Clear Sky Science · en
Enhancing biomedical signals through genetic algorithm optimized Exponentiated transmuted weibull denoising techniques
Why cleaning up medical signals matters
Every heartbeat on an electrocardiogram and every ripple in a brainwave trace can hold clues to serious disease. Yet in real hospitals and clinics, these signals are often buried under layers of unwanted noise from muscle twitches, blinking, electrical interference, and imperfect sensors. This paper presents a new way to clean up those messy signals so that doctors and computer algorithms can see the important patterns more clearly—without needing huge training datasets or supercomputers.
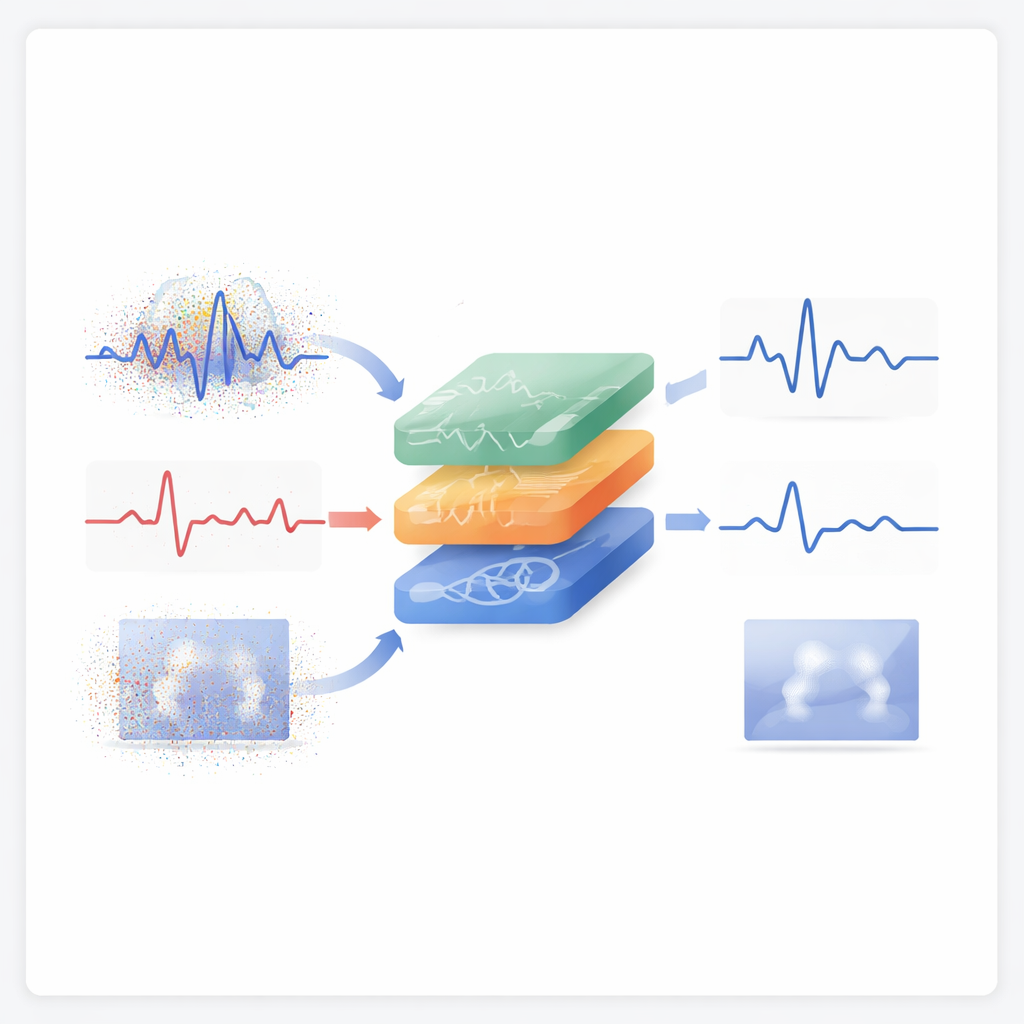
The problem with messy heartbeats and brainwaves
Electroencephalograms (EEG), electrocardiograms (ECG), and medical images like MRI and X‑ray scans are central to diagnosing epilepsy, heart rhythm problems, and structural abnormalities. But these recordings are easily distorted. Eye movements can mask epileptic spikes, body motion can hide key ECG features, and grainy noise can blur small details in images. Traditional cleaning methods—such as simple filters, wavelets, or popular blind source separation techniques—often assume that noise behaves in a neat, symmetric way. In reality, clinical noise is irregular, skewed, and sometimes dominated by rare, extreme bursts. When methods rely on oversimplified noise models, they either fail to remove enough interference or accidentally smooth away the very details clinicians care about.
