Clear Sky Science · en
Splicing retention and enhancer divergence govern the evolutionary fate of ohnologues following whole-genome duplication in rainbow trout
Why Extra Gene Copies Matter in Fish
Rainbow trout, like many other fish, carry an unusual legacy in their DNA: at some point in their history, their entire genome was duplicated. This means many of their genes exist in pairs, raising a basic question about evolution: what happens to all those extra copies over millions of years? This study looks at how those duplicated genes are used, trimmed, and repurposed in trout, focusing on how they are turned on and off and how their messages are cut and rearranged before making proteins. The answers help explain how complex vertebrate genomes can innovate without falling apart under their own complexity.
From One Genome to Two
Roughly 80–100 million years ago, the ancestors of salmon and trout went through a whole-genome duplication. Instead of having just one copy of each gene, they suddenly had two, called “ohnologues.” Extra copies create opportunities: one can keep the original job while the other experiments with new roles. But they also create risks, because the cell must keep the overall balance of gene activity under control. Using a high-quality rainbow trout genome, gene activity data from six organs, and comparisons to a closely related fish (Northern pike) that did not experience the same duplication, the authors tracked nearly ten thousand pairs of duplicated genes to see how they have fared over deep time.
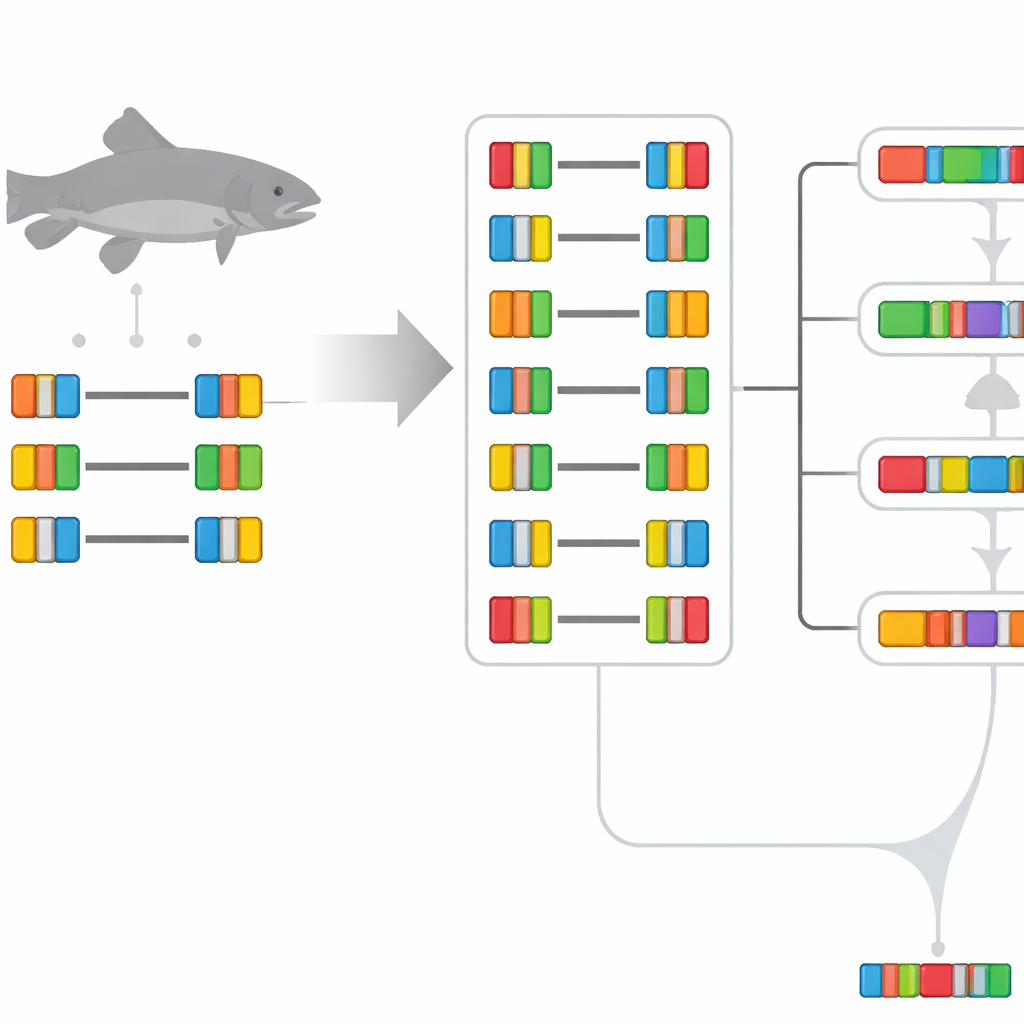
Keeping, Tweaking, or Reinventing Gene Jobs
The team found that the most common fate of duplicated trout genes is surprisingly conservative. More than 70% of gene pairs still show expression patterns similar to the inferred ancestral gene, meaning both copies kept roughly the same role and activity across tissues. A smaller fraction of pairs show one member diverging to take on a new expression pattern (neofunctionalization) or both copies drifting away together into distinct patterns (specialization). Classic splitting of the original job neatly between the two copies (subfunctionalization) turned out to be rare. When the researchers extended their comparison to other fish that went through older duplication events, they saw that, as time passes, more gene pairs tend to drift toward new roles, especially in species with smaller, more streamlined genomes.
