the right piece of DNA to cut and how it avoids making accidental, harmful breaks.
Capturing DNA in the Act of Twisting
The researchers used single-particle cryogenic electron microscopy, a technique that images frozen molecules at near-atomic resolution, to visualize Top6 bound to tiny circular pieces of DNA that were purposely made supercoiled—like an overwound rubber band. They solved structures of Top6 in both uncut and cut states while the enzyme held these coils. In all cases, Top6 clasped a 74-base-pair stretch of DNA bent into a tight loop resembling the tip of a supercoiled plectoneme, the kind of sharp bend that naturally appears in overwound DNA. This explains why Top6 prefers supercoiled DNA over relaxed, straight pieces: its binding pocket is shaped to fit a strongly curved segment.
How the Enzyme Feels and Chooses the Right Spot
Closer inspection showed that Top6 does not simply grab any bent DNA. It favors a particular region whose underlying sequence makes it especially easy to deform. By analyzing the density maps, the team inferred which base pairs were present at the cut site and then calculated how flexible different stretches of the circular DNA should be. The cleavage site sits at a boundary where a comparatively rigid segment meets a very flexible one, and the exact base steps where the cuts occur are themselves unusually bendable. Around this spot, the DNA is forced into a more compressed, slightly non-standard shape. The enzyme appears to use this combination of local flexibility and sharp bending as a physical signature, allowing it to home in on safe cutting sites without “reading” the genetic code in the usual letter-by-letter way.
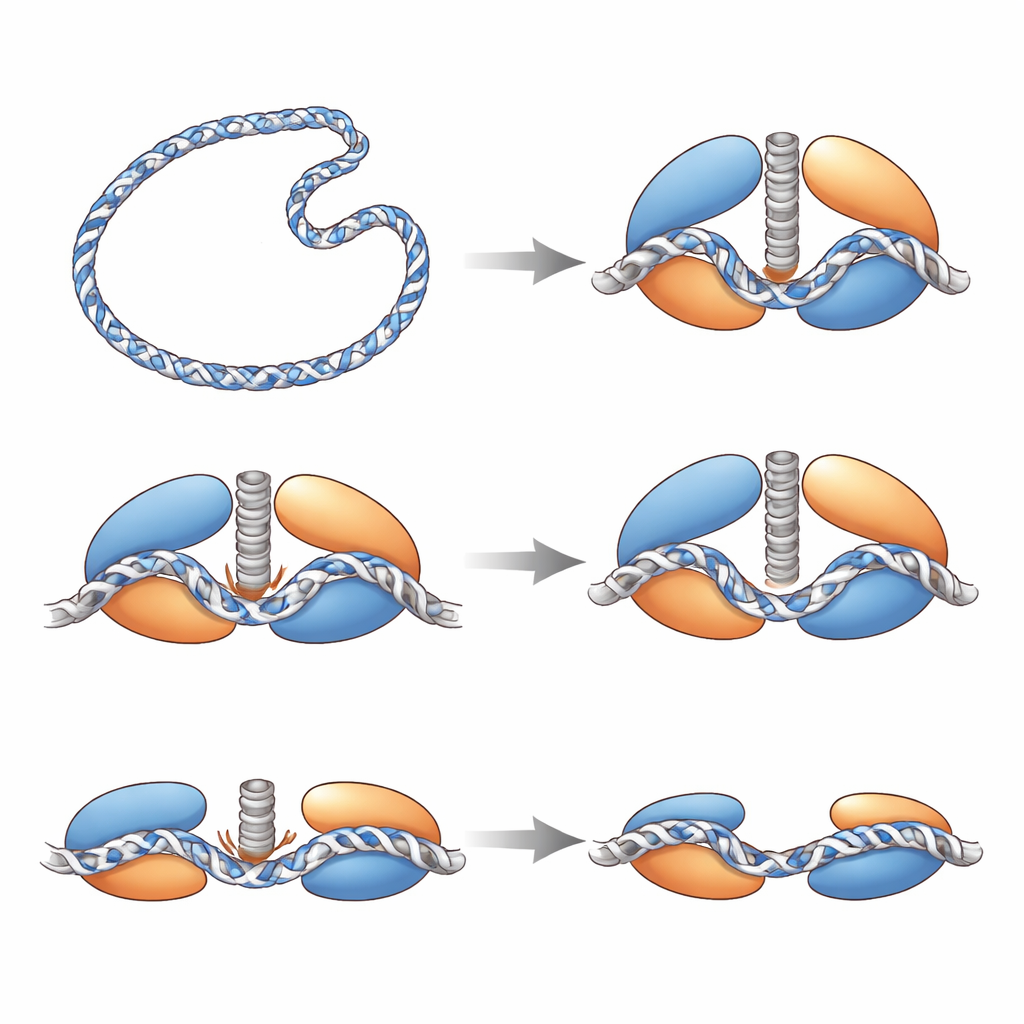
A Safety Latch and a Tension Sensor
The structures also reveal internal safeguards that keep Top6 from cutting DNA too readily. Within the DNA-processing half of the enzyme, a flexible loop becomes ordered only when DNA is tightly bent in place; it acts like a latch, bracing the active site against the DNA. A pair of oppositely charged amino acids forms an “electrostatic clasp” that physically restrains the catalytic tyrosine residue from reaching the DNA backbone. When the team weakened this clasp, the enzyme produced many more breaks, showing that the clasp normally suppresses cleavage. In the ATP-powered half of Top6, a long helical stalk that links motion in the energy-harvesting region to the DNA-cutting region behaves like a tension sensor: when intact and straight, it allows a tight, cleavage-ready configuration; when partially unfolded or destabilized by mutation, the enzyme struggles to relax supercoils, indicating that controlled flexibility in this stalk coordinates energy use with safe cutting.
DNA as an Active Mechanical Partner
Putting these observations together, the study portrays Top6 and DNA as partners in a mechanical dance. Supercoiling stores energy in the DNA, driving certain sequences to bend sharply. Top6 recognizes these stressed shapes, clamps around them, and—only when its internal latch, clasp, and stalk are correctly positioned—makes a carefully controlled double-strand break before resealing it. This work shows that DNA’s physical properties, not just its sequence of letters, help direct where and when cuts occur. By clarifying how Top6 senses and responds to supercoiled DNA, the findings deepen our understanding of chromosome maintenance, meiotic recombination, and some bacterial defense systems, and they highlight how life exploits the physics of twisted molecules to keep the genome both compact and intact.
Citation: Richman, D.E., Wendorff, T.J., Rashid, F. et al. Supercoiled DNA recognition and cleavage control in topoisomerase VI. Nat Commun 17, 3092 (2026). https://doi.org/10.1038/s41467-026-69491-0
Keywords: DNA supercoiling, topoisomerase VI, DNA mechanics, cryo-electron microscopy, genome stability