A flexible way to describe difficult noise
The authors tackle this by introducing a highly adaptable noise model called the Exponentiated Transmuted Weibull Distribution, or ETWD. Instead of assuming a single fixed shape for noise, ETWD uses three main knobs to reshape its curve, allowing it to mimic a wide family of well‑known distributions and to match both heavy‑tailed and asymmetric patterns. In practice, this means the same mathematical framework can describe eye-blink artifacts in EEG, motion spikes in ECG, and grainy speckle in medical images. The team goes further by deriving a special “score function” from ETWD and embedding it inside a fast signal separation algorithm (FastICA). That score function guides the algorithm toward the most plausible hidden sources, unmixing signal and noise in a way that adapts to the true statistics of the data.
Letting evolution tune the cleaner
Because ETWD is so flexible, choosing its parameters by hand would be like trying to tune a complex musical instrument by ear in a crowded room. To automate this, the researchers use a genetic algorithm—a search procedure inspired by natural selection. Candidate parameter sets are treated like individuals in a population; the best performers, judged by how well they fit the data, are gradually refined over generations. This evolutionary step avoids getting stuck in poor solutions and does not require computing complicated derivatives. The same optimization also adjusts sparsity rules in the wavelet domain, which selectively shrink small, noise-dominated coefficients while preserving large, structure-rich ones. Together, ETWD, FastICA, genetic search, and sparsity form a single denoising pipeline that is training‑free yet strongly data‑driven.
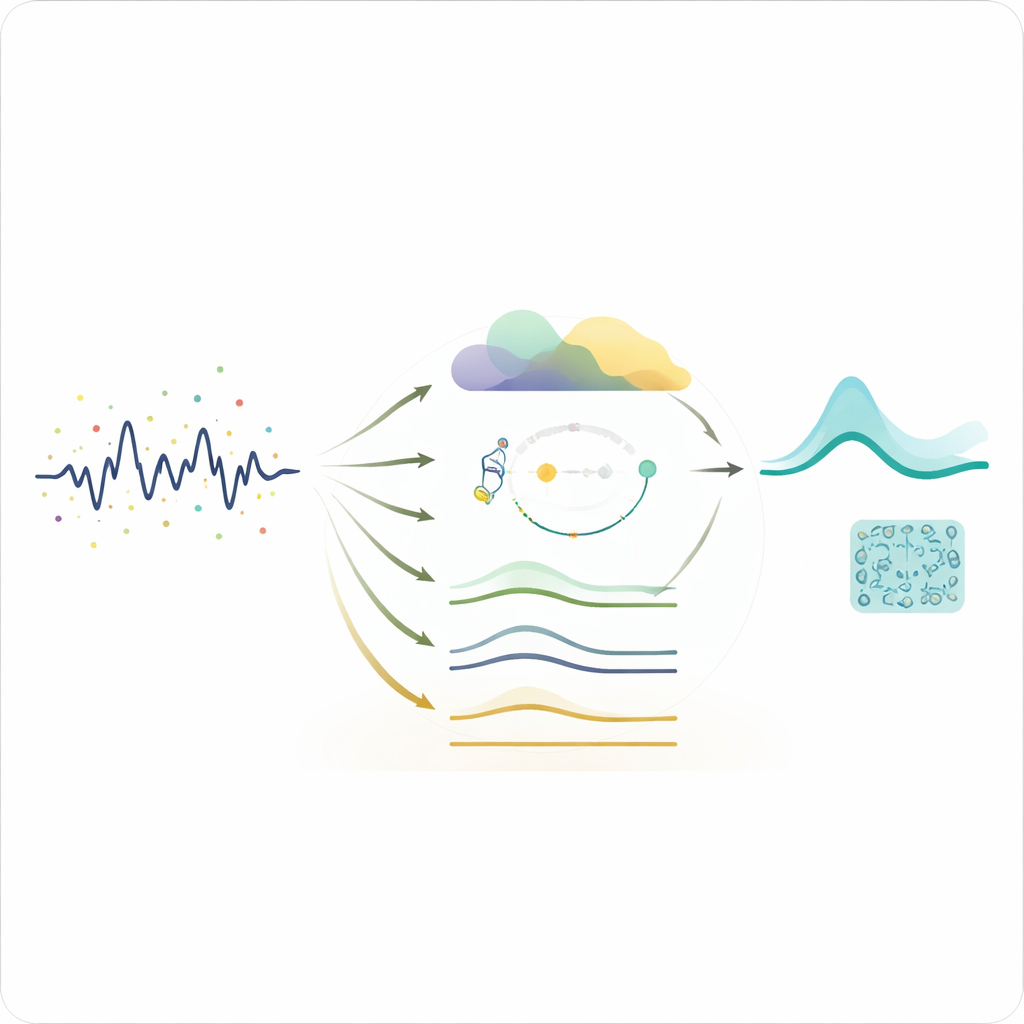

Proving the method on brain, heart, and images
To test the approach, the authors apply it to three different types of biomedical data. For EEG, they clean recordings contaminated by muscle and eye artifacts and then run a spike detection routine on the results. Detection accuracy jumps from 71% on the noisy data to 94% after denoising with the sparsity‑enhanced ETWD method, while false alarms fall sharply. For ECG, they evaluate how well standard heartbeat detectors can find the characteristic R peaks. After