Clear Sky Science · en
JanusDDG: a physics-informed neural network for sequence-based protein stability via two-fronts attention
Why this research matters
Proteins are the tiny machines that keep our cells alive, and even a single change in their building blocks can make them work better, worse, or not at all. Being able to predict how such changes affect a protein’s stability is crucial for understanding genetic diseases and for designing better medicines and industrial enzymes. This paper introduces JanusDDG, a new artificial intelligence model that predicts how mutations alter protein stability using only the protein’s sequence, while also obeying the basic physical rules that govern how proteins fold.
The problem of fragile protein machines
When a protein folds into its three-dimensional shape, it balances many forces, like a tent held up by many ropes. Mutations can tighten some ropes or loosen others, making the structure more or less stable. Experimental tests of these effects are slow and expensive, so researchers rely heavily on computer models to estimate changes in stability, known as ΔΔG. Existing tools often work best when they have access to detailed 3D structures, and they may quietly break the rules of thermodynamics, leading to predictions that look accurate on paper but are physically inconsistent or hard to trust for new proteins.
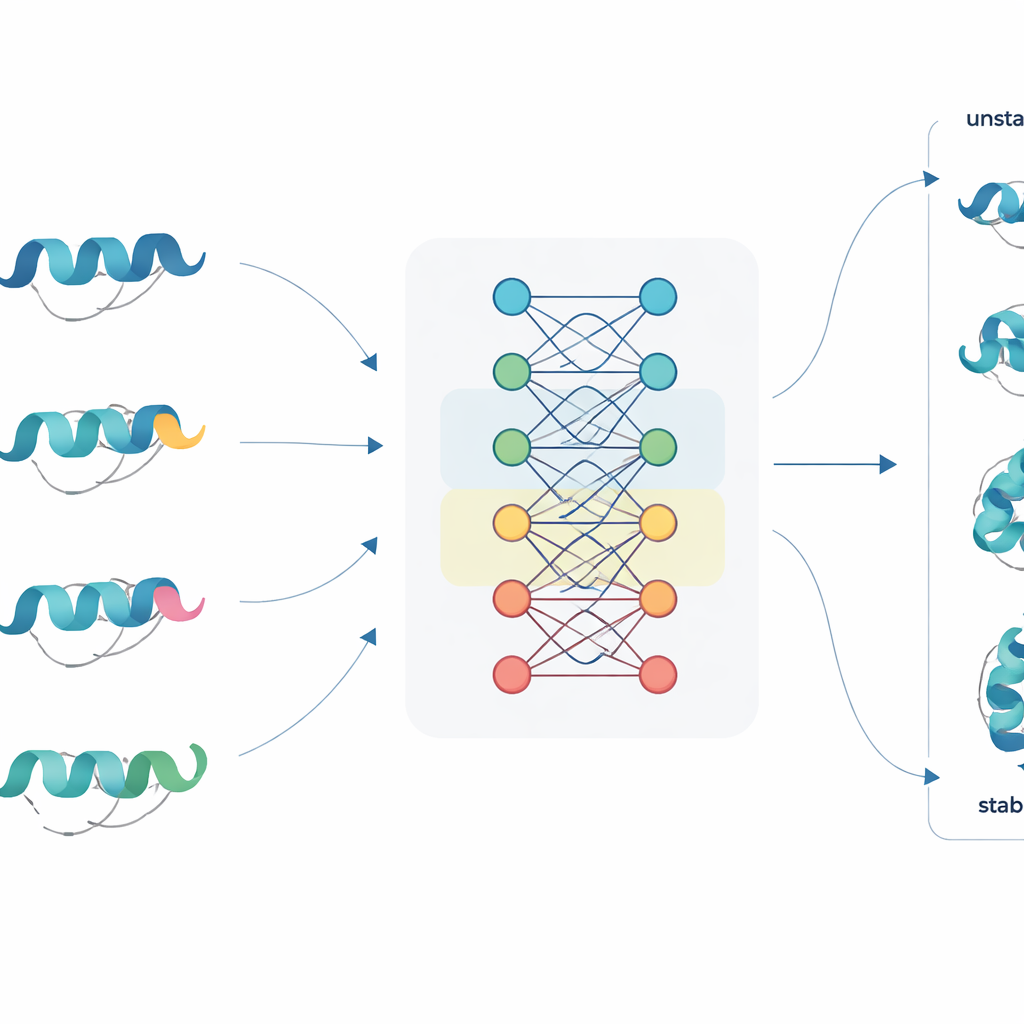
A new way to read protein sequences
JanusDDG tackles this challenge by starting from protein language models, a class of large neural networks trained on millions of protein sequences, somewhat like how language models learn from text. These models convert each amino acid into a rich numerical representation that captures patterns from evolution and typical folding behavior. JanusDDG takes the sequence of the original protein and the sequence of its mutant, compares their learned representations, and uses a specialized attention mechanism that focuses on how the mutation perturbs the surrounding context. Because it only needs sequences, JanusDDG can be applied to proteins whose 3D structures are unknown or difficult to determine.
Building physics into artificial intelligence
A key innovation of JanusDDG is that it is designed to respect fundamental physical principles. The authors focus on two properties of Gibbs free energy, the quantity that underlies protein stability. First, antisymmetry means that if going from one variant to another changes stability by a certain amount, the reverse change must undo that effect. Second, transitivity means that the total effect of going from one variant to a second, then to a third, must equal the direct jump from the first to the third. JanusDDG’s architecture enforces antisymmetry by running two mirrored copies of the network on swapped inputs and combining their outputs so that forward and reverse predictions are exact opposites. Transitivity is encouraged during training by adding a special loss term that pushes the model to make consistent predictions when mutational paths are broken into steps.
