Clear Sky Science · en
Large-area 3D reconstruction of corneal tissues from oscillating focus confocal microscopy
Seeing the Eye in Three Dimensions
The clear window at the front of your eye, the cornea, is packed with delicate nerves and immune cells that can reveal early signs of diseases such as diabetes, multiple sclerosis, or even long COVID. Doctors already use a special type of microscope placed gently on the eye to view these structures, but they mostly see them as flat slices. This study presents a new way to turn thousands of such slices into a detailed three-dimensional map over a wide area of the cornea, opening the door to tracking tiny cells and nerve changes in both health and disease.
Why Flat Pictures Are Not Enough
Standard in vivo confocal microscopy produces very sharp images of the cornea, but each picture captures only a small patch and a single depth. Doctors often stitch many of these pictures together into wider two-dimensional mosaics to examine the corneal nerve network or count immune cells. However, both nerves and immune cells are truly three-dimensional: they branch, bend, and move not only sideways but also deeper or closer to the surface. With current methods, movement in depth is largely invisible. Existing three-dimensional reconstructions are limited to very small areas roughly the size of a single image, which is too small for reliably tracking many cells or for capturing the full variation of the nerve layer.
Scanning a Wide, Deep Region of the Cornea
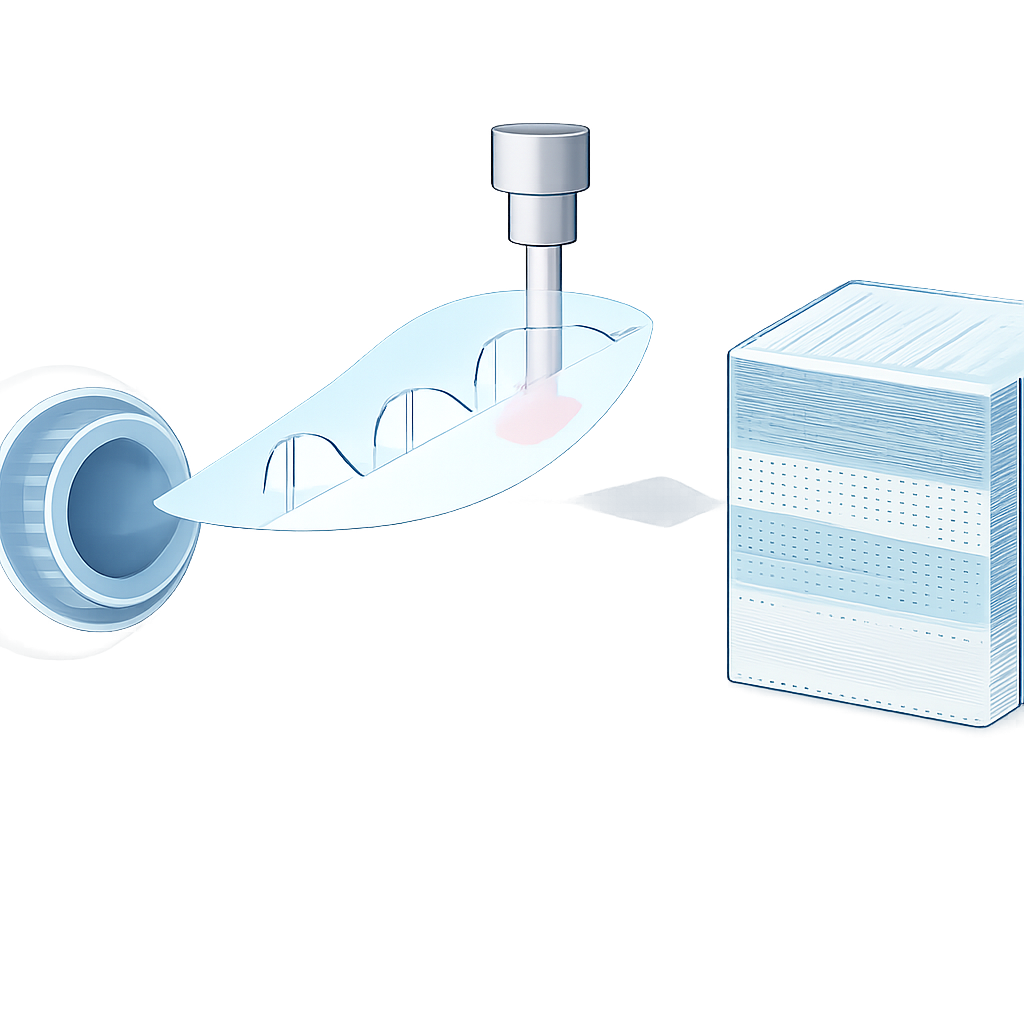
The research team builds on a customized confocal microscope system that gently touches the eye with a capped tip, while a second device guides the patient’s gaze along a planned path. As the eye slowly moves, the microscope sweeps over a large area of the cornea. At the same time, the focus of the microscope is continuously driven up and down in a triangular pattern through the tissue. This means that, instead of imaging just one flat layer, the system repeatedly scans a stack of depths as it travels across the cornea, covering both the subbasal nerve plexus and neighboring layers. The raw data are thousands of small, partially overlapping images acquired at different positions and depths during this oscillating scan.
