Clear Sky Science · en
Mechanistic insights into the disaggregation of amyloid-β fibrils by EPPS via replica-exchange molecular dynamics simulations
Why this research matters
Alzheimer’s disease slowly robs people of memory and independence, and one of its hallmarks is the buildup of sticky protein fibers in the brain called amyloid plaques. A small laboratory chemical known as EPPS has been shown in mice to help clear these plaques and improve memory, but how it actually attacks the fibers at the microscopic level has remained a mystery. This study uses advanced computer simulations to watch, atom by atom, how EPPS latches onto these harmful fibers and begins to pry them apart, offering clues that could guide the design of future Alzheimer’s drugs.
The problem of stubborn protein clumps
In Alzheimer’s disease, fragments of a protein called amyloid-beta can fold into flat, sheet-like structures that stack into long, rope-like assemblies known as fibrils. These fibrils tangle together to form the plaques seen in patients’ brains and are thought to damage nearby nerve cells. The stability of these fibers comes from a combination of water‑repelling regions that stick together and precisely aligned backbones that link up through many tiny hydrogen bonds, like teeth in a zipper. Because these forces are so strong and numerous, once fibrils are formed, they are notoriously difficult to pull apart, which is why finding ways to destabilize them is such an important therapeutic goal.
A closer look at how EPPS meets the fiber
To probe the interaction between EPPS and amyloid fibers, the researchers turned to replica‑exchange molecular dynamics, a powerful simulation technique that lets them explore many possible motions and shapes of molecules over time. They started with a pre‑assembled mini‑fiber composed of five stacked amyloid strands surrounded by water and dissolved salt, then added an equal number of EPPS molecules placed randomly around it. By running multiple simulations at closely spaced temperatures near body temperature, they could track where EPPS prefers to bind and how its presence alters the motion of the protein strands, capturing early events that cannot yet be easily seen in experiments.
Where EPPS grabs on first
The simulations revealed that EPPS is not drawn uniformly to the entire fiber. Instead, it is strongly attracted to the outermost strands, especially at specific negatively charged sites on the protein surface. Here, the positively charged part of EPPS forms tight ionic contacts, acting almost like a magnet snapping onto an exposed screw at the fiber’s edge. These favored docking spots are more accessible to the surrounding water than the tightly packed interior, explaining why EPPS seldom ventures into the middle of the bundle. Once attached, EPPS tends to wedge its flexible tail between neighboring strands, positioning itself in a way that can interfere with the regular, sheet‑like packing that keeps the fibril rigid.
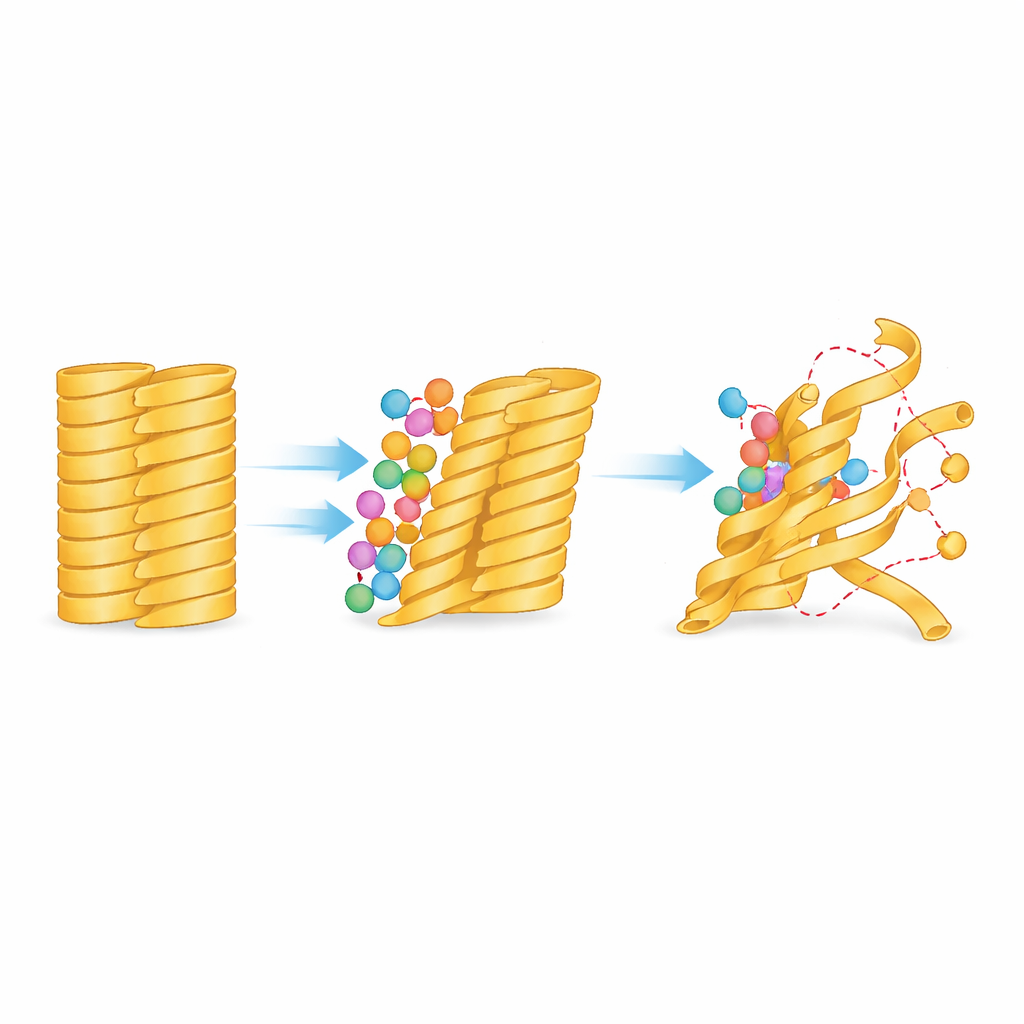

How edge tugging loosens the whole bundle
Binding at the edges has far‑reaching consequences. The team measured how much each part of the fiber wiggled over the course of the simulations and found that segments at the periphery became noticeably more mobile soon after EPPS settled in. This increase in motion did not remain confined to the outer strands; over time it propagated inward, indicating that the disturbance created at the surface ripples through the structure. Detailed analysis showed that the number of stabilizing hydrogen bonds between adjacent strands decreased, particularly where EPPS’s tail slipped between them, and that portions of the protein lost their orderly, sheet‑like character. In some regions, the preferred backbone shapes shifted substantially, signaling that the strands were bending and twisting away from the tightly locked fibril arrangement.
Implications for future Alzheimer’s treatments
Taken together, the simulations paint EPPS as a small but strategic saboteur: it targets accessible charged spots on the outside of amyloid fibers, uses strong electrical attraction to anchor itself, then leverages its flexible structure to pry apart neighboring strands and erode the fiber’s internal glue. Although the computer runs were too short to capture complete breakup of the fibrils, the observed loss of order and bonding strongly suggests that continued EPPS attack would eventually lead to disassembly. For non‑specialists, the key message is that EPPS does not simply dissolve plaques in a vague way; it follows a specific microscopic playbook that can now inform the design of new, more potent molecules aimed at loosening and clearing Alzheimer’s‑related protein deposits.
Citation: Choi, KE., Pae, A.N. & Cho, NC. Mechanistic insights into the disaggregation of amyloid-β fibrils by EPPS via replica-exchange molecular dynamics simulations. Sci Rep 16, 13034 (2026). https://doi.org/10.1038/s41598-026-42391-5
Keywords: Alzheimer’s disease, amyloid plaques, protein aggregation, molecular simulations, drug design