Clear Sky Science · en
Euglena gracilis as a high-throughput screening platform for antibacterial activity, cytotoxicity and membrane permeability in a one-step and cost-effective assay
Why A Tiny Green Cell Matters
Deadly bacterial infections are becoming harder to treat as more germs outsmart our existing antibiotics. Yet finding new drugs is slow, expensive, and often frustrating. This study introduces an unexpectedly simple helper: Euglena gracilis, a common pond-dwelling microbe. By watching how this tiny green cell changes color, researchers built a one-step test that can quickly flag promising antibiotic candidates, weed out toxic ones, and check whether they can slip through multiple biological barriers—all in the same affordable assay.
A Simple Color Change With Big Implications
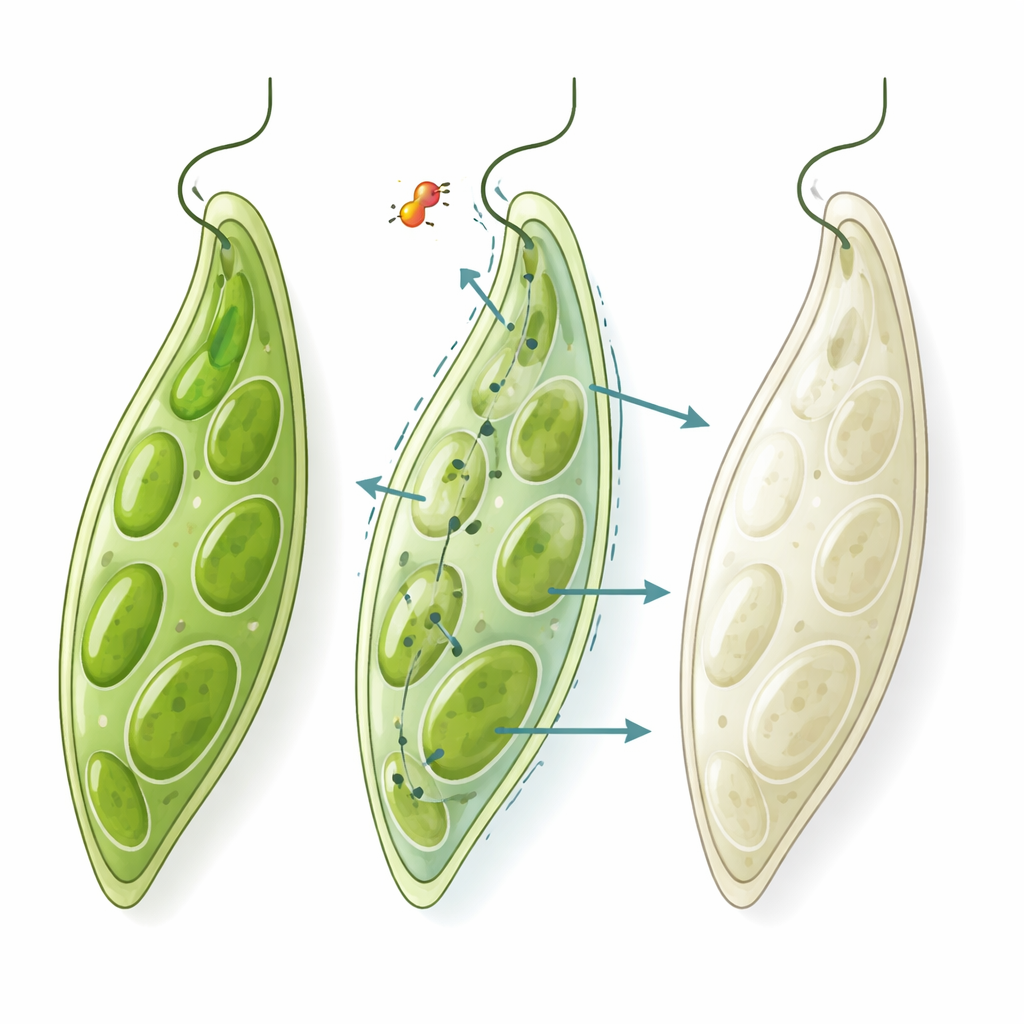
Euglena looks like a small green slipper under the microscope. Its color comes from chloroplasts, the same kind of light-harvesting structures found in plant cells. Long ago, those chloroplasts originated from bacteria, and they still retain some bacterial traits. That quirk makes them vulnerable to several antibiotics, which can damage or remove the chloroplasts and turn Euglena from green to white—a process known as bleaching. At the same time, Euglena is a flexible feeder: it can live on dissolved nutrients even when it loses its chloroplasts. This means that if a compound bleaches Euglena but the cells stay alive and keep dividing, it is likely hitting an antibacterial target without being broadly poisonous to complex cells like ours.
