Clear Sky Science · en
A preclinical CT and MRI Liver Imaging Dataset with Anatomical, Functional and Segmentation Data
Why this matters for liver disease and medical imaging
Chronic liver diseases such as fatty liver, fibrosis, and liver cancer are rising worldwide, yet much of the detailed research data that could speed up new diagnostics and treatments is locked away in individual labs. This article presents a rare resource: a large, openly shared collection of CT and MRI scans from mice with different liver diseases, along with precise outlines of organs and rich metadata. By making these data freely available, the authors aim to accelerate imaging research, improve computer tools for reading scans, and ultimately support less invasive ways to monitor liver health.
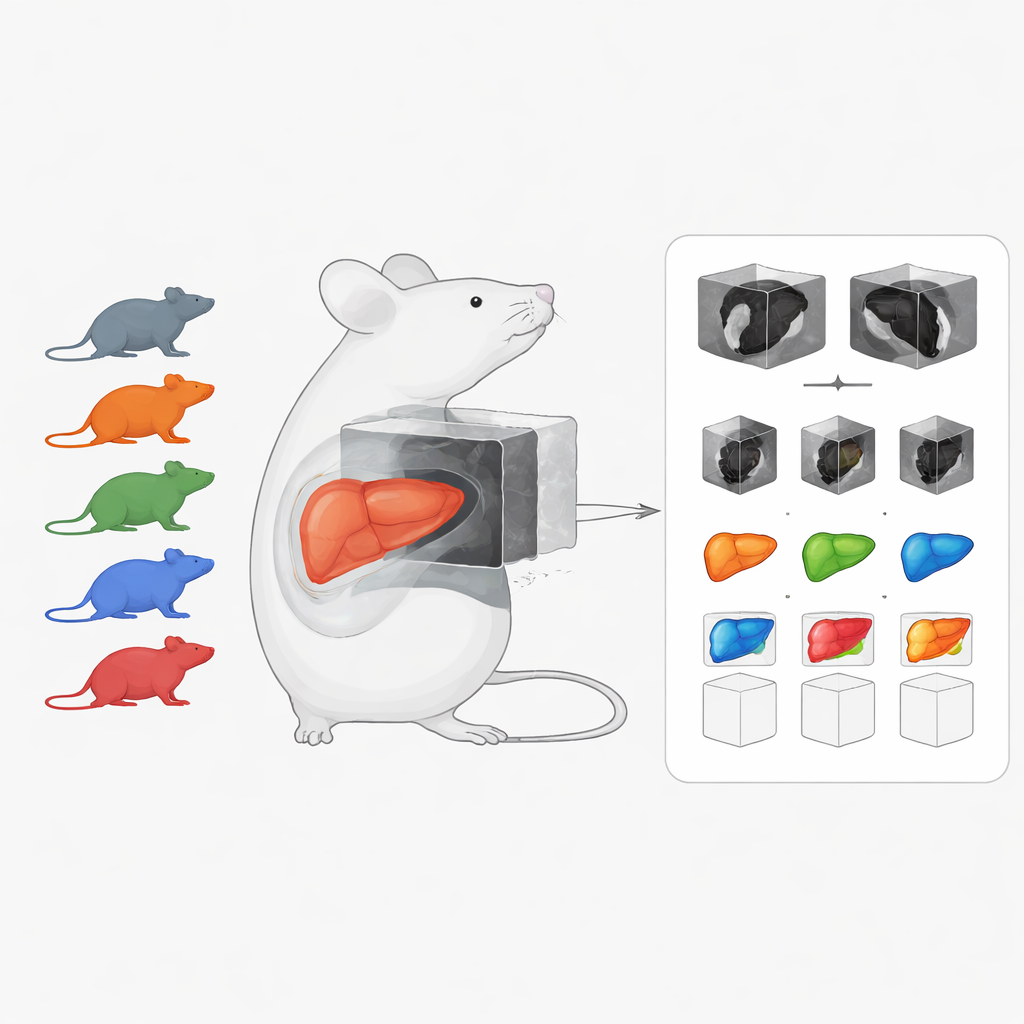
A shared picture library of diseased and healthy livers
The heart of the work is a curated preclinical imaging dataset focused on chronic liver disease in mice. It combines 222 three-dimensional CT and MRI scans covering several common conditions: liver cancer (hepatocellular carcinoma), fatty and inflamed liver (metabolic dysfunction–associated steatohepatitis, or MASH), and liver fibrosis, plus scans from healthy animals. Unlike many existing mouse imaging resources that target healthy organs or single imaging methods, this collection emphasizes diseased livers and brings together different scan types in one place. Many scans are accompanied by hand-drawn organ outlines, known as segmentations, which are crucial for training and testing computer algorithms.
Multiple scan types to capture structure and function
The dataset is built from five separate animal studies that were pooled to cover a broad range of liver damage and imaging approaches. In one study, mice developed liver cancer in a chronically injured liver, and high-field MRI was used repeatedly over weeks to follow how tumors appeared and grew. Another study used “multiparametric” MRI, in which each mouse underwent several kinds of scans: some highlighting anatomy, others measuring how water diffuses through tissue, and still others tracking how injected contrast agents move through the liver. These different measurements report on key features such as scarring, fat content, liver cell function, tissue damage, and immune cell activity.
CT scans to map body fat, liver size, and genetics
To complement MRI, the authors added several CT-based studies. In one, mice were fed a Western-style high-fat diet, with or without oat beta-glucan, a dietary fiber thought to protect the liver. Whole-body CT scans were used to measure liver volume and fat distribution, and the livers were manually outlined for precise volume estimates. A second CT study examined mice lacking a functional ICAM-1 gene, which is involved in inflammation, to see how this genetic change influenced fat distribution and liver size when the animals were fed a high-fat diet. In both CT studies, body fat, liver, and other structures were segmented so that researchers can directly analyze how diet and genes shape organ size and body composition.
