Clear Sky Science · en
A chromosome-level genome assembly of the South African indigenous, Kolbroek pig, Sus scrofa domesticus
A Tough Little Pig with a Big Genetic Story
The Kolbroek pig may not look like a high-tech animal, but this hardy South African farm pig carries a genetic toolkit that helps smallholder farmers cope with heat, parasites, and poor-quality feed. Until now, its DNA had never been mapped in full detail. This study delivers the first complete, chromosome-level genome for the Kolbroek, creating a blueprint that can help conserve the breed, unlock its useful traits, and guide more sustainable pork production in regions facing tough environmental conditions.
Why This Local Pig Matters
South Africa imports much of its pork, and large commercial farms tend to use exotic pig breeds that grow fast and produce large litters. In contrast, smallholder farmers often rely on indigenous pigs like the Kolbroek. These pigs are smaller and less prolific, but they are well adapted to local heat, diseases, and low-cost feeds, including fibrous plants and tannin-rich grains such as red sorghum. Past genetic surveys showed that indigenous pigs in southern Africa hold rich genetic diversity, but they are also at risk: crossbreeding with commercial animals can dilute their unique traits, and limited breeding programs mean that inbreeding is a growing concern.
A Genetic Map to Protect a Heritage Breed
To safeguard the Kolbroek and make better use of its strengths, breeders need a high-quality reference genome—a complete, ordered list of its DNA that can be compared to other pigs. Existing commercial pig genomes, such as that of the Duroc breed, do not accurately reflect South African indigenous animals, making genetic tests less reliable. The new Kolbroek genome changes this. It provides a detailed view of the animal’s chromosomes, allowing researchers to spot the DNA regions that shape important traits like disease resistance, heat tolerance, and the ability to thrive on rough forage. It also lays the groundwork for designing genetic tests tailored to African pigs, instead of relying on tools built from European and Asian breeds.
How the Genome Was Built
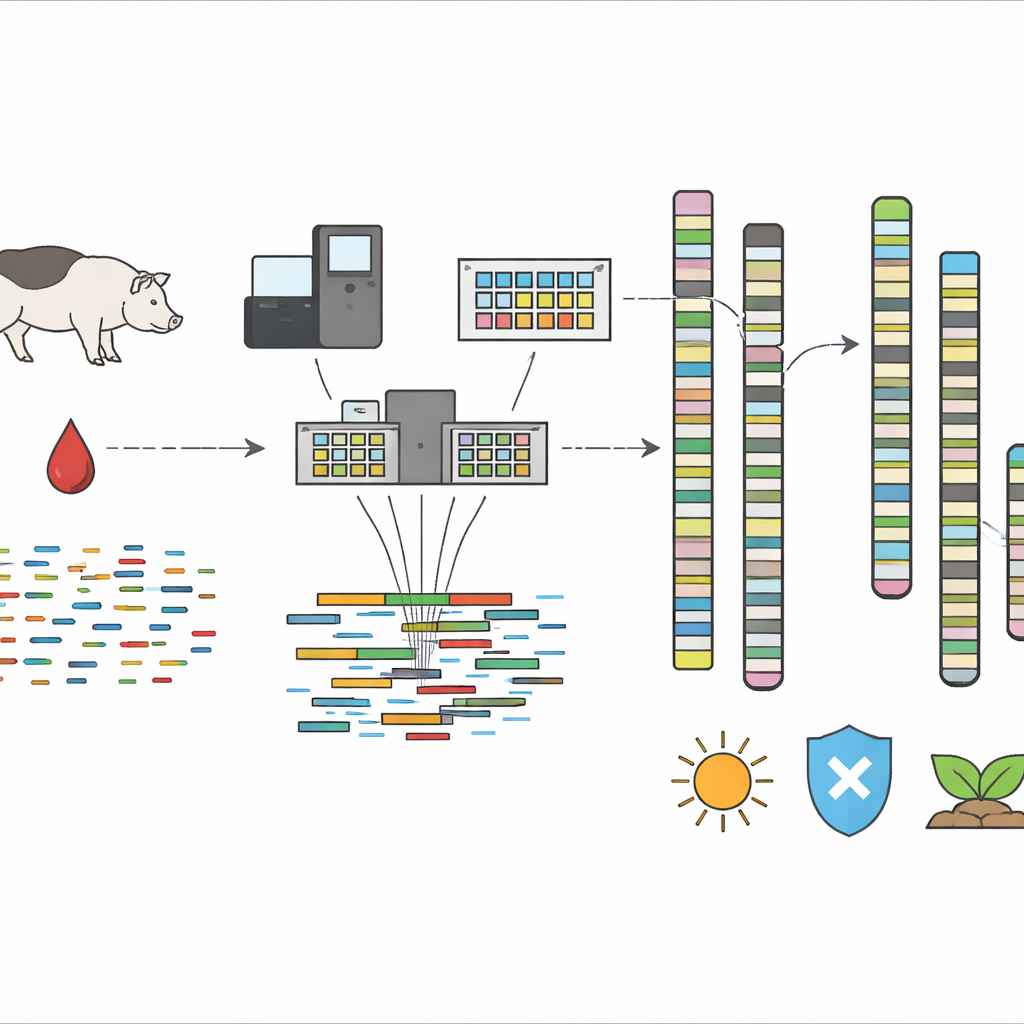
The researchers started with a single pure-bred Kolbroek sow from a stud farm in South Africa. From a carefully collected blood sample, they extracted very long strands of DNA and sequenced them using two advanced technologies. One, known as HiFi sequencing, reads long stretches of DNA with high accuracy. The other, Omni-C, captures how DNA folds inside the cell, helping to link distant pieces into whole chromosomes. A specialized assembly pipeline, developed by the Vertebrate Genome Project and run on the Galaxy Europe computing platform, stitched billions of DNA fragments together, removed contaminants like stray mitochondrial sequences, and arranged the pieces into 19 chromosome-scale scaffolds plus additional fragments.
What the DNA Reveals
The finished genome spans about 2.6 billion DNA letters, similar in size and structure to other domestic pigs. Quality checks showed that more than 95% of expected genes are present and correctly assembled. The team found that repeat elements—stretches of DNA that occur many times in the genome—make up roughly 38% of the Kolbroek’s DNA, a pattern that matches other mammals. Using a machine-learning–based gene prediction tool, they identified 22,025 protein-coding genes. When they compared the Kolbroek’s genome to standard pig references, they saw that its chromosomes line up closely overall, but also harbor structural differences and unique genetic variants that likely underlie its special adaptations to African environments.
From DNA Map to Better Pigs and Better Farms
By turning the Kolbroek’s DNA into a high-quality, public genome, this study gives breeders and scientists a powerful new tool. The assembly can be used to design affordable local genetic tests, search for variants linked to fertility, growth, meat quality, and resilience, and include South African pigs in global “pangenome” efforts that capture the full diversity of the species. In practical terms, this means future breeding programs can aim to improve productivity without losing the Kolbroek’s toughness and environmental fit—helping secure both a unique genetic heritage and more sustainable pork production for smallholder and commercial farmers alike.
Citation: Smith, R.M., Molotsi, A.H., Nesengani, L.T. et al. A chromosome-level genome assembly of the South African indigenous, Kolbroek pig, Sus scrofa domesticus. Sci Data 13, 635 (2026). https://doi.org/10.1038/s41597-026-07002-y
Keywords: Kolbroek pig, genome assembly, indigenous livestock, pig breeding, South Africa