Clear Sky Science · en
Genetically engineered Streptomyces viridosporus ATCC 14672 strains for the discovery of novel moenomycins
Why this work matters
Antibiotic resistance is rising so fast that many once-routine infections are becoming difficult to treat. The study described here explores how to tweak a powerful but imperfect antibiotic, moenomycin, by re-engineering the soil bacterium that makes it. By changing the bacterium’s genes, the researchers create new versions of moenomycin that may ultimately lead to better drugs for fighting stubborn, hospital-acquired infections.
A powerful but problematic antibiotic
Moenomycin is a natural compound made by the bacterium Streptomyces viridosporus. It blocks a key step in building the bacterial cell wall and is the only known drug that hits this target directly at extremely low doses. It has been used safely in animals for decades without widespread resistance, making it very attractive as a starting point for new human medicines. However, moenomycin has two major drawbacks in our bodies: it is not absorbed when taken by mouth, and it lingers in the bloodstream for a very long time. Both issues are linked to its unusually long greasy “tail” made of 25 carbon atoms.
Redesigning the bacterial factory
The team focused on the genes that build this tail in Streptomyces viridosporus strain ATCC 14672, the best-studied producer of moenomycin. Two genes, called moeO5 and moeN5, carry out early steps that attach and then extend the lipid tail. The researchers used modern genetic tools to delete each gene separately, creating two new bacterial strains. One mutant, named dO5, completely lost the ability to make any moenomycins. The other, called M12, still made related compounds but with a shorter 15-carbon tail instead of the original 25-carbon version.
Turning mutants into discovery tools
The dO5 strain, which cannot start building the tail on its own, became a clean test bed for trying out replacement genes from other microbes. When the scientists introduced similar genes from two other antibiotic-producing bacteria, moenomycin production came back, showing that these enzymes can fill in for the missing step. But a more distant relative from an insect-associated bacterium did not restore activity, hinting that it may act on different starting materials or be shaped differently. Computer-based structure models and evolutionary analyses supported this idea, grouping this enzyme into a separate family. Together, these experiments show that dO5 can be used as a living sensor to tell whether new enzymes from genome databases are able to launch moenomycin-like chemistry.
New molecules with shorter tails
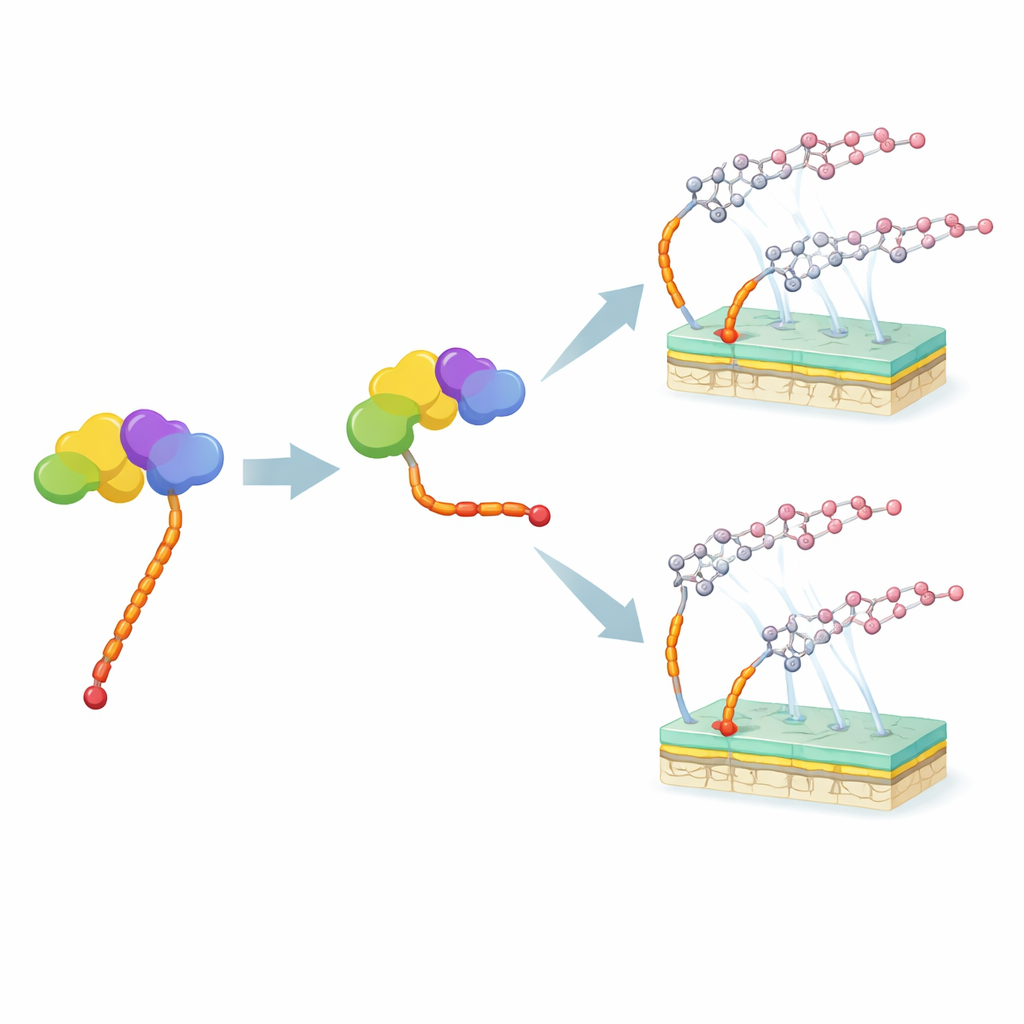
The M12 strain, lacking the moeN5 gene, offered a different advantage: it naturally accumulated new moenomycin variants with a shorter tail. Using advanced mass spectrometry, the researchers identified two such compounds, closely related to known members of the moenomycin family but bearing the 15-carbon tail. They purified these molecules and compared their ability to stop the growth of the pathogen Staphylococcus aureus with that of the original moenomycin. The shorter-tail versions were far less potent—up to a hundredfold weaker—though certain other features on the sugar portion of the molecule could partially restore activity.
What this means for future antibiotics
This work shows that simply shortening moenomycin’s tail, while attractive for improving how the drug behaves in the body, comes at a steep price in antibacterial strength. At the same time, the study delivers two valuable genetic tools: one strain that can host and test tail-building genes from many sources, and another that reliably produces new, shorter-tail molecules for detailed study. Together, these engineered bacteria form a platform for exploring a wide range of moenomycin-like compounds. Over time, this approach may help chemists and microbiologists balance potency with safer, more manageable drug properties, nudging moenomycin-inspired antibiotics closer to use in the clinic.
Citation: Ostash, B., Makitrynskyy, R., Fedchyshyn, M. et al. Genetically engineered Streptomyces viridosporus ATCC 14672 strains for the discovery of novel moenomycins. Sci Rep 16, 12851 (2026). https://doi.org/10.1038/s41598-026-43988-6
Keywords: antibiotic resistance, moenomycin, Streptomyces, genetic engineering, natural products