Clear Sky Science · en
Machine learning assisted single-molecule sensing towards standard-free quantification of per- and polyfluoroalkyl carboxylic acids
Why this matters for everyday water safety
Invisible industrial chemicals known as PFAS have seeped into rivers, drinking water, and even our blood. Among them, per- and polyfluoroalkyl carboxylic acids (PFCAs) are especially worrisome, yet they are extremely hard to measure accurately because reference standards exist for only a tiny fraction of the thousands of PFAS in use. This study introduces a new way to "count" individual PFCA molecules one by one as they pass through a tiny biological hole, using machine learning to recognize their electronic fingerprints—without needing a matching lab standard for each compound.
Hidden chemicals in a complex family
PFAS are a sprawling family of fluorine-rich chemicals used in products from nonstick pans to firefighting foams. Many differ only by a few atoms, but these tiny structural tweaks can dramatically change how they move through the environment, build up in living organisms, or affect health. Traditional techniques such as liquid or gas chromatography coupled to mass spectrometry can detect many PFAS with great sensitivity, but they usually require a pure standard of each chemical to identify and quantify it reliably. So far, commercial standards exist for just over a hundred PFAS—less than one percent of those known—leaving regulators and scientists largely in the dark about the rest.
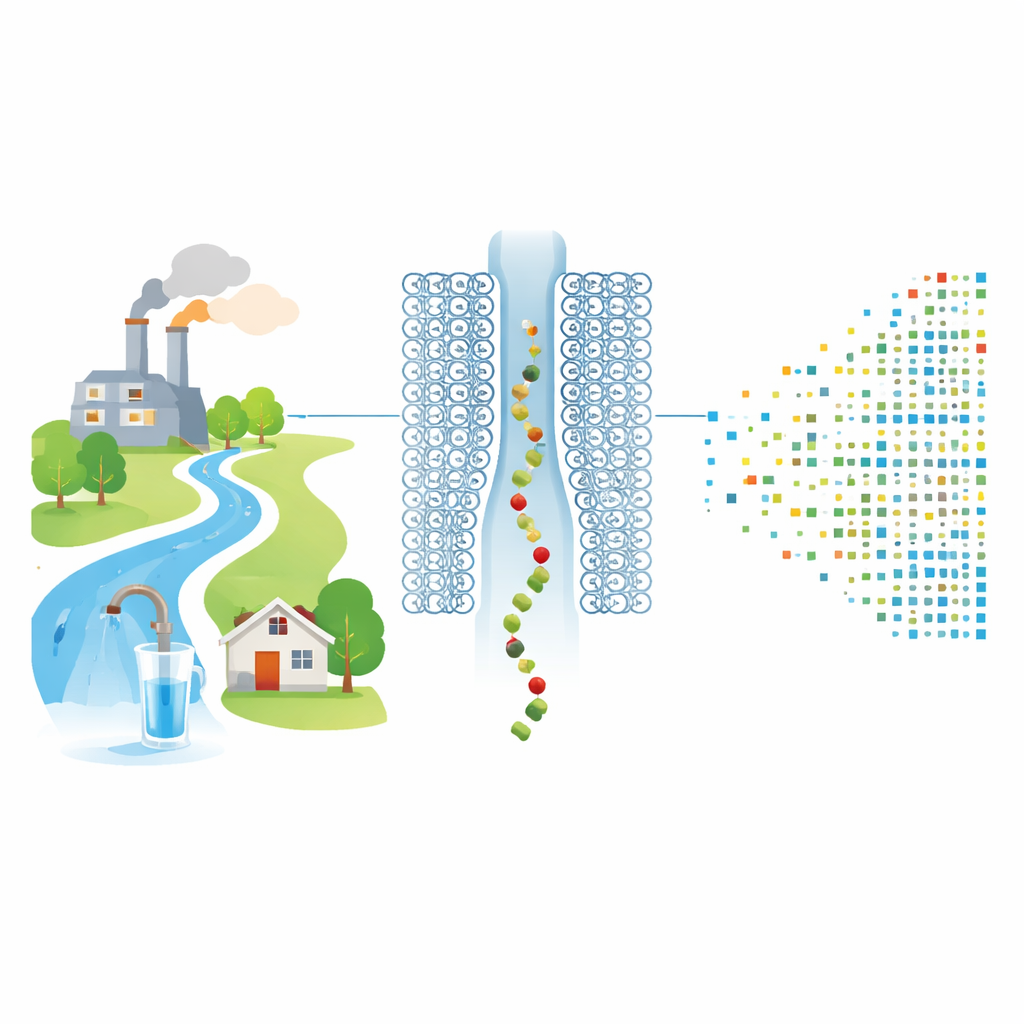
Counting single molecules through a tiny gateway
The researchers tackle this gap using a protein nanopore: a donut-shaped molecule that forms a single hole in a lipid membrane. When a voltage is applied, ions flow through the pore and create a steady electrical current. The team chemically tethers individual PFCA molecules to short positively charged peptide "leaders" that are drawn into the nanopore like beads on a string. As each PFCA–peptide pair enters and occupies the pore, it partially blocks the ion flow, causing a brief dip in current whose depth and duration depend on the size and shape of the molecule in the pore.
