Clear Sky Science · en
A convolutional attention model classifies copy number variants from whole exome sequencing
Finding Hidden Clues in Our DNA
Doctors increasingly use DNA sequencing to look for genetic changes that can explain disease, but some of the most important clues are not single “typo” mutations—they are chunks of DNA that are missing or duplicated. These changes, called copy number variants, can be hard to spot in the data most hospitals already generate. This study introduces a new computer model that reads noisy DNA coverage patterns and spots these missing or extra pieces more accurately and consistently across different sequencing machines, potentially sharpening a tool that is already common in medical genetics.
Why Extra or Missing DNA Matters
Copy number variants are stretches of DNA that occur in fewer or greater copies than usual. A segment might be deleted entirely or copied several times. Such changes can shape everyday traits, influence the risk of conditions like cancer or neurodevelopmental disorders, and affect how human populations evolve over time. Clinically, finding these variants is crucial in both rare disease diagnosis and tumor profiling. Many patients already undergo whole-exome sequencing, which focuses on the parts of the genome that code for proteins. Leveraging these existing exome tests to also detect copy number variants could make genetic workups more informative without requiring additional, more expensive assays.
Why Current Tools Struggle
Detecting copy number variants from exome data is technically difficult. The exome capture process samples the genome unevenly, leading to jagged, noisy read depth—how many sequencing reads cover each region. Traditional software tools smooth this noise using statistical tricks and hand‑crafted rules, then apply thresholds to decide whether a region is normal, deleted, or duplicated. While useful, these methods often falter when coverage is low, when sequencing is done on different machines or chemistries, or when subtle patterns across neighboring regions and chromosomes matter. As a result, sensitivity can suffer, especially for smaller or noisier events, and performance may not transfer well between laboratories or platforms.

A New Way to Read Noisy Signals
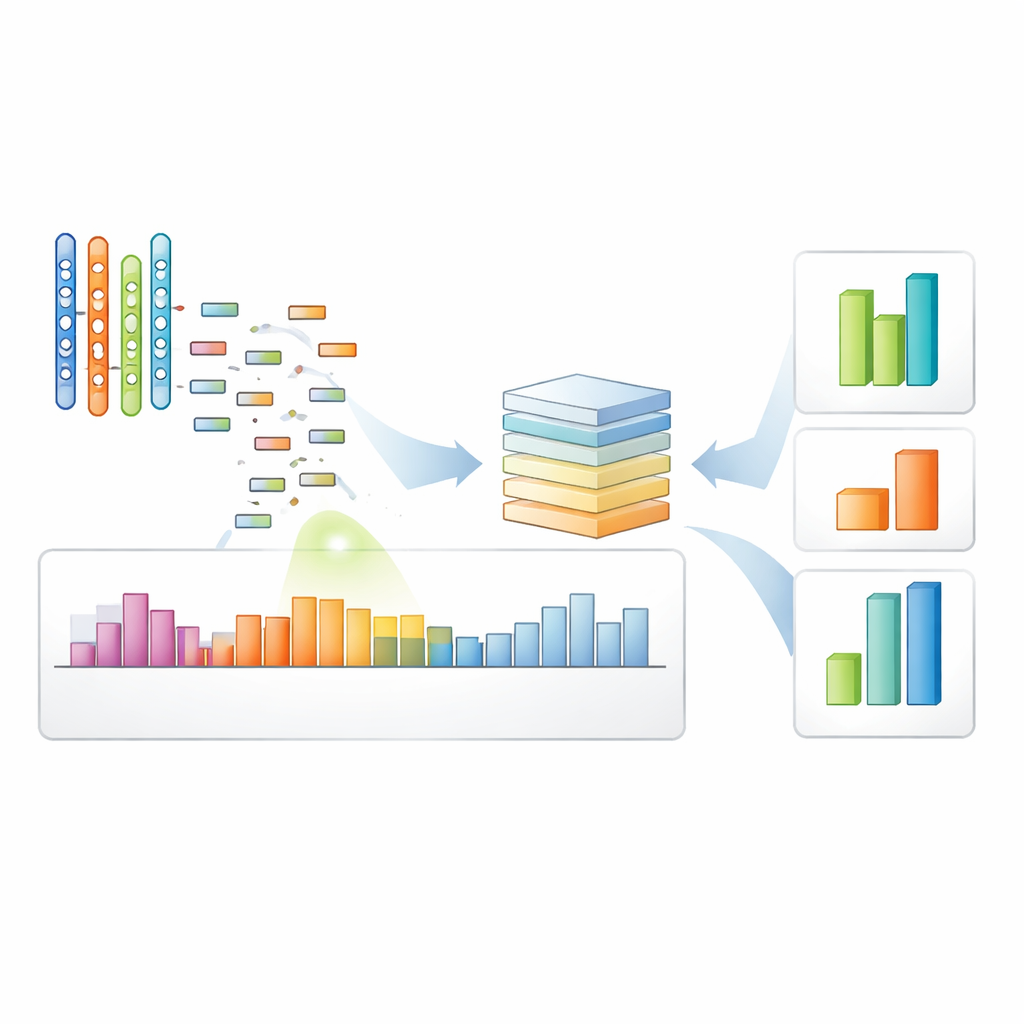
The authors designed a deep learning model, called CNN‑Att, that learns directly from the raw coverage patterns instead of relying mainly on fixed rules. For each protein‑coding segment (an exon), the model takes in a standardized snapshot of read depth across the exon and its surrounding region, along with its genomic start and end positions. It also receives an encoded tag indicating which chromosome the exon comes from. Convolutional layers—originally popularized for image analysis—scan along this one‑dimensional signal to capture local shapes in the coverage pattern, such as dips that might indicate deletions or subtle bumps suggesting duplications. An attention mechanism then highlights the most informative features, particularly faint signals that could correspond to small or noisy events, before the model makes a three‑way decision: normal, deletion, or duplication.