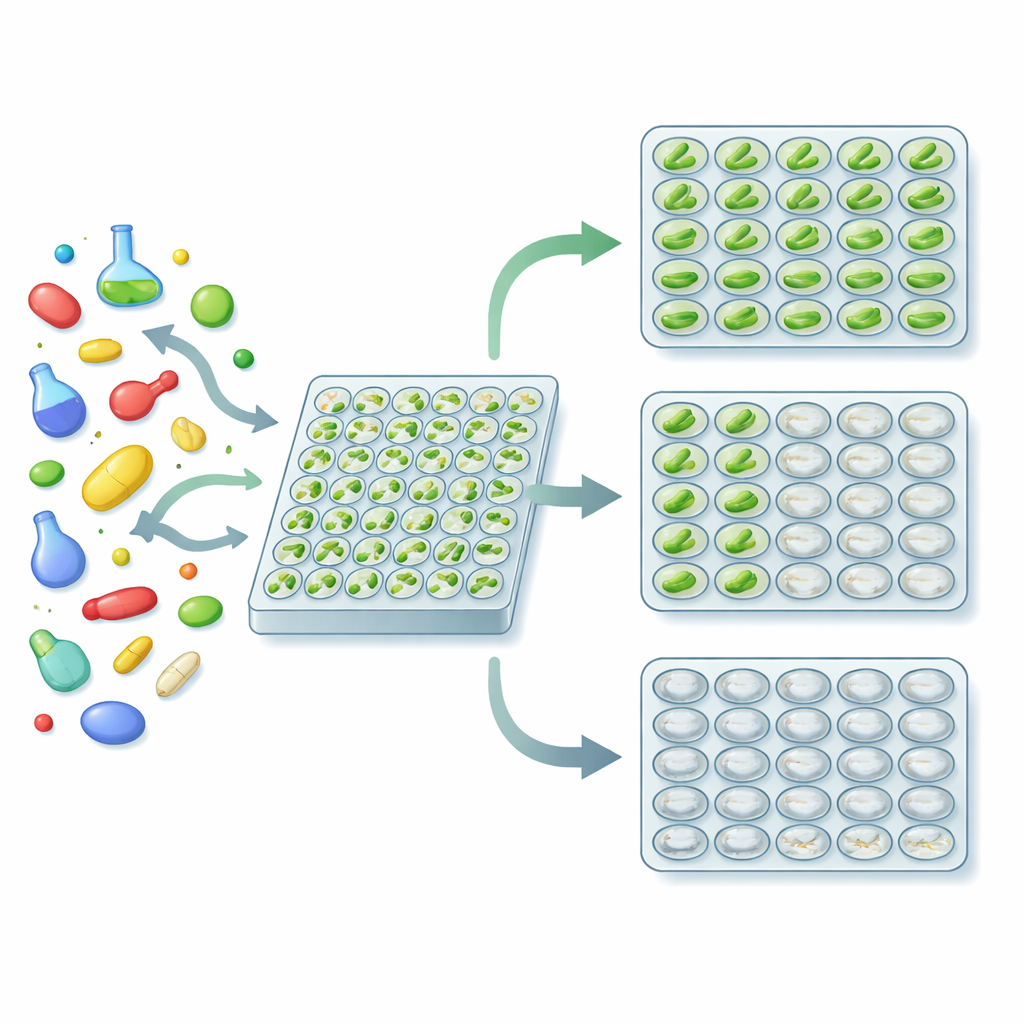
Three Outcomes In One Easy Test
The authors turned this biology into a practical high-throughput screen. They grew Euglena in standard 96-well plates, added a different test compound to each well, and waited a few days. At the end, each well reliably fell into one of three visible states: green (the compound did nothing important), white but cloudy (the cells survived but lost chloroplasts, hinting at antibacterial activity), or clear (the cells were killed, suggesting general toxicity). Using a plate reader that measures how much light the wells absorb at different wavelengths, the team defined numerical cutoffs that automatically distinguish these states. A particular ratio between light absorbed in the red and green regions of the spectrum reports bleaching, while the overall signal strength reports whether the culture grew at all.