Turning pore signals into a molecular ruler
A key breakthrough of this work is that these current dips behave like a precise measuring stick. By combining experiments with molecular simulations, the authors show that, for a series of PFCA molecules up to 14 carbons long, the depth of the current blockade increases in an almost perfectly straight line with the molecule’s volume. In other words, once the nanopore and peptide are chosen, volume alone predicts how strongly the current is blocked. This allowed the team to forecast the electrical signature of other, more complex PFCAs—such as those with hydrogen or chlorine substitutions, side chains, or aromatic rings—and confirm experimentally that the predictions matched reality within the tiny margin of measurement error.
Machine learning that spots look‑alike pollutants
Because many PFCA cousins are so similar in size that their blockades overlap, the scientists then exploited the full richness of the nanopore signal. They extracted dozens of features from each event, including how long it lasts, how noisy it is, and how its shape changes when the signal is digitally filtered at different frequencies. Using machine-learning models trained on these multi-dimensional fingerprints, they achieved near-perfect (about 99.9%) identification accuracy across 13 PFCA types, including closely related isomers. By carefully selecting the most informative 21 features, they reduced model complexity while actually improving performance, even when the target chemical was buried among interfering PFCAs at 100 times higher concentration.
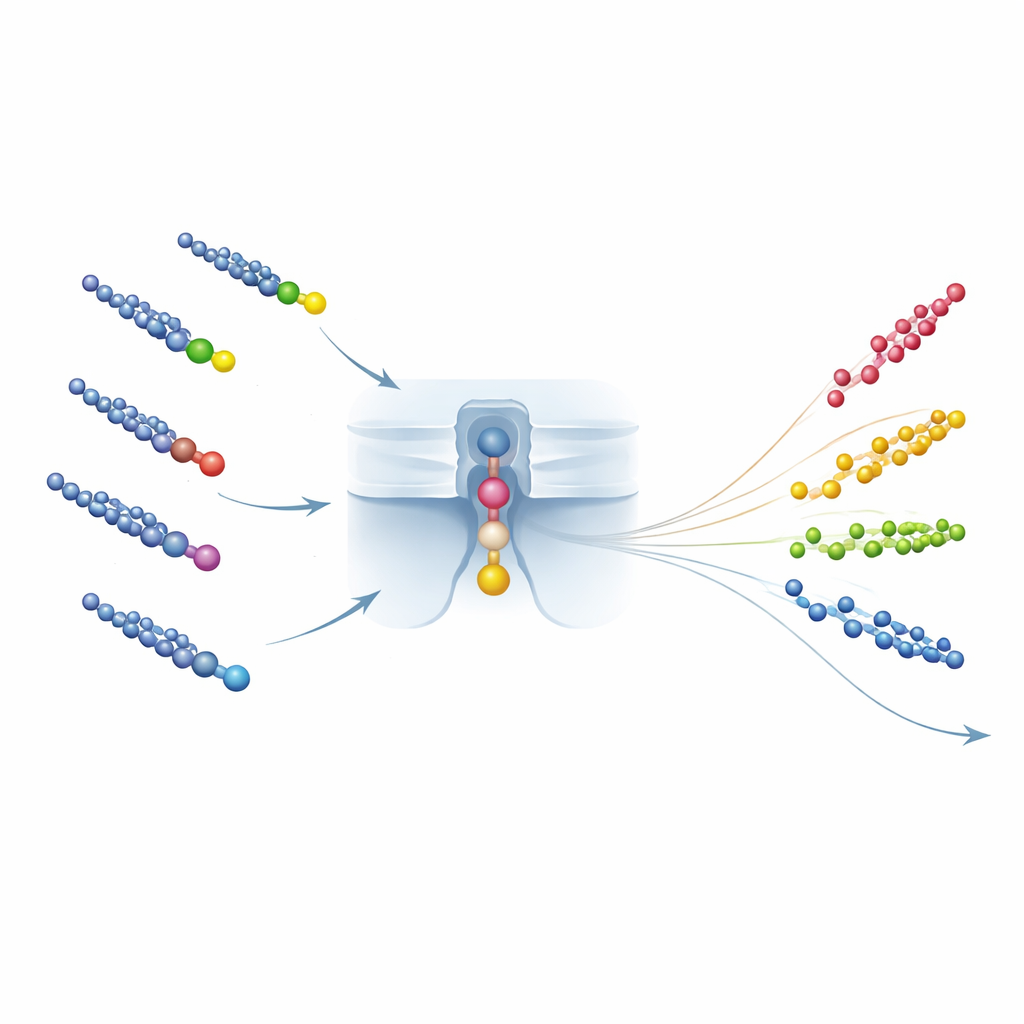
From single-molecule counts to real-world water tests
