Clear Sky Science · en
Personalized signaling pathway analysis of gastrointestinal tumors for patient stratification and drug target evaluation using clinically derived core biopsies
Why this research matters to patients
Cancer doctors are increasingly trying to match treatments to the unique biology of each person’s tumor. For cancers of the digestive system—such as those of the pancreas, colon, liver, or bile ducts—this is especially urgent, because they are common, often diagnosed late, and difficult to treat. This study explores a new lab method that can read the activity of many cancer-related proteins from tiny biopsy samples, with the goal of helping doctors choose more precise therapies for individual patients.
From DNA lists to living signals
Today, most “personalized” cancer care relies on reading DNA changes in a tumor. While powerful, DNA alone does not show which signals inside the cell are actually switched on and driving growth. Those signals are carried by proteins, many of which can be directly targeted by drugs. The researchers used a high‑throughput technique called DigiWest, a modern spin on the classic Western blot, to measure around 130–200 proteins and their activated forms at once. Importantly, this method needs only about as much material as a single needle biopsy, making it suitable for real‑world clinical use.
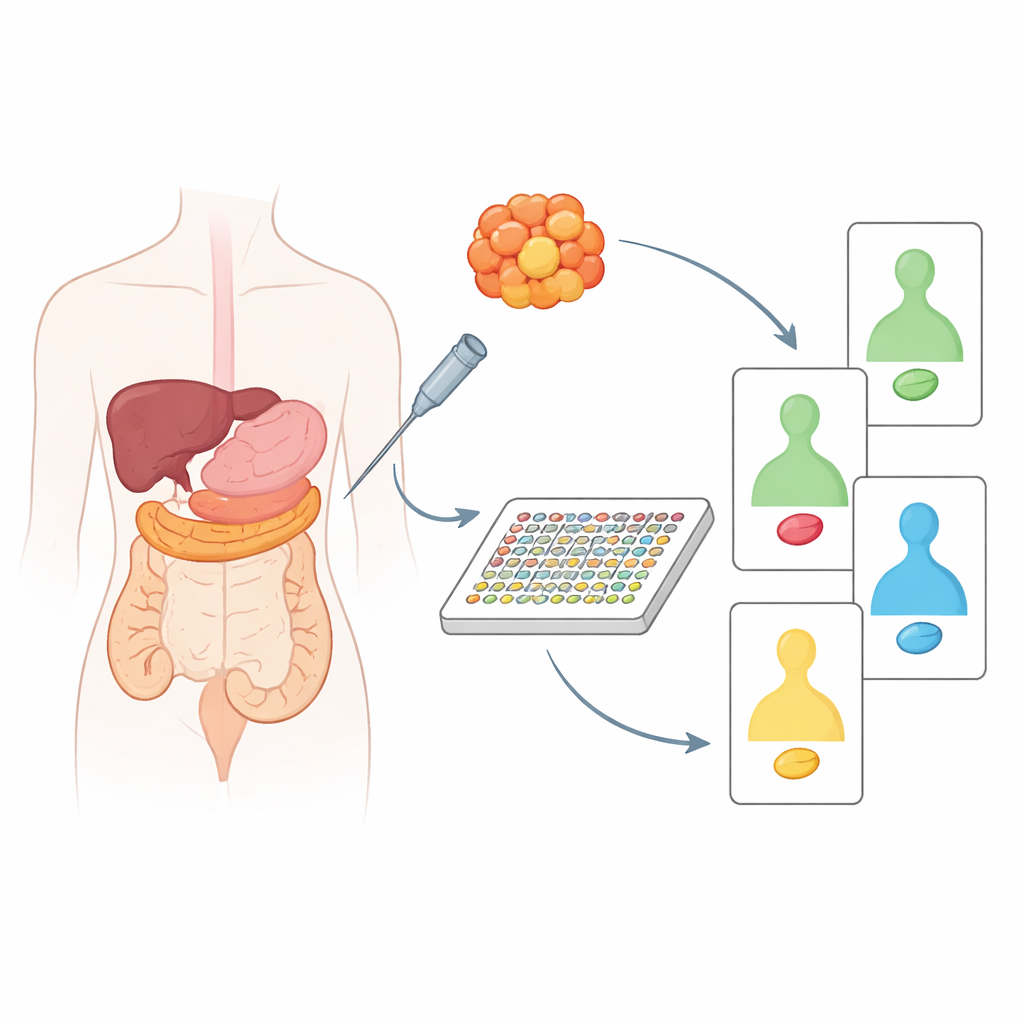
Comparing tumor tissue to its healthy neighbor
The team first analyzed stored tissue from 20 patients with pancreatic or colorectal cancer, always paired with nearby non‑cancerous tissue from the same person. By comparing tumors to their own healthy counterparts, they could see which proteins were truly altered by cancer rather than just reflecting normal differences between organs. This revealed clear contrasts in the behavior of well‑known cancer guardians and drivers, such as p53, Ras, PTEN, and others. Pancreatic tumors, for example, tended to show boosted growth‑promoting signals and loss of protective proteins, whereas colon tumors had their own distinct pattern of disrupted pathways. When the researchers clustered the samples based on these protein changes, they could divide pancreatic cancers into two biologically different groups and detect meaningful differences among colon cancers related to patient age and tumor location in the bowel.