Putting The Screen To The Test
To see whether this approach worked beyond theory, the researchers first challenged Euglena with a commercial library of 79 well-characterized compounds. Their color-based system cleanly separated drugs that were known to kill cells from those that specifically interfere with bacterial-style processes. Eight compounds triggered bleaching without wiping out the cultures. Most of these were classic antibiotics that block protein or DNA production in bacteria, spanning several distinct chemical families. Importantly, not all drugs from a given class bleached Euglena, revealing that the chloroplast’s bacterial heritage overlaps only partly with modern bacteria. A few compounds yielded unusual light-absorption patterns because they themselves were colored, reminding users that strongly tinted molecules may require manual checking of the results.
Discovering Signals In Natural Products
The team next applied the assay to 88 rare natural products isolated from myxobacteria and fungi, organisms known to make chemically exotic molecules. Eighteen compounds stopped Euglena from growing, consistent with known actions on energy production, cell scaffolding, or cellular recycling machinery. Strikingly, one compound—argyrin C, a cyclic peptide from myxobacteria—produced strong bleaching rather than outright killing. Argyrin C is already known to block a protein “elongation factor” involved in protein synthesis inside mitochondria, the cell’s power stations that also trace their ancestry to bacteria. Its bleaching effect in Euglena confirms that the assay can spot antibacterial-style mechanisms that go beyond the usual ribosome- and DNA-targeting drugs.
Promise And Limits Of The Approach
This color-based screen will miss some important antibiotic types. For example, drugs that attack bacterial cell walls have nothing to latch onto in chloroplasts and therefore leave Euglena green. Even among compounds that block protein production, only about 40% caused bleaching, underscoring the evolutionary distance between chloroplasts and modern pathogens. On the other hand, the method is exceptionally cheap, robust, and easy to scale. Euglena grows in simple salt-and-ethanol medium without animal serum, can be maintained for weeks with minimal care, and requires only basic lab equipment. Because its chloroplast derives from a Gram-negative ancestor, the assay is especially well suited to finding drugs that can penetrate the multiple membranes typical of some of today’s most worrisome resistant bacteria.
What This Means For Future Antibiotics
To a lay observer, this work shows how watching a single-celled organism change from green to white can help guide the search for tomorrow’s antibiotics. The Euglena assay is not a stand-alone answer to drug resistance, but a powerful early filter: in one step it highlights molecules that look antibacterial, seem relatively safe for complex cells, and can cross several membrane barriers. Those hits can then move on to deeper testing directly in disease-causing bacteria and animal cells. As multi-drug-resistant infections continue to rise worldwide, such smart, low-cost screening tools may speed discovery by letting scientists focus scarce resources on the most promising leads.
Citation: Pereira, L., Löffler, LS., H. Kirsch, S. et al. Euglena gracilis as a high-throughput screening platform for antibacterial activity, cytotoxicity and membrane permeability in a one-step and cost-effective assay. J Antibiot 79, 376–385 (2026). https://doi.org/10.1038/s41429-026-00911-5
Keywords: antibiotic discovery, high-throughput screening, Euglena gracilis, drug resistance, natural products