Clear Sky Science · en
Transcriptome and resequencing reveal immune-related genes and molecular markers associated with Aeromonas salmonicida resistance in Salmo trutta fario
Why hardy trout matter to mountain rivers
High mountain rivers on the Tibetan Plateau support unique fish that local communities rely on for food and income. One of these fish, the brown trout Salmo trutta fario, is now farmed in the region but suffers heavy losses from a bacterial gill disease caused by Aeromonas salmonicida. This study asked a practical question with big ecological and economic stakes: can we pinpoint the fish that are naturally better at fighting this infection, and use that genetic know-how to raise hardier stocks for future farms? 
Fish on the roof of the world under pressure
The Yarlung Zangbo River and its tributaries flow across what is sometimes called the roof of the world. Hydropower development and other human activities are increasing stress on these cold, fast rivers, making fish conservation more urgent. Brown trout introduced more than a century ago have become an important aquaculture species, yet outbreaks of bacterial gill disease can wipe out large numbers of fish. Instead of relying only on drugs or vaccines, the authors explored whether the trout themselves hold genetic clues that explain why some survive infections while others die, opening the door to breeding naturally tougher fish.
Watching genes respond to infection
The team first exposed hundreds of trout to the disease-causing bacterium. Based on whether they became sick, died, or stayed healthy, the fish were grouped as susceptible, resistant, or uninfected controls. From a key immune organ called the head kidney, the researchers measured which genes turned up or down in each group using a technique that reads out thousands of RNA messages at once. They found thousands of genes whose activity changed after infection, particularly those involved in immune alarms, cell communication, and responses to stress. Several stood out, including versions of interferon-stimulated genes and switches in two major defense routes known as PI3K and NF kappa B pathways, which together help control how immune cells respond to invaders.
Linking DNA variants to tougher trout
Gene activity alone cannot explain why some trout shrug off disease. To dig deeper, the scientists sequenced most of the DNA from the livers of resistant and susceptible fish and searched for tiny spelling differences called single nucleotide polymorphisms, or SNPs. They then overlaid these DNA variants with the list of infection-sensitive genes. This combined view highlighted 104 SNPs inside immune-related genes that showed clear differences between resistant and susceptible groups across nearly all chromosomes. From these, six SNP sites were chosen as the most promising markers, many in or near genes tied to the PI3K and NF kappa B control systems. 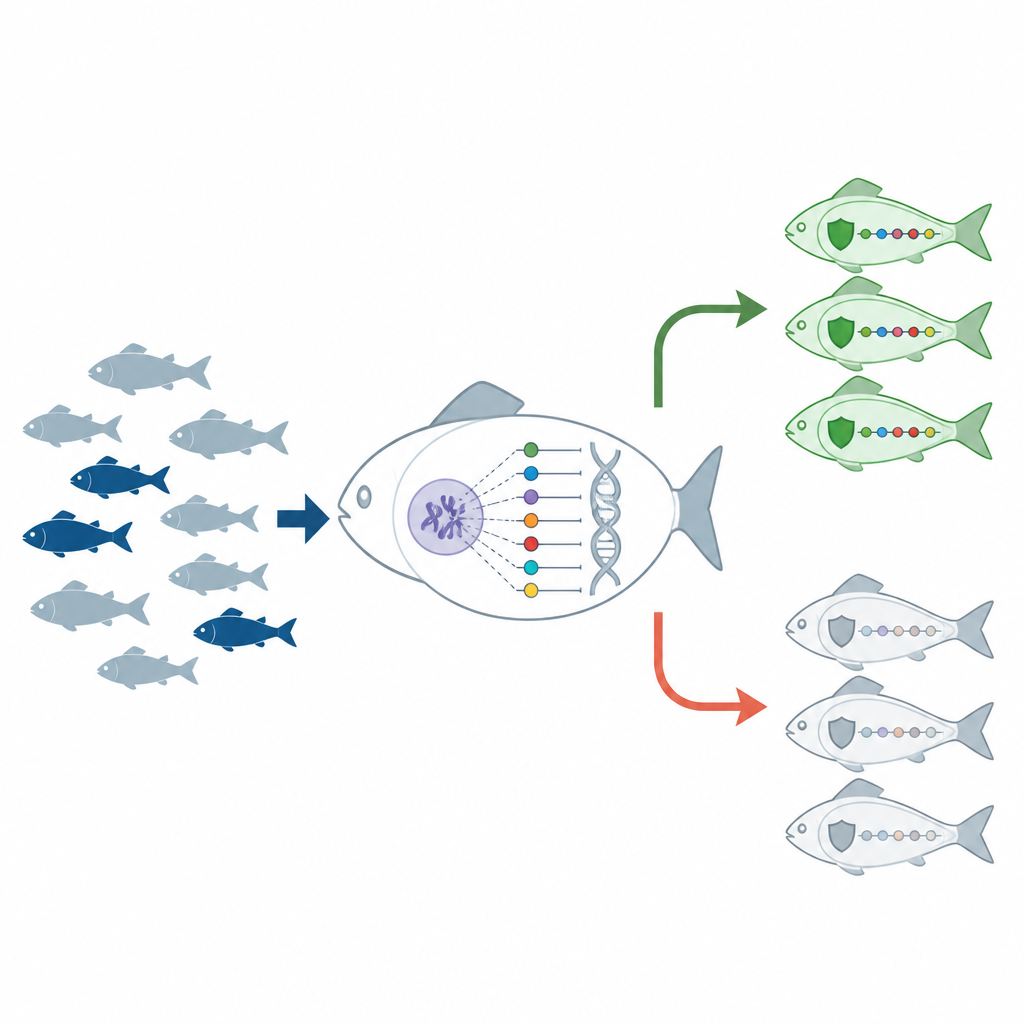
Testing a genetic screening tool
To see if these DNA markers could actually sort fish by their real-world toughness, the team challenged a new batch of 360 trout with the bacterium. After the outbreak, they used a laboratory method called multiplex PCR to read all six SNPs at once from a small fin clip. In the susceptible group, most fish carried SNP combinations linked to weakness, while in the resistant group, most carried the opposite pattern. Overall, the genetic test correctly classified about 88 percent of fish, and the match between DNA pattern and survival was statistically strong. Around 15 percent of the entire population carried a fully resistant genotype, suggesting that a disease-resilient strain is already starting to emerge.
What this means for future trout farms
For non-specialists, the core message is simple: by listening to which genes switch on during infection and scanning trout DNA for tiny, consistent differences, scientists have identified a short list of genetic markers that help distinguish hardy fish from vulnerable ones. These markers could let farmers screen young trout before they ever face disease, gradually building stocks that suffer fewer losses and require fewer treatments. While more work is needed to confirm these results in larger and different populations, the study shows how careful genetic detective work can support healthier aquaculture and help safeguard fish in some of the world’s highest rivers.
Citation: Zhou, J., Sun, S., Wang, W. et al. Transcriptome and resequencing reveal immune-related genes and molecular markers associated with Aeromonas salmonicida resistance in Salmo trutta fario. Sci Rep 16, 14909 (2026). https://doi.org/10.1038/s41598-026-45045-8
Keywords: brown trout, fish disease resistance, aquaculture genetics, Aeromonas salmonicida, SNP markers