How Messages Are Cut and Pasted
Genes do not just turn on or off; their RNA messages can be edited in different ways through a process called alternative splicing, which mixes and matches sections of the message to make multiple protein variants. In trout, more than four out of five genes are alternatively spliced, with an average of about seven distinct message forms per gene. Contrary to earlier ideas that extra copies quickly lose these extra message forms, gene duplication in trout is often accompanied by a gain in splicing complexity. Duplicated genes created by whole-genome duplication tend to have more splice variants than single-copy genes, and they lose those variants only gradually over evolutionary time. The authors show that the relationship between family size and splicing is not simply “more copies, fewer variants,” but instead follows a hump-shaped pattern: moderate-sized gene families tend to be the most richly spliced.
Different Paths for Splicing Evolution
To understand how splicing itself evolves after duplication, the researchers compared trout gene pairs to their single ancestral counterparts in pike. They grouped pairs into three scenarios. In the “accelerated” model, the duplicates together have more message variants than the ancestor; in the “function-sharing” model, the two copies split the ancestor’s variants between them; and in the “independent” model, overall splicing stays similar to the ancestor. In trout and Atlantic salmon, all three patterns occur, but a key insight is that accelerated splicing appears common soon after duplication, whereas the independent model dominates in the long run. Over time, ohnologues slowly lose splice variants while other kinds of duplicated genes, created one-by-one rather than by a whole-genome event, tend to gain variants as they age.