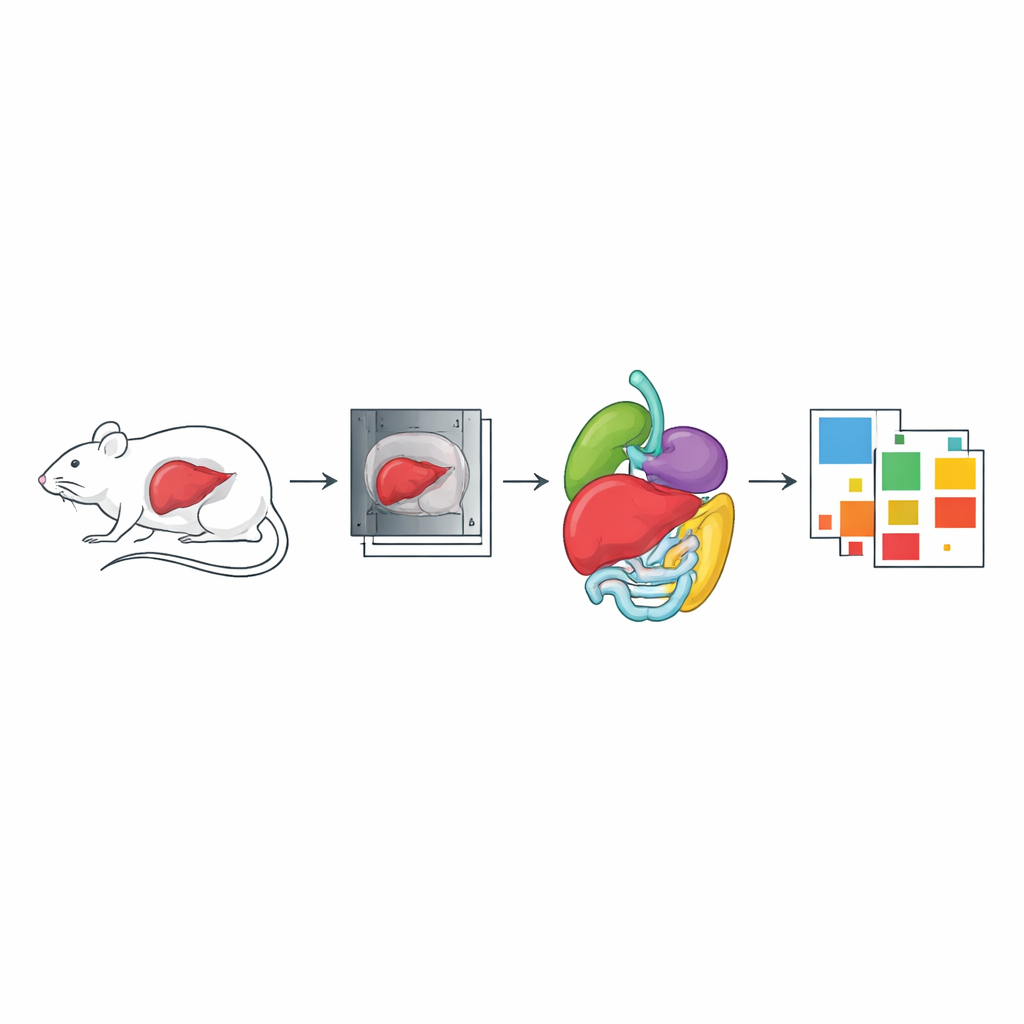
Bridging CT and MRI in the same animals
Another component of the dataset focuses on healthy nude mice scanned with both CT and MRI in the same imaging holder. Here, the goal was to create overlapping views of the same anatomy so that images from the two technologies can be precisely aligned. The animals were scanned with whole-body CT and then with a special MRI sequence that separates water and fat signals. Many organs—including liver, lungs, heart, kidneys, spleen, stomach, intestines, brain, and spinal cord—were segmented in either CT, MRI, or both. Such paired scans are valuable for tasks like building attenuation maps for PET-MR scanners and for testing image-registration methods that align different imaging modalities.
Rich metadata to support reuse and reproducibility
Beyond the images themselves, the authors invested heavily in standardized metadata so that others can understand and reuse the data. For each animal and scan, they record information such as mouse strain, age, disease model, imaging modality, device brand, contrast agents, and exact scan settings like field of view, voxel size, and timing. Organ segmentations are linked to clear definitions of what each label represents (for example, liver, visceral fat, bladder, or spine). All of this information is stored in ISA-Tab files following a tailored metadata profile, and the entire package is hosted on Zenodo as open data. The scans are provided in standard medical formats (DICOM or NIfTI) with segmentations in widely used segmentation file types.
How this resource can speed up future discoveries
To a lay reader, the main takeaway is that this work does not present a new drug or diagnostic test, but rather builds the foundation for many such advances. By releasing a large, well-annotated library of liver CT and MRI scans in mice, the authors make it easier for researchers to train computer models that automatically outline organs, detect tumors, or predict how liver disease will progress—without subjecting new animals to every experiment. The dataset can serve as a reference for imaging studies, reduce redundant animal use, and support open, reproducible science. In short, it turns years of specialized imaging work into a shared community asset for tackling chronic liver disease.
Citation: Schraven, S., Gonzalez, C., Baskaya, F. et al. A preclinical CT and MRI Liver Imaging Dataset with Anatomical, Functional and Segmentation Data. Sci Data 13, 446 (2026). https://doi.org/10.1038/s41597-026-07003-x
Keywords: chronic liver disease, preclinical imaging, MRI, CT, mouse models