Clear Sky Science · en
Metagenomic profiling of antimicrobial resistance in wastewater from metropolitan cities of India
Why Dirty Water Matters to Everyone
In big cities, what we flush and wash away doesn’t simply vanish. It collects in drains and sewers, forming a swirling mix of human waste, industrial runoff, and countless microbes. This hidden world can act as an early warning system for public health—especially for tracking bacteria that no longer respond to antibiotics. This study peered into the wastewater of four of India’s largest metropolitan areas to see which microbes are present, which antibiotic resistance genes they carry, and how those genes might be spreading. The findings help explain how our shared environment can quietly fuel the global crisis of drug-resistant infections.
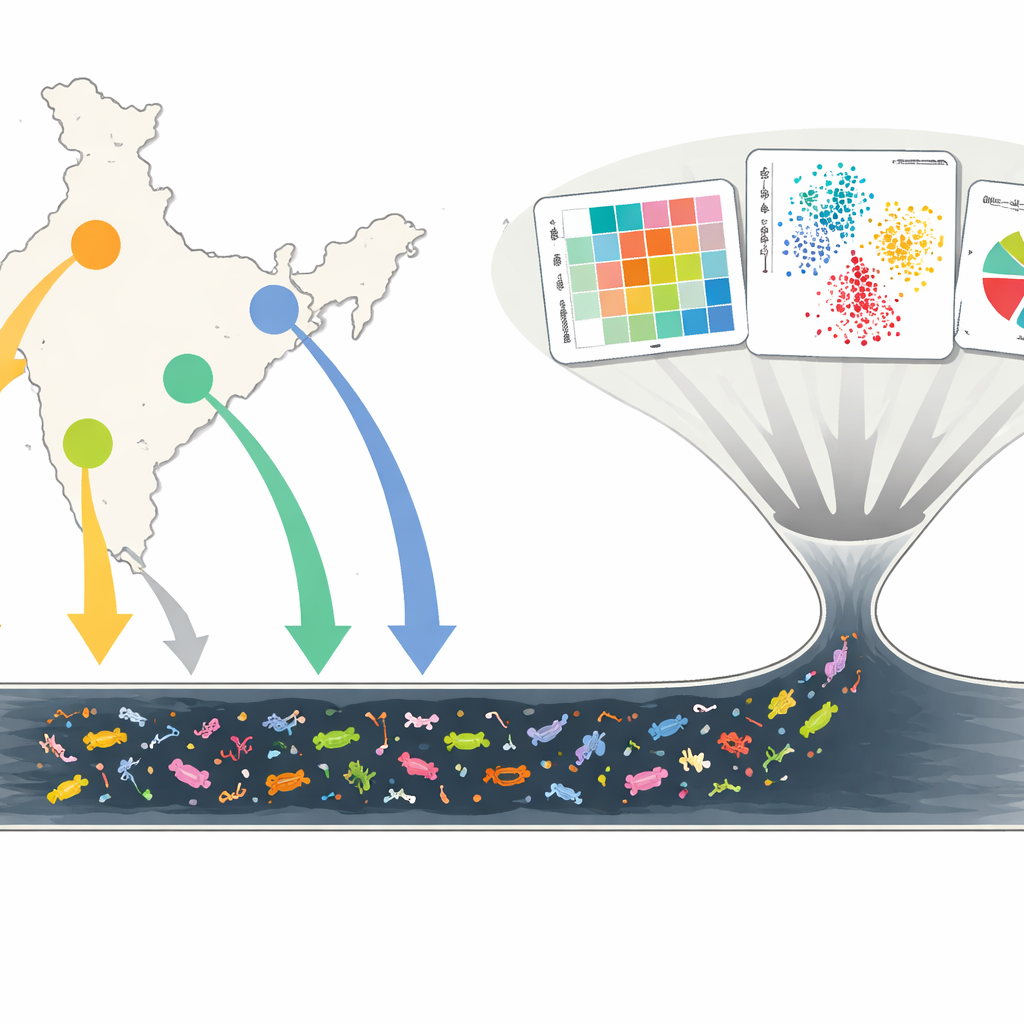
Peeking Into the City’s Underground Soup
Researchers collected nearly 450 wastewater samples from 19 large open drains in Delhi, Mumbai, Chennai, and Kolkata, month after month for two years. Instead of growing microbes in the lab, they used shotgun metagenomic sequencing, a method that reads all DNA in a sample at once. This allowed them to catalog which bacteria were present (the microbiome), which resistance genes appeared (the resistome), and how often those genes were linked to mobile genetic elements—small pieces of DNA that can hop between microbes. By comparing samples over time and across cities, they built a detailed picture of how urban wastewater reflects the health risks and antibiotic pressures of millions of people.
