Clear Sky Science · en
Aptamer-functionalized magnetic beads combined with hybridization chain reaction for detection of Bacillus cereus
Why this matters for everyday eating
Food poisoning from takeout rice, milk, or ready‑to‑eat meals is often blamed on a hardy microbe called Bacillus cereus. Traditional lab tests to spot this germ can take days, which is far too slow for busy kitchens, food companies, or safety inspectors. This study presents a fast, highly sensitive lab method that could one day help keep foods like milk and rice safer by detecting even very low levels of B. cereus without expensive equipment or complicated genetics work.
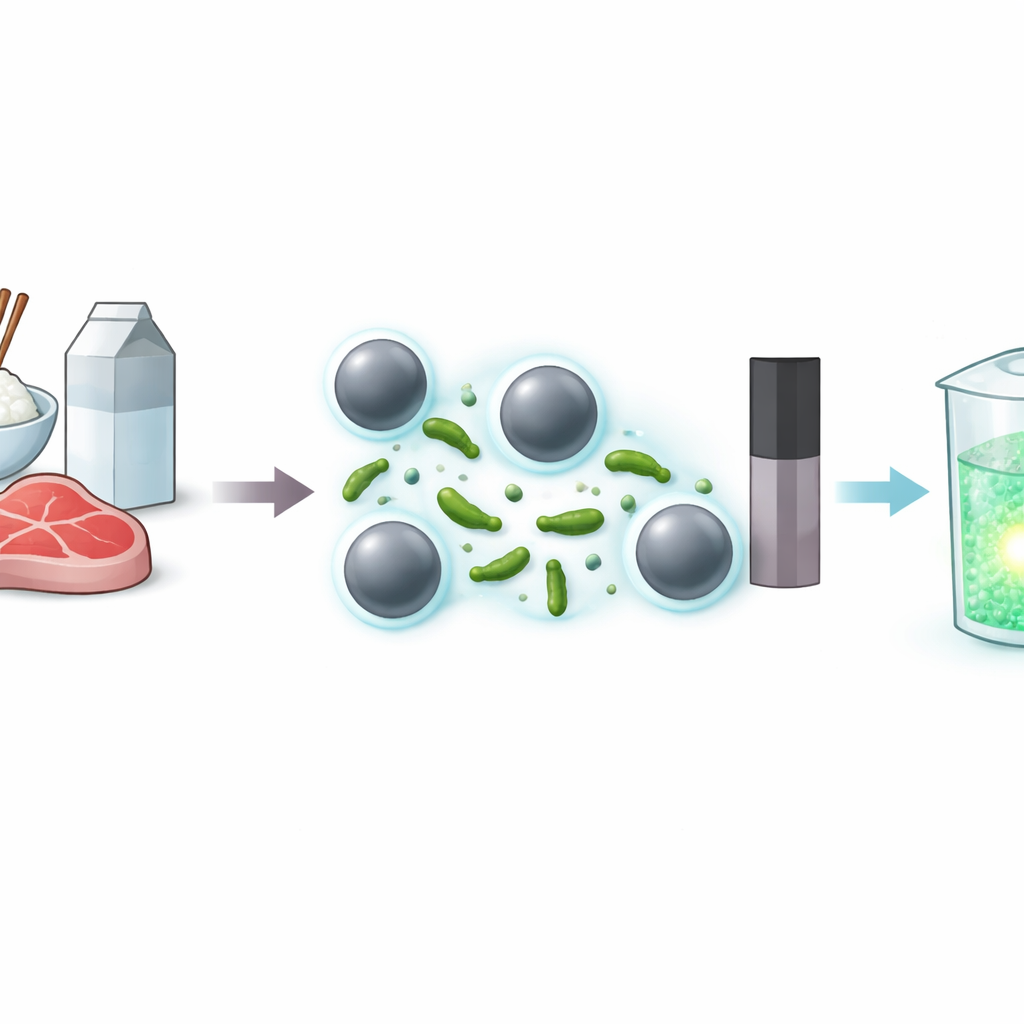
The hidden threat in common foods
Bacillus cereus is widespread in everyday foods such as rice, milk, and meat. It can survive harsh conditions, form tough spores, and produce toxins that trigger vomiting and diarrhea. Standard detection relies on growing the bacteria on plates, which is accurate but slow and labor‑intensive, making it impractical for rapid on‑site checks. Molecular tests like PCR are faster, but they depend on careful DNA extraction, specialized instruments, and strict contamination control. Other antibody‑based tests can be quicker but often lack the needed sensitivity or remain too costly for routine screening.