Individual tumor “barcodes” of signaling activity
Looking beyond group averages, the scientists built a detailed protein profile for each tumor. These profiles highlighted which signaling routes—such as those involving mTOR, MAPK/Erk, Wnt, or immune‑related factors—were especially active or shut down. Many of the measured proteins are the direct targets of existing drugs, or sit just downstream of such targets, allowing the team to infer which medicines might interfere with a tumor’s main growth engines. In three quarters of the retrospective cases, they could point to one or more pathways that likely fueled tumor progression. They also picked out tumors packed with immune‑cell markers, suggesting “hot” cancers that might respond to immunotherapy, and unusual cases with strikingly unique signatures.
Putting the method to work in real patients
To test its usefulness at the bedside, the researchers then applied DigiWest to fresh needle biopsies from 14 patients with various gastrointestinal cancers whose cases were being reviewed by a Molecular Tumor Board. These patients had complex disease histories and, often, prior treatments. Because matching healthy tissue was not available, each tumor’s protein levels were compared to the median level across the group to define what counted as abnormally high or low. Even with this stricter setup, 12 of the 14 tumors showed clear, treatment‑relevant patterns of pathway activity. In two detailed examples, the protein data confirmed a DNA‑level amplification of the FGFR2 gene in a colon cancer and loss of an mTOR brake in a liver cancer, strongly supporting the Board’s consideration of FGFR‑blocking or mTOR‑blocking drugs. Overall, DigiWest findings agreed with key genetic drivers and suggested drug targets in most evaluable cases.
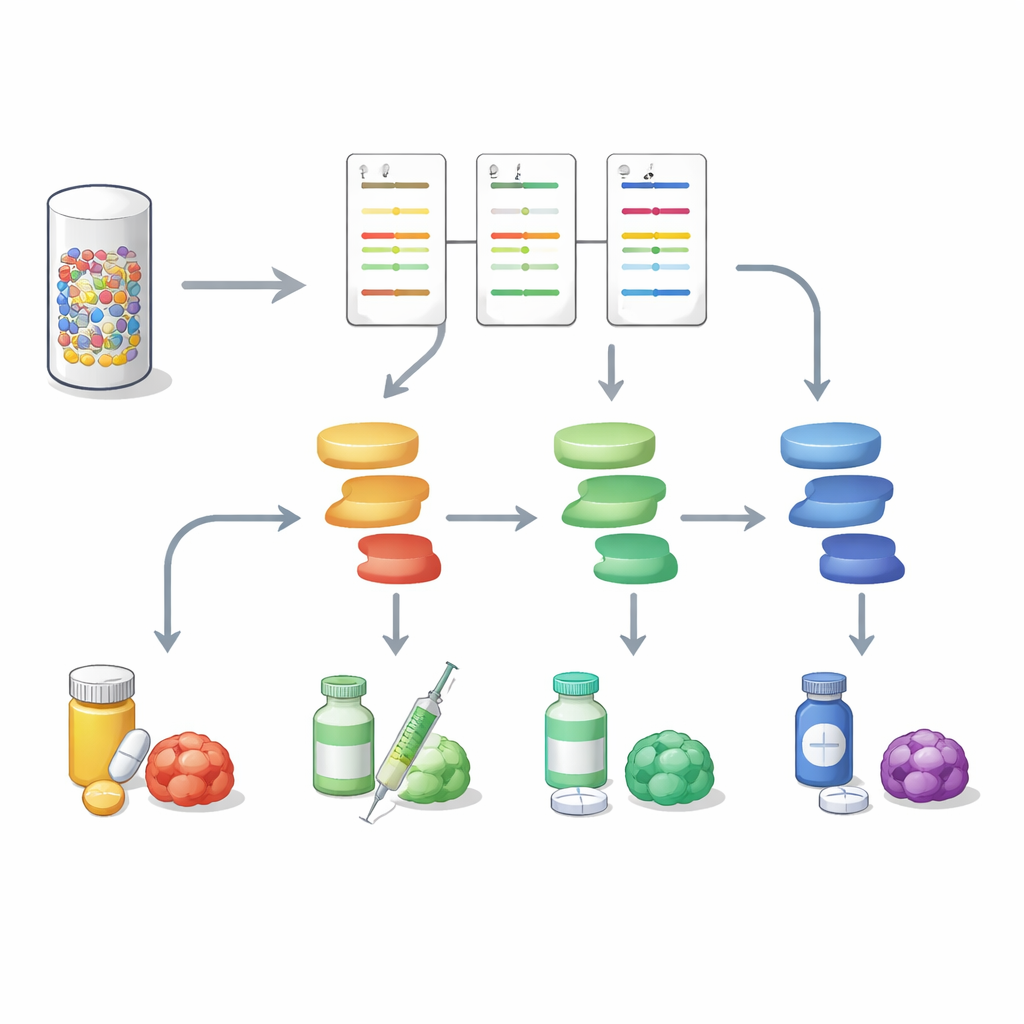
Toward more complete tumor portraits
This work shows that analyzing protein signaling in parallel with DNA sequencing can give a richer and more actionable picture of gastrointestinal tumors. By turning a tiny biopsy into a multi‑pathway activity map, DigiWest helps distinguish which molecular switches are truly on in a given cancer and which drugs might best hit them, and may also flag emerging resistance routes. While larger studies are still needed, the approach offers a practical way to bring high‑content protein profiling into everyday precision oncology and move closer to treatment plans tailored to each patient’s living tumor, not just its genetic blueprint.
Citation: Stahl, A., Büringer, K., Missios, P. et al. Personalized signaling pathway analysis of gastrointestinal tumors for patient stratification and drug target evaluation using clinically derived core biopsies. npj Precis. Onc. 10, 124 (2026). https://doi.org/10.1038/s41698-026-01304-5
Keywords: precision oncology, gastrointestinal cancer, proteomics, biopsy profiling, targeted therapy