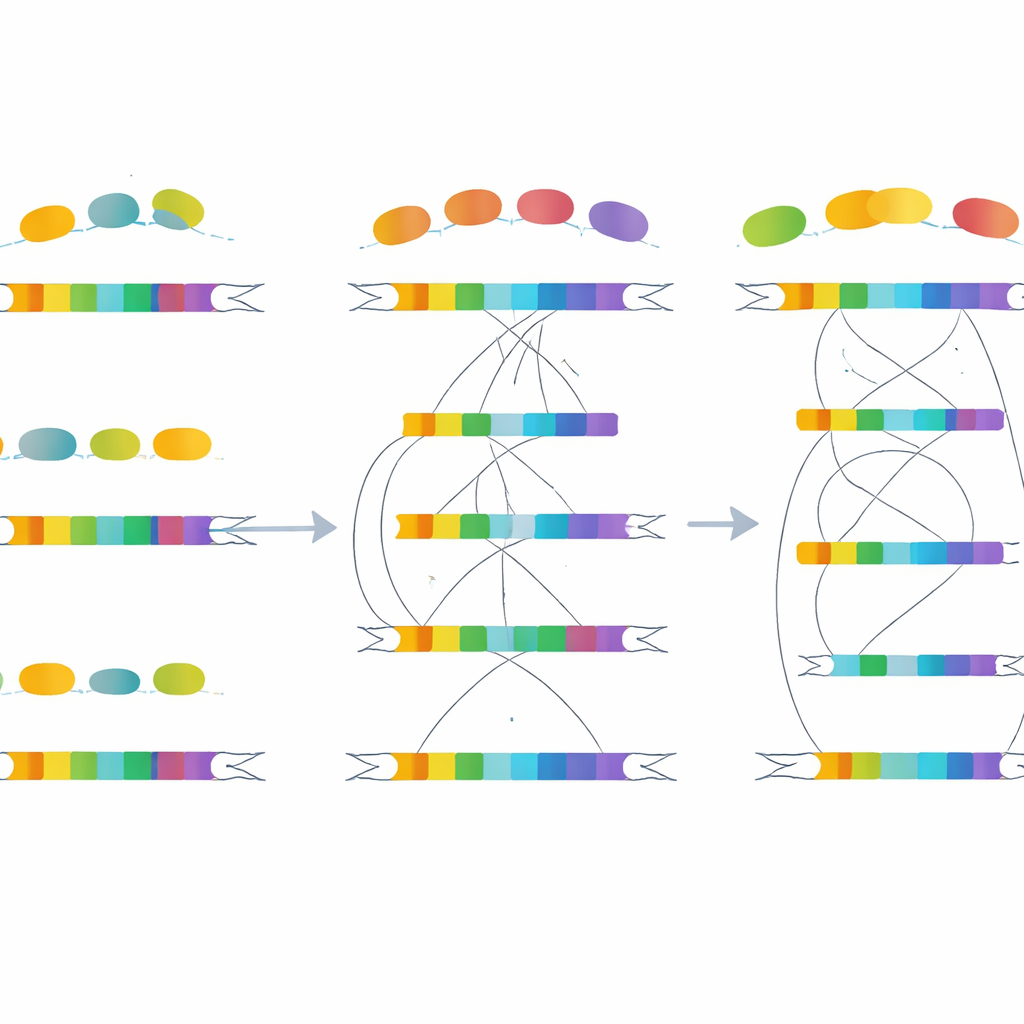
Epigenetic Switches and Enhancer “Rewiring”
The story does not end with DNA sequences. The team overlaid maps of chemical tags on DNA-packaging proteins—histones—that act as flags for active or silent regions in the genome. They found that duplicated genes generally carry strong marks of active control elements, especially in regions that act like enhancers, boosting gene activity at a distance. Gene pairs with rapidly evolving splicing patterns show particularly high levels of enhancer-associated marks and lower levels of repressive marks. Meanwhile, conserved gene pairs show more similar histone patterns between their two copies. This suggests that changes in regulatory elements, rather than in the gene bodies alone, help steer both expression and splicing differences between duplicates.
What This Means for Evolution
In everyday terms, this work shows that when a whole genome duplicates, evolution often first saves both copies and lets them experiment with extra ways of cutting and using their messages. Only slowly are these options trimmed back. Many genes retain their original roles for tens of millions of years, while a subset branch off into new functions or expression patterns. Crucially, the study overturns the simple idea that splicing diversity must quickly collapse after duplication and highlights a major role for independent, long-term maintenance of splicing levels. By linking these patterns to changes in enhancer activity and chromatin marks, the authors provide a mechanistic picture of how extra genetic “hardware” can be stabilized, repurposed, and fine-tuned, helping fish—and by extension other vertebrates—expand their biological toolkit.
Citation: Ali, A., Al-Tobasei, R., Zhou, H. et al. Splicing retention and enhancer divergence govern the evolutionary fate of ohnologues following whole-genome duplication in rainbow trout. Sci Rep 16, 13265 (2026). https://doi.org/10.1038/s41598-026-44703-1
Keywords: whole genome duplication, alternative splicing, rainbow trout, gene regulation, epigenetics