denoising, the detector’s F1‑score, a balance of hits and misses, rises from 0.89 to 0.97, indicating that crucial heartbeats are both better preserved and easier to detect. In medical images with added Gaussian, speckle, or Rician noise, the method consistently improves peak signal‑to‑noise ratio compared with several standard filters, recovering subtle anatomical details more faithfully and in less time.
Standing up to modern deep learning
Deep neural networks have become popular for denoising, but they require large labeled datasets and significant training time, and they sometimes oversmooth away fine structure. The authors compare their framework with convolutional autoencoders trained on the same EEG, ECG, and image data. Across tasks, the ETWD-based pipeline matches or outperforms these deep models in key quality measures, while avoiding any separate training phase. It also shows good robustness when applied, without retuning, to entirely new datasets acquired with different protocols—performance drops only slightly and still beats standard Gaussian-based baselines, suggesting the method can transfer reasonably well across sites.
What this means for patients and clinicians
In plain terms, this work delivers a smarter noise eraser for medical signals. By using a highly flexible description of noise and letting an evolutionary algorithm automatically tune it, the framework manages to strip away clutter while keeping the tiny spikes, sharp heartbeats, and faint image edges that often carry diagnostic value. The result is clearer EEG traces, cleaner ECG rhythms, and crisper medical images, achieved without massive training collections or black‑box models. If validated broadly in clinical settings, this kind of adaptable, training‑free denoising could make everyday monitoring devices and hospital systems more reliable, helping both humans and algorithms make safer, more confident diagnoses.
Citation: Sayedelahl, M.A., Farouk, R.M. & Adam, A.M. Enhancing biomedical signals through genetic algorithm optimized Exponentiated transmuted weibull denoising techniques. Sci Rep 16, 13409 (2026). https://doi.org/10.1038/s41598-026-46088-7
Keywords: biomedical signal denoising, EEG and ECG noise removal, medical image enhancement, independent component analysis, genetic algorithm optimization