City Microbes With Local Signatures
The bacterial communities in each city’s wastewater were distinct. While some broad groups of bacteria were common everywhere, such as Proteobacteria and Bacteroidetes, the exact mix of species varied from city to city and sometimes from month to month. Mumbai had the richest variety of species, while Kolkata had the least. Certain bacteria tied to human disease, including strains of Escherichia coli, Klebsiella pneumoniae, and Pseudomonas aeruginosa, were abundant in at least one city, underlining the presence of clinically important pathogens in everyday wastewater. The team also reconstructed thousands of near-complete bacterial genomes from the data and found that more than half could not be matched to known species, suggesting that India’s sewers harbor a vast, mostly uncharted microbial world.
Resistance Genes That Ignore City Borders
When the researchers turned from whole microbes to the resistance genes themselves, a different pattern emerged. Unlike the microbes, antibiotic resistance genes did not cluster neatly by city. Many of the same genes appeared across all four locations, hinting at a shared pool of resistance shaped by similar antibiotic use and environmental pressures. Genes that protect bacteria against multiple classes of drugs were especially common, as were those conferring resistance to macrolides, tetracyclines, rifamycins, and aminoglycosides. However, genes guarding against some of our most critical last-line antibiotics, such as certain beta-lactams and colistin, were relatively rare in these samples, offering a small measure of reassurance.
How Genes Hitchhike and Form Hidden Alliances
The team next asked how easily these resistance genes might spread. They searched bacterial genomes for mobile genetic elements—plasmids and transposons—that act like shuttles, moving DNA from one microbe to another. Only about one in ten resistance-bearing DNA fragments also carried such mobile elements, but this subset is crucial: it represents genes that are poised to jump. Resistance to common drug families like tetracyclines and beta-lactams was often found on these mobile segments, whereas many macrolide resistance genes were more static. By building “co-occurrence networks,” the researchers also showed that each city’s microbes formed their own interaction patterns, and that certain groups of bacteria contributed disproportionately to the local load of resistance genes.
What This Means for Public Health
Altogether, the study shows that city wastewater is not just dirty water—it is a sensitive mirror of both local microbial life and a widely shared pool of antibiotic resistance genes. Each city had its own microbial fingerprint, yet the resistance genes themselves were surprisingly similar, highlighting how easily antibiotic pressure can generate a common threat. The discovery of many potentially new bacterial species and mobile resistance combinations underscores how much we still do not know about the invisible ecosystem beneath our feet. By standardizing simple sampling and storage methods, the researchers demonstrate that large-scale wastewater surveillance is practical even in resource-limited settings. Such routine monitoring could help countries like India detect dangerous pathogens and emerging resistance earlier, guiding smarter policies and infrastructure to slow the spread of antimicrobial resistance before it reaches clinics and households.
Citation: Singh, N.K., Garg, P., Kumari, S. et al. Metagenomic profiling of antimicrobial resistance in wastewater from metropolitan cities of India. Nat Commun 17, 4097 (2026). https://doi.org/10.1038/s41467-026-70702-x
Keywords: wastewater surveillance, antimicrobial resistance, metagenomics, urban microbiome, India