Beyond identification, the method also needs to measure how much of each PFCA is present. Here the team exploits the rate at which PFCA–peptide pairs are captured by the nanopore: the average time between events shrinks as concentration rises. Clever peptide design ensures that this capture rate is set mainly by the peptide’s charge and the pore’s electric landscape, and much less by which PFCA is attached. That means a single calibration curve—relating event frequency to concentration—can be shared across many PFCAs, enabling what the authors call "one calibration curve fits all." They validate this universality in mixtures and in complex samples such as tap water and serum, showing accurate counts even in the presence of many other chemicals, and reaching detection limits for the ultrashort PFCA trifluoroacetic acid comparable to the best mass-spectrometry methods.
A new route to tracking PFAS without custom standards
Taken together, this work outlines a path to monitoring a broad swath of PFCA pollutants without needing a tailor-made standard for each one. A carefully engineered nanopore and peptide probe create a simple linear link between molecular size and signal, while machine learning teases apart subtle signal features to distinguish even nearly identical isomers. By further tuning the pore’s entry and exit "barriers," the authors show, in experiments and simulations, how the same strategy could be extended to longer-chain PFCAs and potentially to the wider universe of PFAS. For the general public, this means a promising new way to see and measure chemicals that have long remained largely invisible in our water and environment.
Citation: Zuo, J., Li, HS., Tang, W. et al. Machine learning assisted single-molecule sensing towards standard-free quantification of per- and polyfluoroalkyl carboxylic acids. Nat Commun 17, 3923 (2026). https://doi.org/10.1038/s41467-026-70718-3
Keywords: PFAS, nanopore sensing, single-molecule detection, machine learning, water contaminants