Using smart “locks” and tiny magnets
The authors build on a newer class of biological “locks” called aptamers—short strands of DNA that fold into shapes that recognize a specific target, here B. cereus cells. These aptamers are attached to magnetic beads, creating tiny particles that can selectively grab the bacteria out of a complex sample. When a mixture containing B. cereus is added, the bacteria bind to the aptamer‑coated beads. After a magnet pulls the beads (and any captured bacteria) to the side, the rest of the sample can be poured away, effectively enriching the target and removing much of the interfering background found in real foods.
Turning captured bacteria into a bright signal
The clever twist is how the presence of bacteria is converted into a strong fluorescent glow. A short DNA strand tagged with a fluorescent dye is designed to compete with B. cereus for the same aptamer on the magnetic bead. If no bacteria are present, this fluorescent strand mostly sticks to the beads and is removed with them, leaving little signal. If bacteria are present, they out‑compete the strand, which is released into the solution. That freed strand then triggers a hybridization chain reaction, in which two hairpin‑shaped DNA pieces repeatedly open and link together into long double‑stranded chains carrying many dye molecules. This acts like a built‑in amplifier, turning a tiny molecular event into a bright, measurable signal.
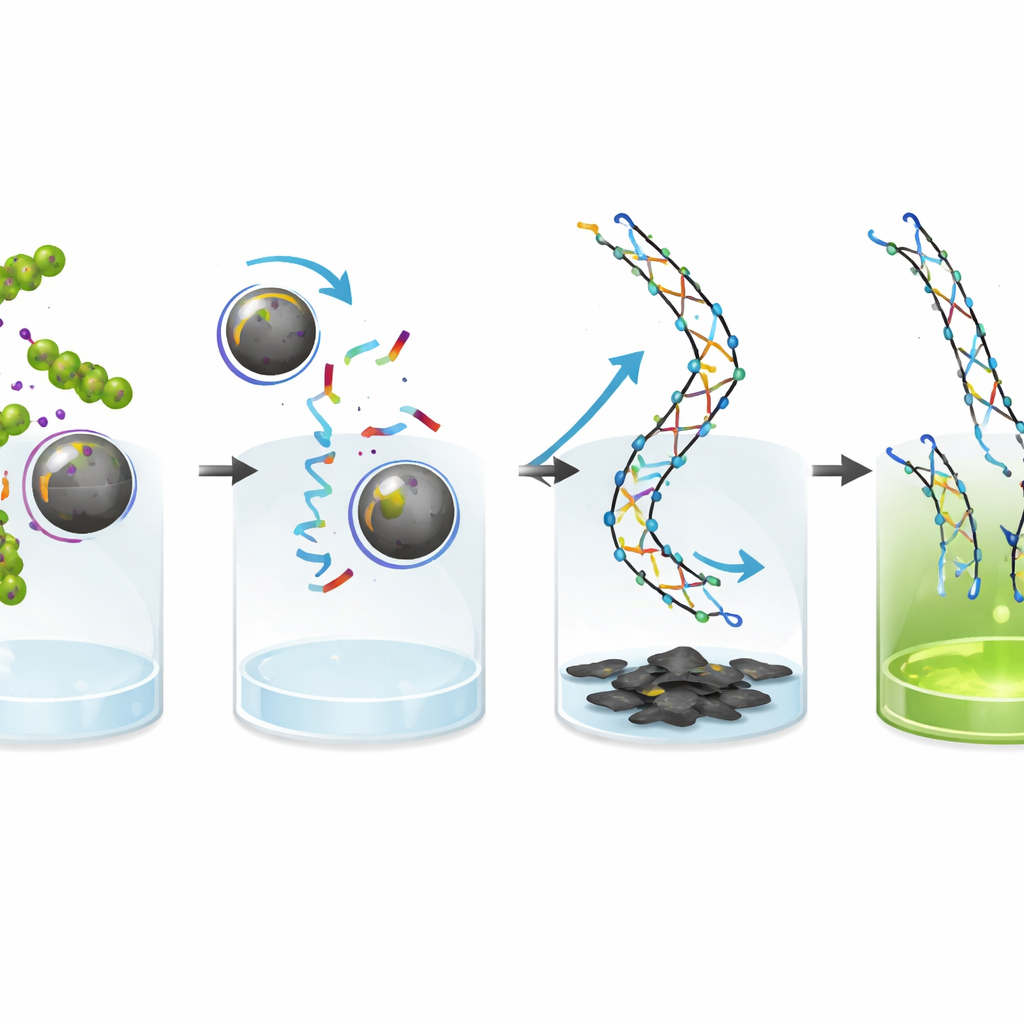
Making the glow clearer with a carbon “eraser”
To further sharpen the readout, the team uses graphene oxide, a sheet‑like carbon material that strongly sticks to loose single DNA strands and efficiently quenches their fluorescence. Short, unreacted fluorescent strands are pulled down onto the graphene oxide surface and their light is effectively shut off. In contrast, the long, stiff double‑stranded products of the chain reaction do not bind well to graphene oxide, so their glow remains bright. This combination greatly improves the contrast between “bacteria present” and “bacteria absent,” helping the system detect very low bacterial concentrations with confidence.
How well the method performs in the lab
