Clear Sky Science · en
Dual deconvolution in multiphoton structured illumination microscopy for deep-tissue super-resolution imaging
Seeing Deeper into Living Tissues
Modern biology often depends on seeing the tiniest details inside thick pieces of tissue, such as brain slices or developing embryos. Unfortunately, as light travels through these crowded environments, it gets bent and scrambled, making images blurry just when scientists most want sharp views. This article introduces a way to digitally "unsmear" those images, allowing a standard advanced microscope to reveal extremely fine structures deep inside tissue without adding costly, complex hardware.
Why Deep Imaging Is So Difficult
Fluorescence microscopes let researchers label specific molecules and watch how cells and tissues are built and behave. Over the past decades, several “super-resolution” methods have beaten the traditional limits of sharpness, revealing details well below 200 nanometers. However, most of these methods only work well near the surface of a sample. In thick tissues such as mouse brain, the light used to excite fluorescent labels and the light emitted back to the detector both get distorted by tiny variations in the tissue’s structure. These distortions, called aberrations, blur the microscope’s focus and erase high‑detail information, especially at depths greater than tens of micrometers.
From Hardware Fixes to a Software Solution
A popular way to fight aberrations is hardware-based adaptive optics, which uses movable mirrors or other devices to reshape the light wavefront and restore sharp focus. While powerful, these systems are expensive, technically demanding, and often only correct distortions for one color or one direction of light travel at a time. The authors instead propose a computational approach that works with a type of microscope already common in many labs: a laser-scanning multiphoton microscope. By swapping the usual single light detector for a camera, they record a rich stack of scanned images that encodes how both the incoming excitation light and the outgoing fluorescence have been distorted by the tissue.
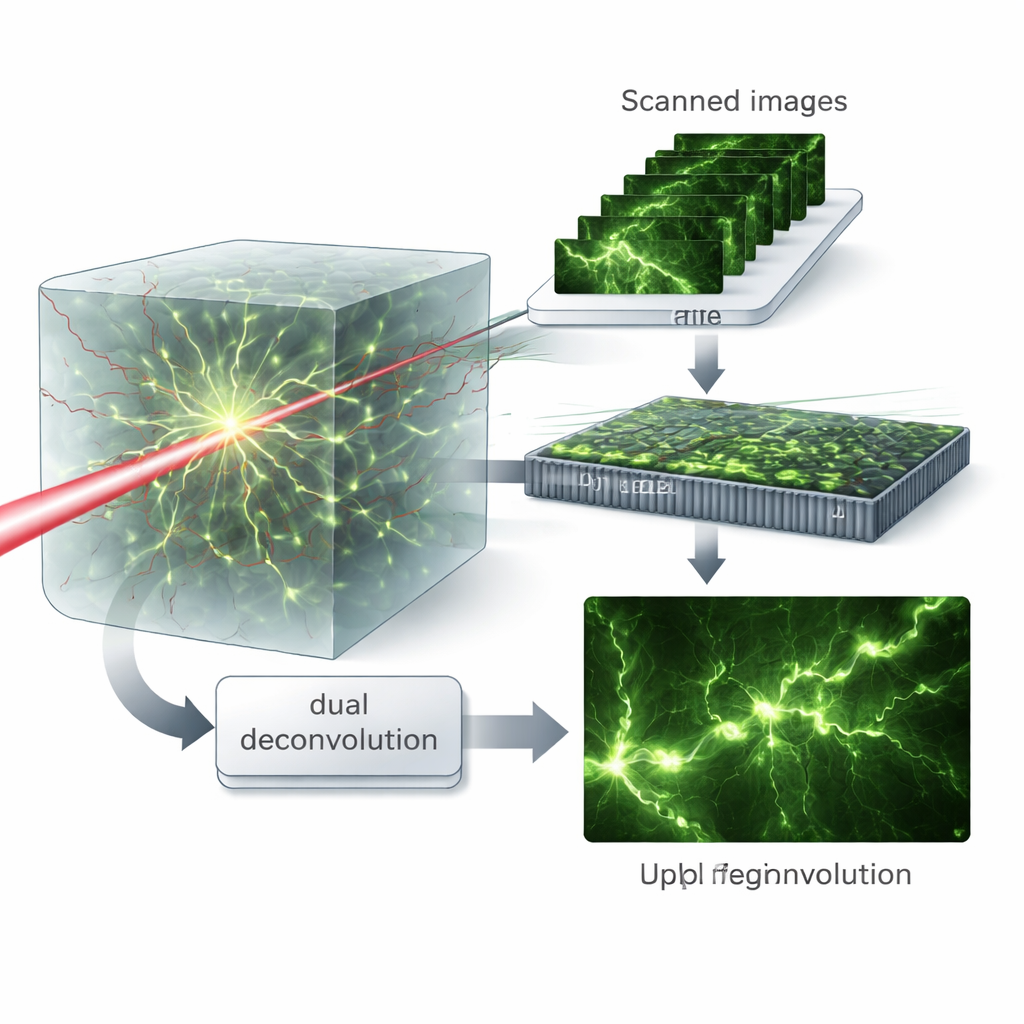
Virtual Patterns and Dual Cleanup
The key insight is to treat the scanned images as if the sample had been illuminated with many different fine patterns of light, a concept the authors call virtual structured illumination. Mathematically recombining these data in the frequency domain lets them separate the roles of the excitation and emission processes. They then introduce a “dual deconvolution” algorithm that alternately estimates and corrects the blurring from each side—the light going in and the light coming out—rather than lumping them together into a single effective blur. This matrix-based treatment preserves more high‑frequency detail and allows the algorithm to recover fine structures even when aberrations are strong.
