Clear Sky Science · en
Immunogenicity assessment and epitope mapping of the ASFV proteome by profiling serum antibodies with ASFV antigen phage libraries
Why this matters for farmers and food security
African swine fever is a deadly viral disease of pigs that has wiped out herds and shaken the global pork market, especially in countries with large pig populations like China. Because there is still no safe, effective vaccine, outbreaks are controlled mainly by mass culling, which is devastating for farmers and food prices. This study asks a simple but powerful question: what exactly does a pig’s immune system “see” when it fights this virus, and how can that knowledge be turned into better tests and, eventually, vaccines?
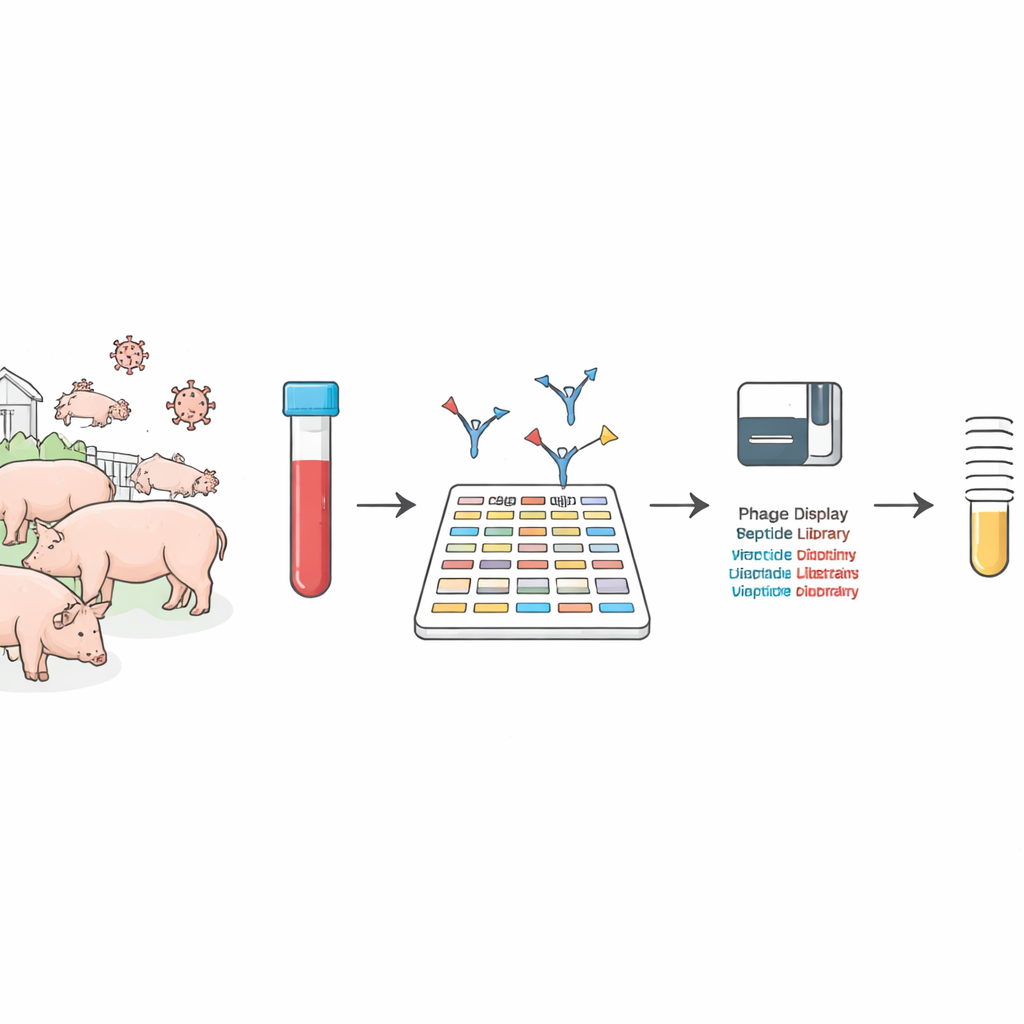
Reading the immune system like a barcode
The researchers used a high‑throughput method called phage display sequencing to take a deep look at antibodies in pig blood. Instead of testing one viral protein at a time, they built two enormous collections of harmless viruses, each one displaying a tiny piece of African swine fever virus or other common pig viruses on its surface. When pig serum is mixed with these collections, antibodies latch onto the pieces they recognize. By sequencing which pieces were grabbed, the team could reconstruct, in one sweep, the entire landscape of what the pig’s immune system was targeting.
Following infection from crisis to recovery
The team analyzed blood from 100 pigs, some never infected and others at different stages of African swine fever: early, mid‑course, and recovered. They found that pigs in the acute, early stage recognized only a few viral pieces, while animals that survived and recovered showed a broad, rich pattern of antibody recognition across many viral proteins. This “spreading” of the response suggests that long‑term control of the virus depends on hitting many targets at once, not just one or two famous proteins. Their maps also confirmed that the method was reliable by matching many known virus regions already cataloged in public databases.
Unearthing a hidden viral target
Beyond confirming well‑studied proteins, the most striking discovery was a previously little‑known viral protein called DP238L. Antibodies against DP238L showed up again and again in pigs that had recovered from infection. When the scientists compared virus sequences from 169 different strains, they found that DP238L is highly conserved—meaning it changes very little as the virus evolves—yet it is also widely recognized by pig antibodies. Structural predictions suggest that most of DP238L is exposed and easy for the immune system to reach, and antibody “heat maps” showed strong responses along much of its length, marking it as a prime candidate for vaccines and diagnostic tests.
Turning epitope maps into real‑world tools
To move from maps on a computer to practical use, the team produced DP238L in bacteria and tested whether real pig sera recognized it. Blood from infected pigs strongly bound to the purified DP238L protein, while blood from healthy pigs did not, confirming that DP238L behaves as a genuine marker of infection. The researchers also stitched together two highly reactive short segments from other viral proteins into a single “multi‑epitope” test protein and showed that infected pig sera recognized this chimera as well. Finally, they immunized pigs with DP238L and observed a strong, long‑lasting antibody response along with balanced immune‑signaling molecules, suggesting that this protein can safely trigger a robust immune reaction.
What this means for tests and vaccines
Seen through a layperson’s lens, this work is like drawing a detailed map of the bull’s‑eyes the virus presents to the pig’s immune system, and then testing which bull’s‑eyes are both stable and easy to hit. The study identifies 29 key viral proteins and pinpoints the most reactive regions within them, with DP238L standing out as a particularly promising target. These insights can guide the design of more sensitive blood tests that rely on several strong, conserved viral pieces rather than just one, making diagnosis more reliable across different virus strains. They also provide a blueprint for building safer, targeted vaccines that aim to mimic the broad, multi‑target immune response seen in surviving pigs—an essential step toward reducing culling, protecting farmers’ livelihoods, and stabilizing the global pork supply.
Citation: Ma, L., Weng, Z., Zhang, Y. et al. Immunogenicity assessment and epitope mapping of the ASFV proteome by profiling serum antibodies with ASFV antigen phage libraries. Commun Biol 9, 448 (2026). https://doi.org/10.1038/s42003-026-09709-5
Keywords: African swine fever, pig immunity, viral epitopes, vaccine design, serological diagnostics