After carefully tuning conditions such as aptamer loading on the beads, reaction times, and the amount of graphene oxide, the authors show that their setup can specifically recognize B. cereus among a panel of other common foodborne bacteria. The fluorescent signal from true B. cereus strains is much higher than from non‑targets or blank samples. In pure bacterial suspensions, the method can detect down to about 5.4 × 101 colony‑forming units per milliliter—comparable to many advanced rapid tests in the field. From initial sample handling to final readout, the full process takes around two hours, much faster than culture‑based methods and simpler than many DNA‑amplification approaches.
Testing in real milk and looking ahead
To see whether the method might work beyond clean lab samples, the researchers spiked commercial milk with known amounts of B. cereus and ran the same protocol. They still observed strong, concentration‑dependent fluorescent signals over a similar range as in pure culture, showing that the approach can function in a realistic food matrix. However, the technique detects only live bacteria with intact cell surfaces and currently depends on a benchtop fluorescence reader, so further engineering is needed to adapt it into portable, field‑ready devices. The authors suggest that, by swapping in different aptamers, the same framework could be retargeted to many other foodborne pathogens.
What this means for safer food
In plain terms, this work demonstrates a promising laboratory method that uses smart DNA locks, magnetic beads, and a self‑building fluorescent signal to spot very small amounts of B. cereus quickly and selectively. While it is not yet a handheld test, it points toward future tools that could screen foods like milk or rice in a couple of hours rather than days, reducing the chance that dangerous batches reach consumers. With further development and adaptation, similar strategies could help build a new generation of rapid tests to keep a wide range of foodborne germs in check.
Citation: Song, X., Shi, C., Lv, Y. et al. Aptamer-functionalized magnetic beads combined with hybridization chain reaction for detection of Bacillus cereus. npj Sci Food 10, 124 (2026). https://doi.org/10.1038/s41538-026-00751-5
Keywords: Bacillus cereus detection, food safety, aptamer biosensor, magnetic beads, hybridization chain reaction