Clear Sky Science · en
Automated detection of fetal vascular malperfusion via data augmentation and algorithm improvement
Why tiny clues in the placenta matter
Before and after birth, the placenta quietly sustains a baby’s life by steering blood, oxygen, and nutrients between mother and fetus. When that flow is disturbed, as in a condition called fetal vascular malperfusion, the risks of poor growth, preterm birth, or even stillbirth rise. Yet today, spotting the telltale damage in placental tissue still relies on experts visually scanning microscope slides, a process that is time‑consuming and can vary from one pathologist to another. This study explores how modern artificial intelligence can help read these subtle tissue patterns more consistently, offering a step toward faster and more reliable pregnancy care.
Hidden patterns in a baby’s lifeline
Fetal vascular malperfusion arises when blood flow from the fetus through the placenta is chronically obstructed, often by problems with the umbilical cord, clots, or high blood pressure. Under the microscope, this shows up as a constellation of small lesions—blocked vessels, damaged or empty villi, and tiny areas of clot or scarring. Recognizing these patterns after delivery can warn doctors about a newborn’s long‑term health risks and guide closer monitoring in future pregnancies. But the lesions vary greatly in size and appearance, and they may be scattered across large placental sections. In busy hospitals or places with few specialists, that makes consistent diagnosis difficult and increases the chance that important warning signs are missed.
Teaching computers to read tissue images
The researchers set out to build an automated system that could detect these lesions from digital images of stained placental tissue. They began with samples from 310 pregnancies complicated by gestational diabetes, creating nearly two thousand cropped images that captured seven common lesion types linked to fetal vascular malperfusion. Because powerful deep‑learning models usually demand far more data than medicine can easily provide, the team turned to a medical imaging framework called MONAI. This software generates realistic variations of existing images—subtle flips, stretches, blurs, sharpening, and changes in color and contrast that mimic real‑world differences in slide preparation and scanning—while preserving the key diagnostic structures. Doubling the training data in this way gave the model a richer “experience” of how the same lesion might appear under slightly different conditions.
How the smarter vision system works
