Clear Sky Science · en
Gene expression dynamics of human and mouse craniofacial development at the single-cell level
How Our Faces Take Shape
The human face starts as a collection of tiny tissue buds that must grow, move, and fuse with exquisite precision. When these early steps go awry, the result can be common birth differences such as cleft lip and cleft palate. This study uses powerful single-cell tools to watch, in unprecedented detail, how thousands of individual cells in the developing face behave in early human embryos and in mice, and how their activity connects to both normal facial variation and disease.
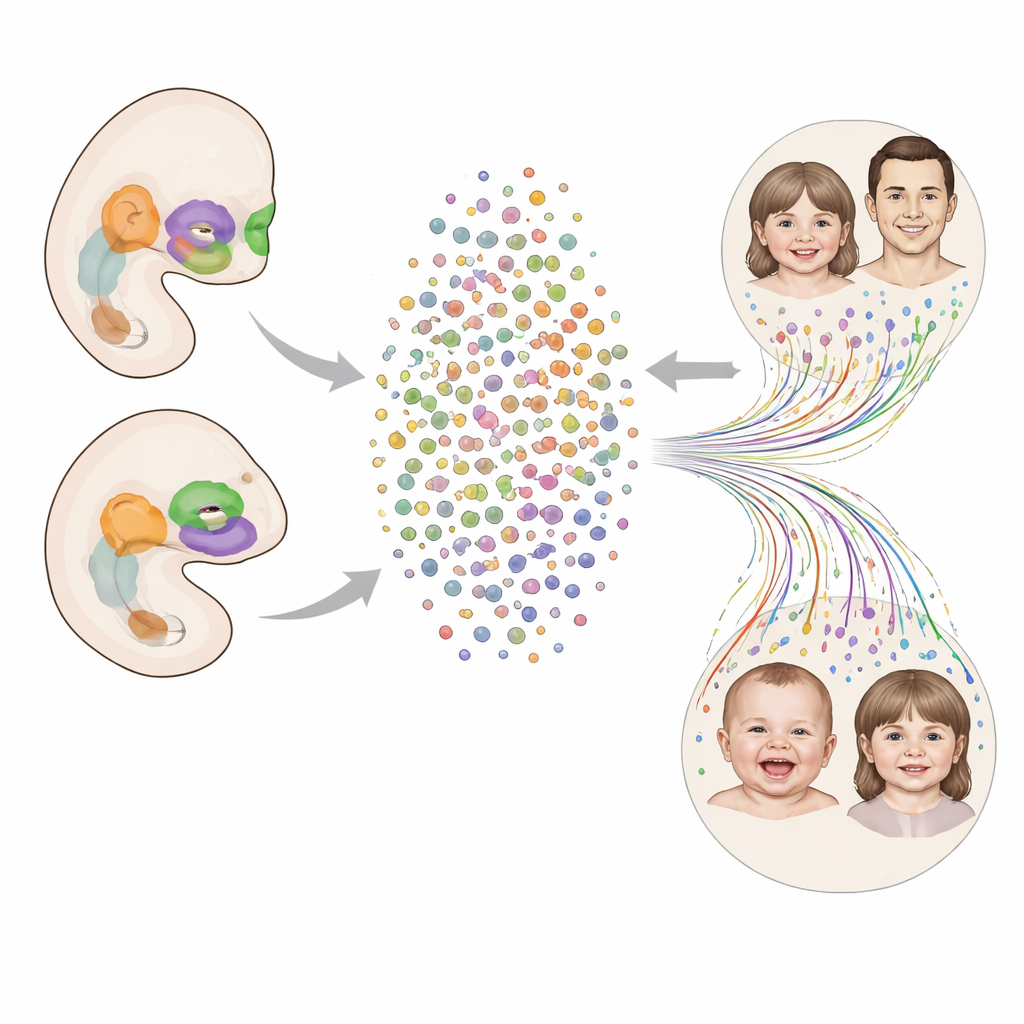
Looking at the Young Face Cell by Cell
The researchers built a detailed map, or atlas, of the developing human face between four and eight weeks after conception—exactly when the upper lip and palate are forming. They isolated nearly 96,000 cell nuclei from the facial region of 24 human embryos and measured which genes were active in each one. This allowed them to sort the cells into eight broad groups, including connective-tissue–like mesenchyme, surface layers derived from ectoderm, blood vessels, blood cells, immune cells, early muscle cells, cranial neural crest cells, and a pool of early progenitors that can still give rise to multiple lineages. By following how gene activity changes over developmental time, they could see how early progenitor cells branch into more specialized types.
Comparing Human and Mouse Faces
To understand which features of facial development are shared across species and which are unique to humans, the team performed matching experiments in mice. They collected face tissue from mouse embryos at stages that roughly align with the human samples and again profiled tens of thousands of cells at single-cell resolution. Most major cell types were strikingly similar between species, using overlapping sets of genes to carry out their roles. However, cranial neural crest cells—migratory cells that build much of the face—showed the least conservation, hinting that changes in this population may underlie evolutionary differences in facial shape. The human samples also contained a distinct group of early progenitor cells that did not appear as a separate cluster in mouse faces, suggesting a possible human-specific feature or difference in sampling.
