Clear Sky Science · en
Reconstructing single-cell resolution from spatial transcriptomics with CellRefiner
Seeing Cells in Their Neighborhoods
Modern biology can now read the genetic activity of hundreds of thousands of individual cells at once, but often loses track of where those cells actually sit inside a tissue. At the same time, new "spatial" methods can map gene activity across a tissue slice, yet typically blur together several cells into each measurement. This article introduces CellRefiner, a computational approach that combines the strengths of both worlds to reconstruct where individual cells are located and how they touch one another inside real tissues. The result is a much sharper picture of how cells are arranged and how they communicate, particularly in the brain, lymph nodes, and tumors.
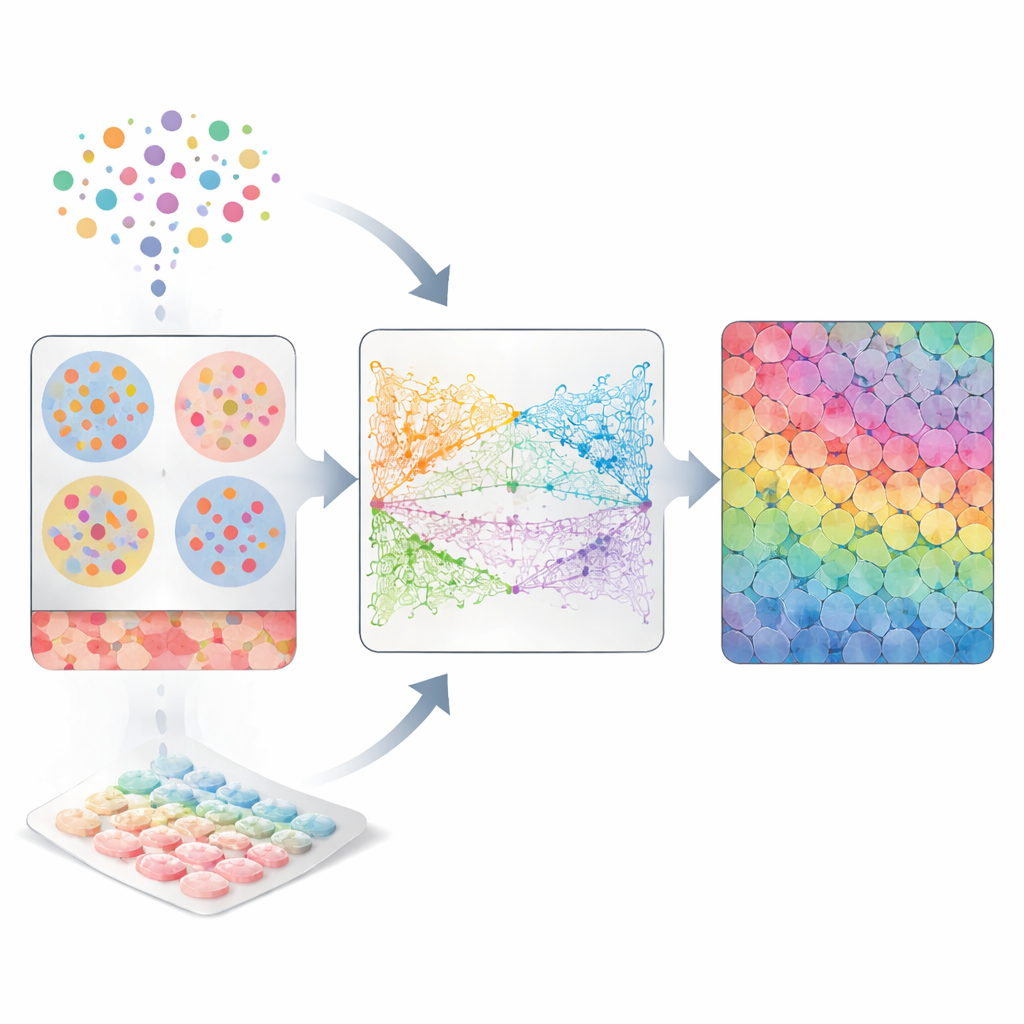
Two Imperfect Views of Living Tissues
Biologists commonly use single-cell RNA sequencing to capture which genes are turned on in individual cells. To do this, tissue must be broken apart, so each cell becomes a free-floating unit with rich molecular information but no address. Spatial transcriptomics takes the opposite approach: it keeps cells in place on a slide and reads out gene activity in small spots across the tissue. However, each spot often contains several mixed cells, and many imaging-based platforms only measure a subset of genes. As a result, neither technology alone can fully answer questions that depend on knowing both what each cell is doing and exactly where it sits, such as which cells are neighbors or which pairs are in physical contact.