On top of this expanded image library, the team improved a fast object‑detection network known as YOLOv11. Standard versions of such networks can struggle with the tiny, fine‑edged abnormalities common in placental disease, especially when each full image covers a large area. The authors added a LocalWindow attention module, which conceptually slices the image features into many small patches and lets the model focus its computing power inside each patch. Within these windows, a series of attention steps emphasizes the most informative shapes and textures—such as subtle changes in vessel walls or villous borders—while downplaying uniform background. This design helps the system home in on small lesions without being overwhelmed by the surrounding normal tissue, much as a pathologist mentally narrows attention to a suspicious corner of a slide.
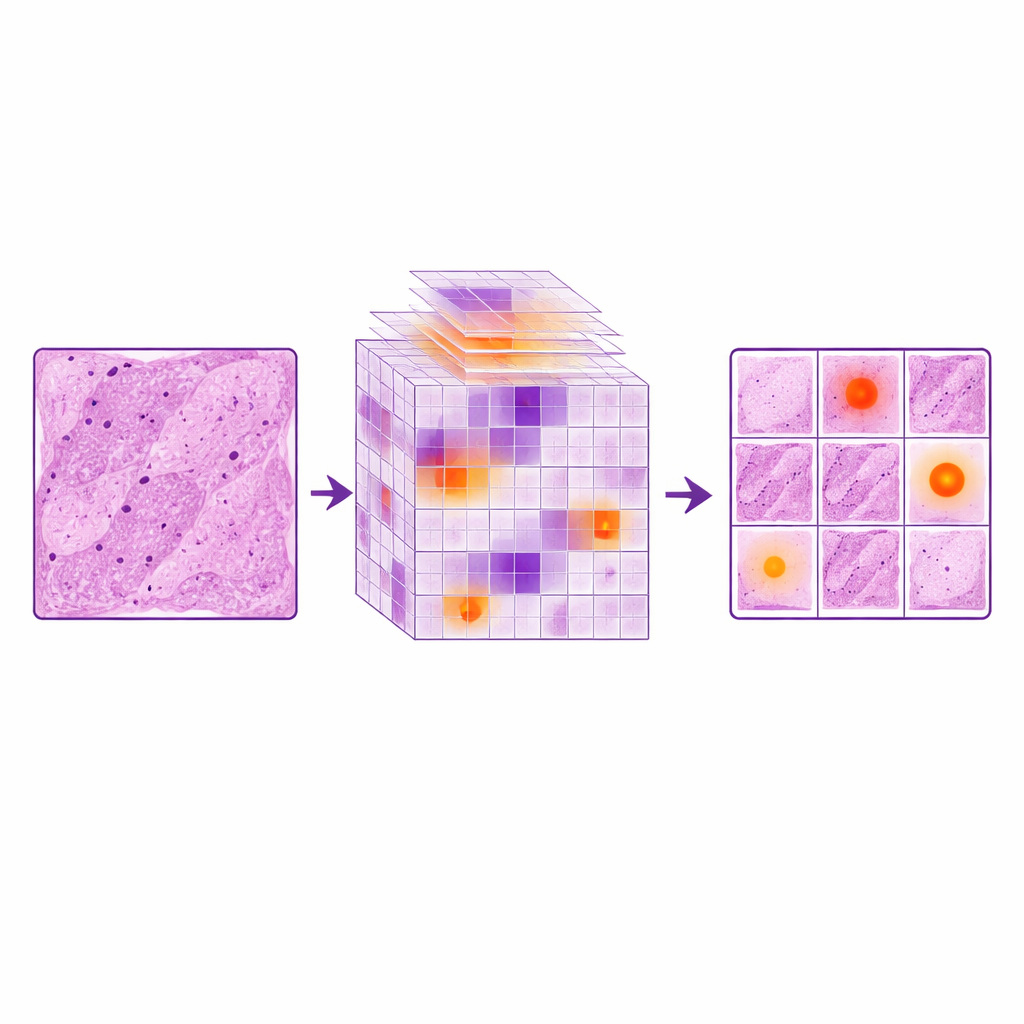
Measuring gains in accuracy and reliability
To see what each ingredient contributed, the researchers compared three setups: the original YOLOv11 model, the same model trained on simple duplicated images, and the enhanced versions using MONAI and LocalWindow attention alone or together. Just repeating images actually hurt performance, suggesting the model began to memorize the limited examples instead of learning general rules. In contrast, MONAI‑based augmentation reduced missed lesions dramatically and improved several accuracy scores. The LocalWindow attention module further boosted the system’s ability to detect lesions of different sizes. When both strategies were combined, the model achieved its best results, with key benchmarks rising by roughly 6–10 percent over the baseline and surpassing a range of other popular detection methods. Its overall performance approached that of an intermediate‑level pathologist, while still running fast enough for practical use.
What this means for mothers and babies
By pairing realistic data expansion with a more focused way of “looking” at tissue, the study shows that computers can learn to flag placental blood‑flow lesions with growing confidence. The authors argue that such tools will not replace human experts, but can act as tireless assistants—pre‑screening slides, highlighting suspicious regions, and helping standardize reporting across hospitals. This could be especially valuable in centers with few trained pathologists, where subtle placental problems are easily overlooked. Although the current work was done at a single hospital and relies on a modest dataset, it lays out a pathway for broader, multi‑center systems that support more consistent, data‑driven care for pregnancies at risk.
Citation: Li, X., Jiang, Z., Chen, F. et al. Automated detection of fetal vascular malperfusion via data augmentation and algorithm improvement. Sci Rep 16, 11042 (2026). https://doi.org/10.1038/s41598-026-39942-1
Keywords: placenta, fetal vascular malperfusion, computational pathology, deep learning, pregnancy outcomes