Clear Sky Science · en
The expression pattern of EXO70 subunits in single-cell stereo-seq of Arabidopsis thaliana leaves using coexistence network analysis
How plant leaves quietly manage traffic
Inside every plant leaf, countless tiny cargo packets shuttle fats, proteins, and signaling molecules to the cell surface. This microscopic traffic keeps leaves growing, sensing light, and coping with stress. The study described here looks at one set of traffic directors, the EXO70 family of proteins in Arabidopsis thaliana, and shows how different versions of these proteins are switched on in different leaf cell types. It also introduces a new way to find patterns in very faint gene activity, opening doors to studying many other hard-to-detect molecules.
Why tiny traffic directors matter
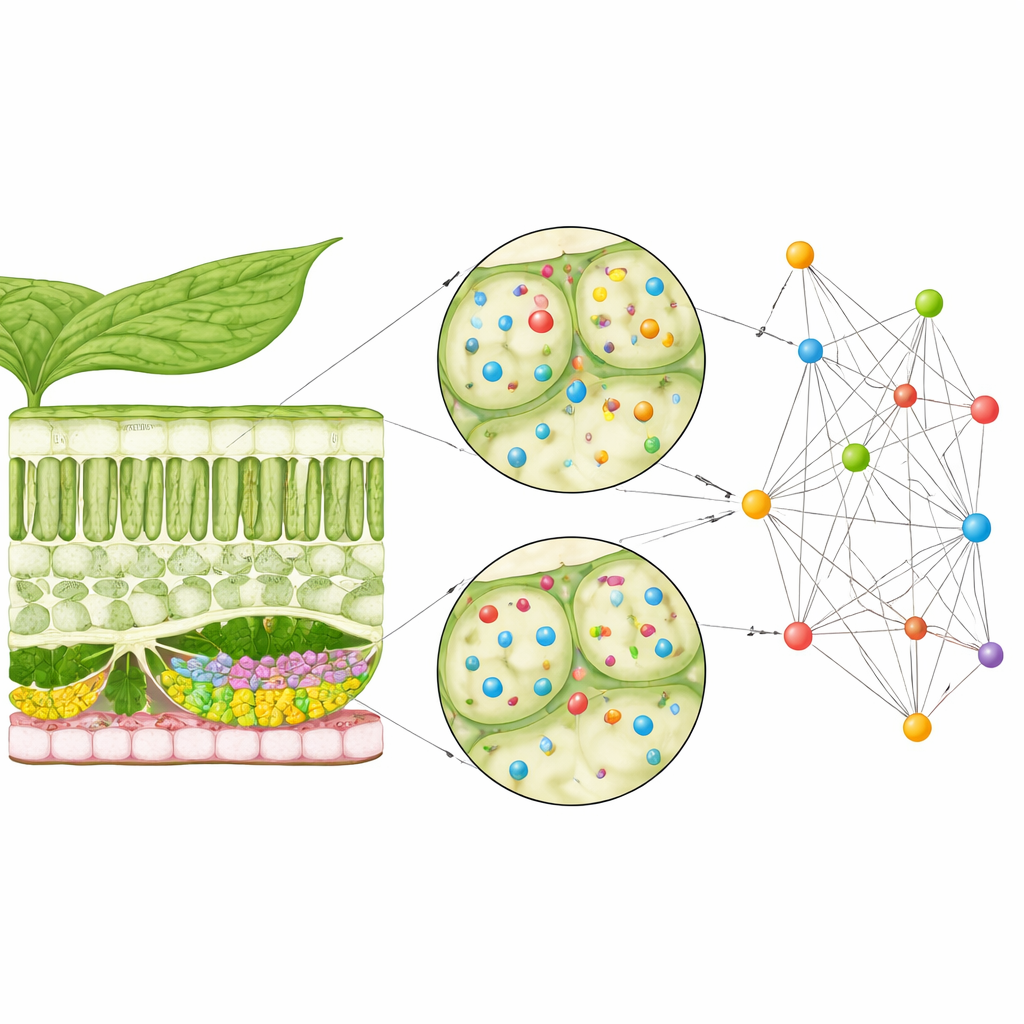
Plant cells rely on a delivery system that sends membrane-bound packets, or vesicles, to specific spots on the cell surface. A large protein assembly called the exocyst helps dock these vesicles. One group of exocyst components, known as EXO70 proteins, exists in many slightly different versions, or isoforms, in Arabidopsis leaves. Earlier work had hinted that different EXO70 isoforms work in different cell types and situations, but it was not clear how their activity varied across the layered structure of the leaf. Because the genes that encode these proteins are often weakly active, they are difficult to study with standard tools that look for strong gene signals.
Looking at individual cells in place
The researchers turned to a powerful approach called single-cell spatial transcriptomics, which reads out which genes are active in hundreds of individual cells while preserving their positions within a leaf cross-section. They reused public “stereo-seq” data from Arabidopsis leaves that captured upper and lower skin layers, two kinds of photosynthetic cells, and vascular cells that carry water and nutrients. Even with this high-resolution method, each cell showed activity for only a small fraction of all genes, and EXO70-related genes were especially faint. Traditional “co-expression” analyses, which rely on comparing how strongly genes turn on together, therefore struggled to find meaningful patterns in these sparse signals.