A Physics-Inspired Mapmaker for Cells
CellRefiner tackles this gap by treating cells like tiny interacting particles that can move within the tissue until they settle into realistic positions. First, it uses existing methods to roughly assign groups of single cells to each spatial spot based on how similar their gene activity is to what is measured in that spot. These assigned cells are initially scattered at random inside each spot. Then CellRefiner applies three kinds of virtual "forces" to gradually rearrange them: one force keeps cells from overlapping or leaving empty gaps, another gently pulls together cells with similar gene activity, and an optional third force pulls together cells that show matching "sender" and "receiver" molecules used in cell-to-cell signaling. Over several iterations, this simulation sharpens a coarse spot-level picture into a plausible single-cell map.
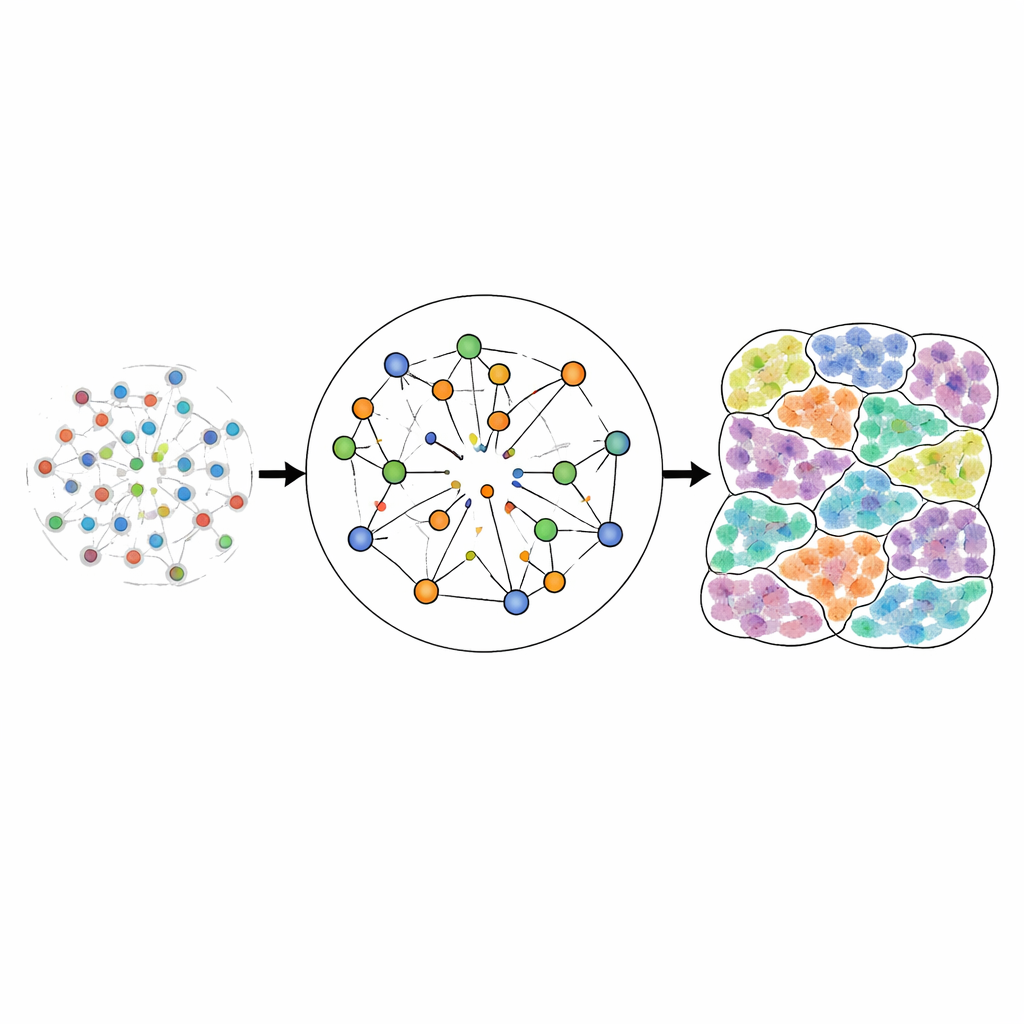
Testing the Method Across Many Tissues
To check whether CellRefiner recovers realistic structures, the authors first created test datasets where the true positions of cells are known. They started from high-resolution imaging-based data in which each cell is individually mapped, then artificially blurred these data into lower-resolution spots that mimic popular spatial platforms. Feeding only the blurred version plus the original single-cell profiles into CellRefiner, they asked how well the method could reconstruct the original fine-scale map. Using several measures of spatial error, CellRefiner consistently improved cell placement over the initial rough assignments and outperformed other leading methods across multiple datasets from mouse brain regions and other tissues. It captured sharp structures such as narrow bands of ependymal cells, layered hippocampal regions, and cortical layers more faithfully than competing approaches.
From Points to Shapes and Conversations
Beyond just placing cell centers, CellRefiner can also reconstruct approximate cell shapes. It represents each cell as a cluster of many small elements connected by virtual springs, which respond to pushing and pulling forces between neighboring cells. This allows the method to infer which cells are in direct physical contact, a key requirement for studying contact-based signaling, where molecules on one cell’s