Sharper Views in Simulations and Real Samples
To test their method, the team first used computer simulations of two-photon microscopy, a deep-imaging technique that uses pairs of lower‑energy photons to excite fluorescence only at the focus point. Under severe simulated distortions, conventional two-photon and standard structured illumination reconstructions produced noticeably blurred images. In contrast, dual deconvolution restored crisp patterns whose resolution approached one-quarter of the fluorescence wavelength—about 130 nanometers—matching theoretical expectations. The authors then built a custom two‑photon setup with a scientific camera and applied their algorithm to real samples, including fluorescent beads, test patterns hidden behind scattering layers, cultured cells, mouse brain tissue, and whole-mount zebrafish. Again and again, structures that appeared smeared or doubled in conventional images emerged as clearly separated features after processing, and fine neuronal elements such as dendritic spines remained resolvable at depths up to 180 micrometers in mouse brain.
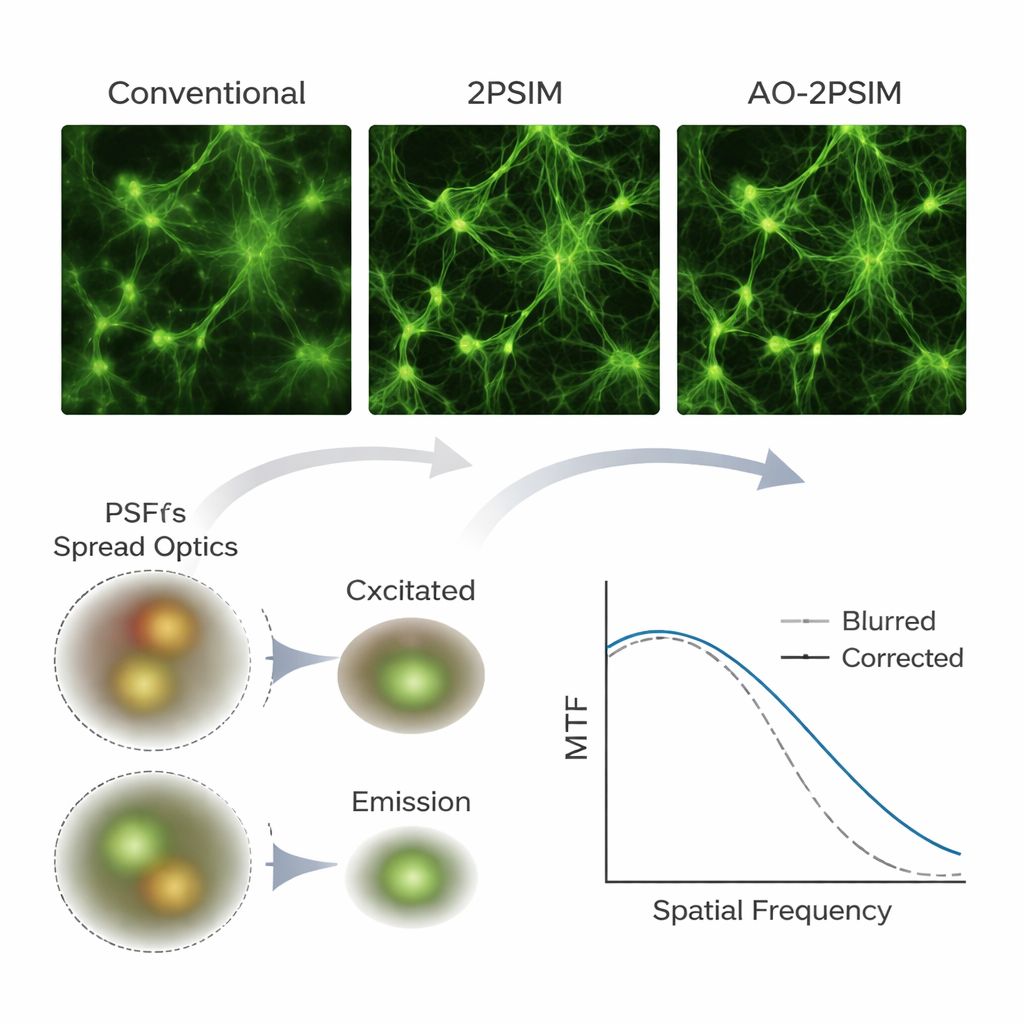
What This Means for Biological Imaging
For non-specialists, the main message is that the authors have shown how to turn existing multiphoton microscopes into much more powerful deep‑tissue imaging tools using primarily software and a camera upgrade. By carefully modeling and correcting how tissues bend light on both the way in and the way out, their dual deconvolution approach doubles the usual resolution limit in challenging, thick samples without relying on complex adaptive mirrors. While the method still depends on collecting enough signal and currently requires relatively slow scanning, it offers a practical, cost‑effective path toward routine, ultra‑sharp views of the brain and other organs in three dimensions, opening the door to more detailed studies of how biological structures are organized and change over time.
Citation: Lim, S., Kang, S., Hong, J.H. et al. Dual deconvolution in multiphoton structured illumination microscopy for deep-tissue super-resolution imaging. Nat Commun 17, 2123 (2026). https://doi.org/10.1038/s41467-026-69798-y
Keywords: super-resolution microscopy, two-photon imaging, adaptive optics, deep tissue imaging, computational imaging