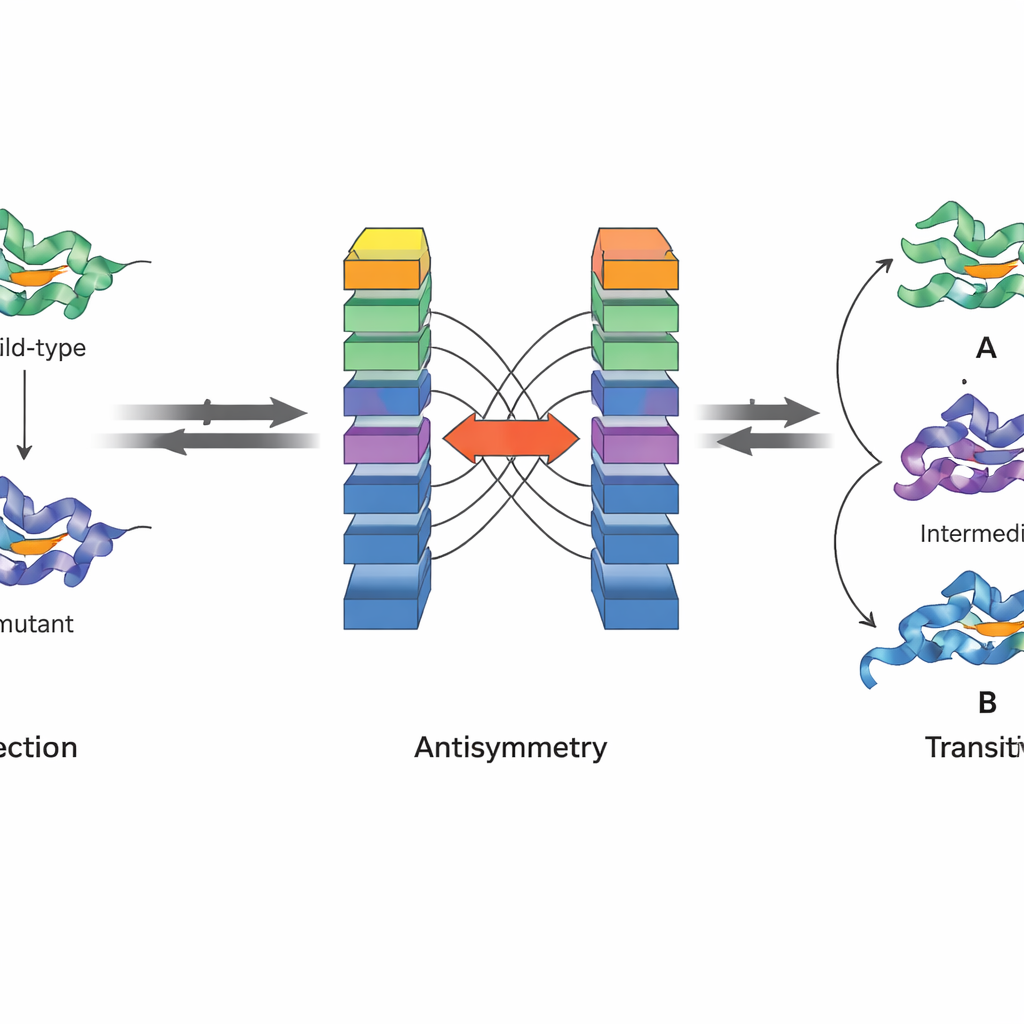
Testing performance on many kinds of mutations
The researchers trained JanusDDG on a curated dataset of thousands of mutations with measured stability changes and then tested it on several independent benchmarks where sequence overlap with the training data was kept very low. This careful design reduces the risk that the model is simply memorizing familiar proteins. Across three widely used collections of single mutations, JanusDDG matched or outperformed both other sequence-based tools and many methods that rely on 3D structures. It also handled multiple simultaneous mutations, a harder scenario where interactions between changes can be non-additive. Remarkably, its accuracy did not drop for pairs of mutations that are close together in space, where earlier models often struggle.
From numbers to useful stability labels
In practical applications, researchers often want to know not just how big a stability change is, but whether a mutation is clearly stabilizing or destabilizing. The authors tested JanusDDG on a dataset focused on distinguishing stabilizing from destabilizing variants. While the model reached solid performance, this task remained more difficult than predicting raw numerical values, especially near the boundary between categories where experimental noise and biological ambiguity are highest. Still, JanusDDG compared favorably with other top methods, suggesting that its physics-aware design and use of rich sequence embeddings help it navigate this uncertainty better than many competitors.
What this means for future protein design
Overall, JanusDDG shows that it is possible to combine the strengths of modern sequence-based AI with the firm constraints of physical law. By treating proteins as sequences that can be read like language, yet insisting that predictions obey antisymmetry and transitivity, the model produces stability estimates that are both accurate and thermodynamically consistent. For non-specialists, the takeaway is that we are getting closer to reliable, structure-free tools that can scan through countless possible mutations, highlighting those most likely to stabilize a protein or flagging risky changes linked to disease, all while staying grounded in the rules of physics rather than mere statistical shortcuts.
Citation: Barducci, G., Rossi, I., Codicé, F. et al. JanusDDG: a physics-informed neural network for sequence-based protein stability via two-fronts attention. Commun Biol 9, 494 (2026). https://doi.org/10.1038/s42003-026-09632-9
Keywords: protein stability, genetic mutations, protein design, machine learning, thermodynamics