How Well the Model Performs
To evaluate CNN‑Att, the researchers trained it on a large benchmark built from the 1000 Genomes Project, where exome data are paired with labels inferred from more comprehensive whole‑genome sequencing. On a separate set of 50 exome samples held out for testing, the model correctly classified about 83 percent of exon windows overall and showed strong ability to distinguish between the three classes, with high scores on both receiver‑operating and precision–recall curves. Deletions were somewhat easier to detect than duplications, reflecting the fact that deletions usually leave a stronger footprint in coverage. The model outperformed a simpler baseline that knew only the genomic coordinates, indicating it was truly learning from the depth patterns rather than memorizing “hotspot” locations where variants are common.
Reliable Across Different Sequencers
Because clinical and research centers use a variety of sequencing machines, a practical tool must behave well across platforms. The authors therefore tested CNN‑Att on exome data from the same reference DNA sample sequenced on four major technologies: HiSeq 4000, NovaSeq 6000, MGISEQ 2000, and BGISEQ 500. Across these diverse instruments, the model’s overall F1‑score—a balance of precision and recall—ranged from 0.89 to 0.96, consistently higher than several widely used traditional tools. In a further experiment, the team fine‑tuned only the final decision layers of the model using a small set of seven samples painstakingly labeled by experts. Even with this limited curated data, fine‑tuning noticeably boosted recall for true deletions and duplications on held‑out samples, at the cost of some extra false positives, a trade‑off often acceptable when questionable calls can be checked with follow‑up tests.
What This Means for Patients and Research
This work shows that a focused deep learning approach can turn the noisy, uneven coverage of routine exome sequencing into a more reliable detector of missing and extra DNA segments. CNN‑Att attains high sensitivity while keeping errors at manageable levels and remains robust across different sequencing machines, making it useful for multi‑site studies and large population projects. Although it still needs validation on larger expert‑annotated cohorts and currently depends on a specific reference genome, the framework points toward exome tests that miss fewer important variants. In practice, that could mean more patients receiving timely, actionable genetic answers from the sequencing they are already getting.
Citation: Ouhmouk, M., Abik, M. A convolutional attention model classifies copy number variants from whole exome sequencing. Sci Rep 16, 14310 (2026). https://doi.org/10.1038/s41598-026-44691-2
Keywords: copy number variants, whole exome sequencing, deep learning genomics, convolutional neural network, clinical genetics