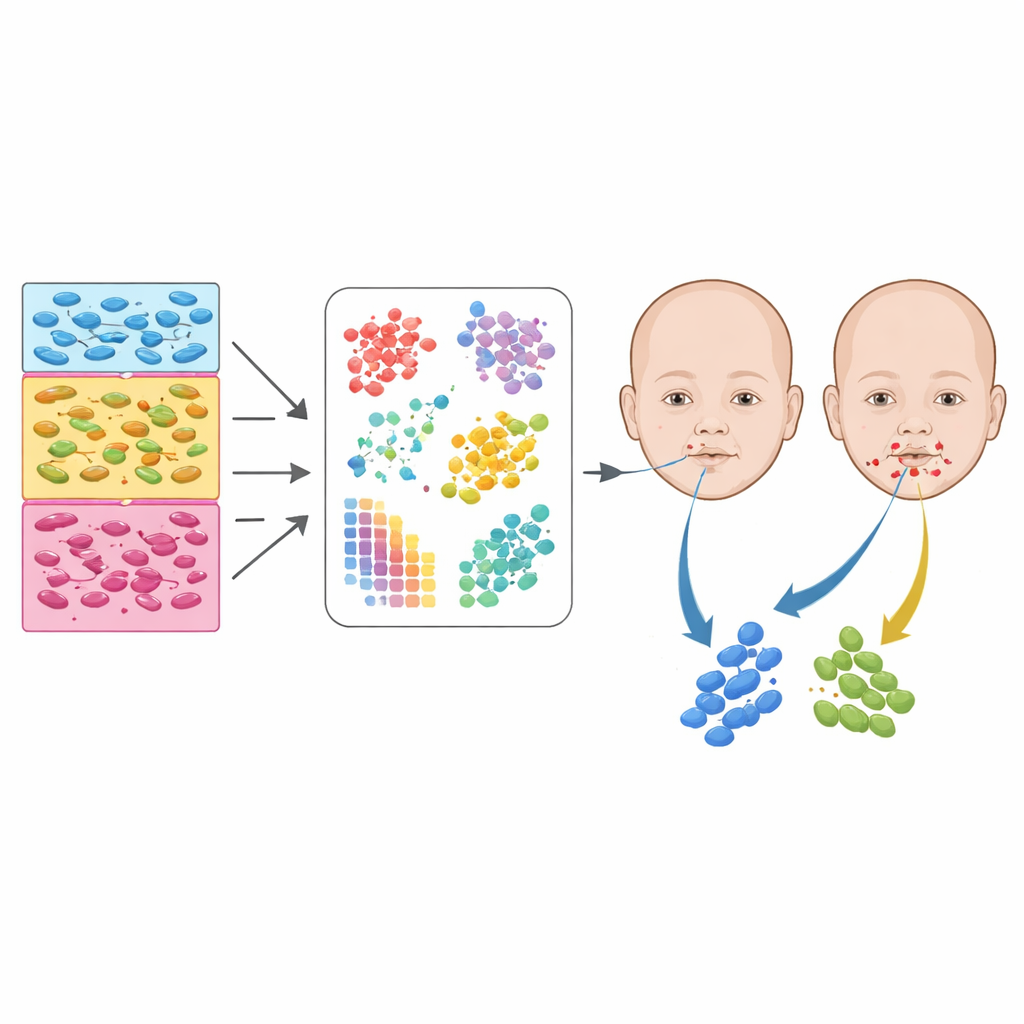
Zooming In on Specialized Cell Neighborhoods
Beyond broad categories, the authors identified dozens of finer subtypes within the mesenchyme, ectoderm, and neural crest–related populations. For example, they separated mesenchymal cells into those destined to form nasal structures, the upper jaw, palatal shelves, cartilage, and bone-forming osteoblasts. Ectodermal cells were parsed into surface layers, inner ear precursors, eye cells, pituitary and thyroid progenitors, and several distinct palate-related surfaces. By combining this atlas with spatial gene expression data—thin sections of embryos where gene activity is mapped back onto physical positions—the team could assign many subtypes to specific regions, such as the frontonasal area, the pharyngeal arches, or the exact zones where facial processes fuse.
From Cell Types to Facial Traits and Birth Defects
The power of this atlas emerges when it is combined with human genetic studies. Using large datasets that link common genetic variants to subtle differences in facial shape, the authors asked which cell subtypes express the genes near those variants. Traits that depend on bone or cartilage, like jaw and chin projection or the spacing of the nose and eyes, were most strongly tied to mesenchymal subtypes. In contrast, measurements of soft tissues—like lip thickness or ear size—were enriched in ectodermal surface cells. When the team overlaid genetic studies of cleft lip and cleft palate, they saw that risk variants clustered in genes active in specific fusion-related mesenchyme and palate surface ectoderm. In parallel, rare, newly arisen protein-altering mutations in children with orofacial clefts were particularly common in genes switched on in certain ectodermal and epithelial subtypes, underscoring the importance of these thin surface layers, even though they make up a small fraction of facial tissue.
What This Means for Understanding Faces
Taken together, the work shows that our facial appearance and our risk for clefting emerge from the combined behavior of many distinct cell subtypes acting in narrow developmental windows. The atlas reveals which exact cell neighborhoods in the early face are most sensitive to genetic changes, offering a roadmap for future studies of how individual genes and variants alter development. For non-specialists, the key message is that birth differences like cleft lip and palate are not caused by a single “face gene,” but by disruptions in intricate cellular communities that build the nose, lips, and palate. By mapping these communities in both humans and mice, this study lays the groundwork for better diagnosis, targeted therapies, and deeper insight into how subtle changes in early development sculpt the diversity of human faces.
Citation: Khouri-Farah, N., Manchel, A., Wentworth Winchester, E. et al. Gene expression dynamics of human and mouse craniofacial development at the single-cell level. Nat Commun 17, 3714 (2026). https://doi.org/10.1038/s41467-026-70232-6
Keywords: craniofacial development, single-cell RNA sequencing, orofacial clefts, neural crest cells, facial genetics