surface bind to partners on a neighboring cell. When applied to high-resolution imaging datasets, the reconstructed shapes closely matched the observed cell outlines and recovered realistic contact networks. Applied to lower-resolution platforms such as Visium, CellRefiner produced detailed contact maps that the original spot-level data cannot provide.
Revealing Hidden Signaling in Brain and Cancer
Using its refined maps and contact networks, CellRefiner was able to rediscover known signaling patterns in human squamous cell carcinoma and mouse cortex. In tumors, it highlighted signaling systems involved in cell adhesion, blood vessel growth, and interactions at the tumor border, including pathways that help cancer cells stick together or engage with their surroundings. In the mouse brain, CellRefiner revealed structured signaling between cortical layers and specific classes of interneurons, supporting roles in neuron migration, circuit wiring, and synapse formation. Importantly, the method showed that what looks like strong signaling in a single mixed spot can actually arise from only a subset of the cells inside that spot, exposing hidden diversity in how nearby cells respond.
Sharper Tissue Maps for Future Biology
In essence, CellRefiner turns blurry spatial measurements into detailed maps where each cell has both a molecular identity and a realistic position and shape. This sharper view enables more trustworthy studies of how cells are organized, how they cluster into layers and regions, and how they communicate through direct contact. While the method depends on the quality of input data and makes simplifying assumptions about cell density and size, it offers a flexible, physics-inspired framework that can be extended to other molecular layers such as proteins or chromatin. For non-specialists, CellRefiner represents a step toward virtual microscopes that show not just where cells are, but how they interact as dynamic communities inside living tissues.
Citation: Bourgain-Chang, E., Kuang, X., Cang, Z. et al. Reconstructing single-cell resolution from spatial transcriptomics with CellRefiner. Nat Commun 17, 3304 (2026). https://doi.org/10.1038/s41467-026-70090-2
Keywords: spatial transcriptomics, single-cell RNA sequencing, cell-cell communication, computational biology, tumor microenvironment