Turning Moving Slices into a Solid Volume
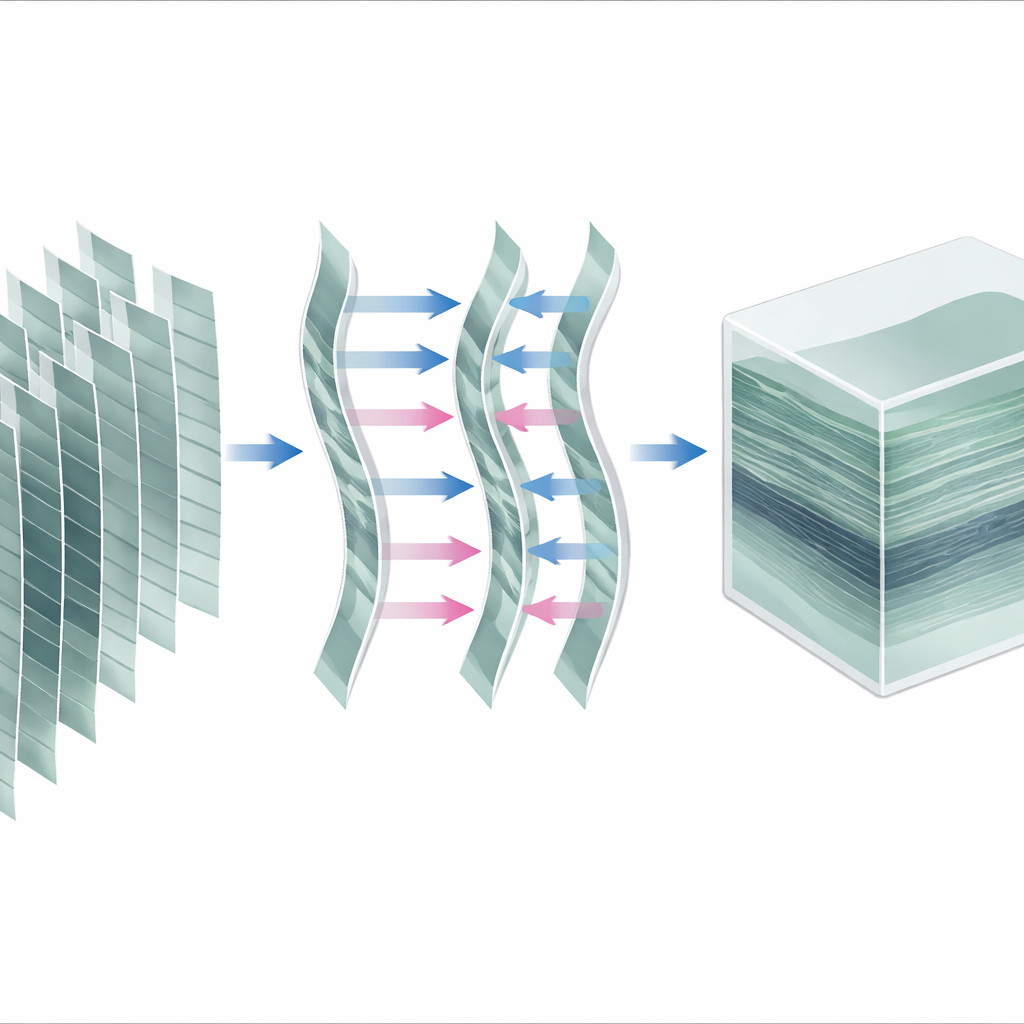
To transform these images into a clean three-dimensional block of tissue, the authors design a step-by-step processing pipeline. First, they correct side-to-side eye movements, making sure every image is placed on a common grid, as if the eye had held perfectly still. Then they divide the continuous scan into many short focus “stacks,” each spanning one pass of the focus from front to back or vice versa. For each stack, they assign a depth value to every image based on the logged focus position and interpolate between slices to fill a regular three-dimensional grid of tiny cubes, or voxels. Finally, they average together all stacks that overlap in space to fuse them into one combined volume of the scanned region.
Correcting for Subtle Motion in Depth
Because the eye and cornea are soft and breathing and other small motions can occur, the tissue may shift slightly toward or away from the microscope during the scan. To handle this, the authors introduce two increasingly sophisticated refinement methods. The simpler method treats each focus stack as a rigid block and aligns stacks along the depth axis using three-dimensional image registration, solving a system of equations to place every stack at its most consistent depth. The most advanced method goes further, breaking stacks into smaller partial stacks and estimating how the tissue’s depth changes smoothly over time. This allows them to compensate stretching or compression effects within stacks, effectively reconstructing an axial motion trajectory and correcting each image individually before the final fusion.
How Well Does It Work and Why It Matters
The team tested all three reconstruction variants on datasets from 15 people with different forms of dry eye disease. Visual inspection of cross-sections through the reconstructed volumes showed similarly good quality across methods, suggesting that, in this particular setup, large depth errors were already limited. However, a specialized, objective image quality measure designed for confocal images detected small but statistically significant improvements when depth motion correction was applied, especially with the most advanced method. Although this higher-precision correction requires substantially more computation time, the authors recommend it for projects that will perform automated analyses or detailed measurements on the volumes.
From Research Tool to Future Clinical Uses
In plain terms, this work shows how to turn a fast, sweeping eye scan into a stable, three-dimensional map of a broad corneal region. For applications like tracking immune cells as they migrate near the nerve layer, or characterizing how the corneal nerve network changes over time or with treatment, this combination of wide coverage and full depth is crucial. The same workflow can be adjusted to focus on different corneal layers, and it may become essential for future versions of the microscope that do not touch the eye at all, where depth motion is more pronounced. Ultimately, such detailed 3D reconstructions could help doctors detect nerve damage earlier, monitor responses to therapies more precisely, and gain deeper insight into how the eye’s surface reflects diseases elsewhere in the body.
Citation: Allgeier, S., Bohn, S., Mikut, R. et al. Large-area 3D reconstruction of corneal tissues from oscillating focus confocal microscopy. Sci Rep 16, 12693 (2026). https://doi.org/10.1038/s41598-026-48735-5
Keywords: corneal imaging, confocal microscopy, 3D reconstruction, corneal nerves, immune cells