From co-expression to coexistence
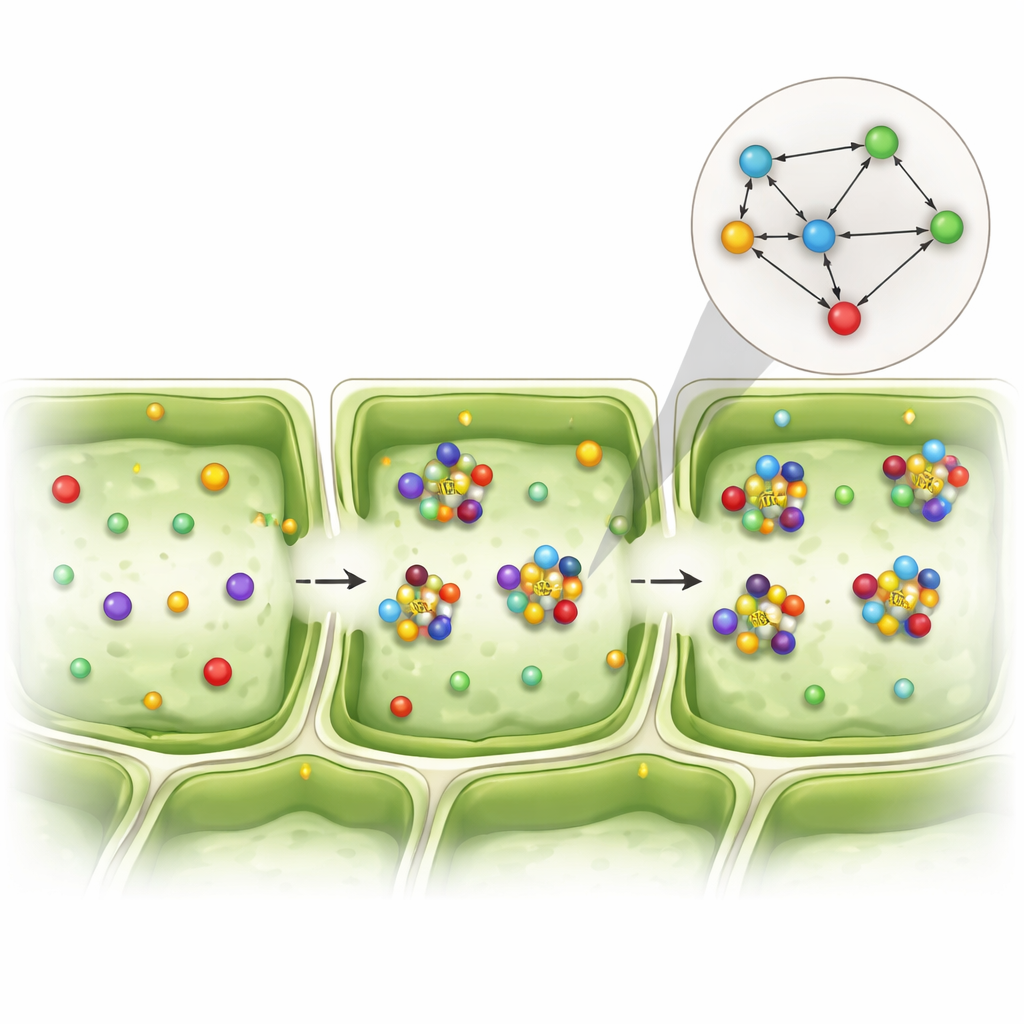
To overcome this hurdle, the team reframed the problem. Instead of asking whether two genes were strongly active together, they asked a simpler yes-or-no question: do these two genes show up in the same cell at all? They converted the data into a grid of zeros and ones that marked mere presence or absence, then multiplied and compared these patterns across thousands of cells. This produced what they call a “coexistence network,” a map of how often pairs of genes tend to be found in the same cells more often than expected by chance. Using statistical shuffling tests, they highlighted gene pairs whose shared appearance was unlikely to be random, even when each gene was only weakly active.
Hidden partnerships inside leaf tissues
This coexistence view revealed distinct interaction patterns among EXO70 isoforms, other exocyst components, and vesicle-related proteins in different cell layers. In particular, certain EXO70 versions and another exocyst subunit, SEC3A, repeatedly appeared together in spongy photosynthetic cells, suggesting that these proteins act as a team to guide vesicle delivery in that specific tissue. Other EXO70 combinations stood out in the upper surface cells of the leaf, hinting at roles in stress responses and cell wall shaping. While the study focused on gene activity rather than protein levels, and many EXO70 proteins are known to be controlled after their RNA is made, these recurring coexistence patterns point to functionally specialized partnerships within the same molecular family.
A new lens on quiet but crucial genes
For non-specialists, the key message is that important biological players are not always the loudest ones. By switching from measuring how “strongly” genes turn on together to simply asking whether they tend to appear in the same cell, this work recovers subtle but meaningful connections among low-activity genes. The authors show that EXO70 proteins in leaves are not interchangeable parts, but members of tailored teams assembled in specific cell types. Their coexistence network approach offers a general strategy for uncovering similarly quiet but essential regulators, from transcription factors to signaling proteins, in many kinds of tissues.
Citation: Yu, X., Shang, J. The expression pattern of EXO70 subunits in single-cell stereo-seq of Arabidopsis thaliana leaves using coexistence network analysis. Sci Rep 16, 10694 (2026). https://doi.org/10.1038/s41598-026-44501-9
Keywords: exocyst, spatial transcriptomics, Arabidopsis, vesicle trafficking, single-cell biology