Clear Sky Science · en
Near telomere-to-telomere diploid genome assembly of Acrossocheilus wenchowensis
Why this mountain stream fish matters
Acrossocheilus wenchowensis, sometimes called a freshwater grouper, may look like an ordinary river fish, but it is prized in southern China both as food and as an ornamental species. It grows in cool mountain streams, has flavorful meat rich in healthy fats, and shows striking differences in growth between males and females—traits that make it attractive for fish farming. To understand and eventually manage these traits, scientists need a complete instruction manual for the fish: its genome. This study delivers one of the most complete fish genomes to date, stitched together nearly from one chromosome tip to the other.
A complete playbook from end to end
The authors set out to build a near “telomere-to-telomere” genome for A. wenchowensis. Telomeres are the protective caps at the ends of chromosomes, and reaching from one end to the other with no gaps is the gold standard for genome quality. Using several cutting-edge DNA sequencing technologies, they assembled two full sets of chromosomes—one for each parental contribution in this diploid species. Each set, or haplotype, spans about 860 to 870 million DNA letters, with long, continuous stretches that far surpass earlier fish genomes. For most chromosomes, the team could trace the sequence from one telomere to the other, with only a handful of tiny gaps remaining.
Finding the hidden anchors of the genome
Beyond just laying out the DNA, the researchers looked for key structural features that organize the chromosomes. They discovered a repeated DNA sequence of 262 bases that appears once on every chromosome and marks the centromere, the central anchor point needed for proper chromosome separation during cell division. This repeat comes from a mobile genetic element, revealing how jumping DNA has helped shape the chromosome backbone. Around these centromeres, genes become sparse, while other repetitive elements cluster densely, a pattern also seen in humans and other animals. Having this level of detail for a fish genome is unusual and opens the door to exploring how chromosome structure influences traits over evolution.
Spotting differences between the two chromosome sets
Because the assembly keeps the two parental haplotypes separate, the team could compare them directly and catalogue how they differ. They found millions of single-letter changes, hundreds of thousands of small insertions and deletions, and thousands of larger rearrangements, such as segments that have been inverted, duplicated, moved, or copied in different numbers. Many of these changes overlap with mobile genetic elements, hinting that these restless sequences are major drivers of structural change. When variants overlapped genes, most lay within the gene bodies themselves, and many small insertions or deletions were predicted to alter the resulting proteins. This fine-grained map shows how much variation can exist even within the genome of a single individual.
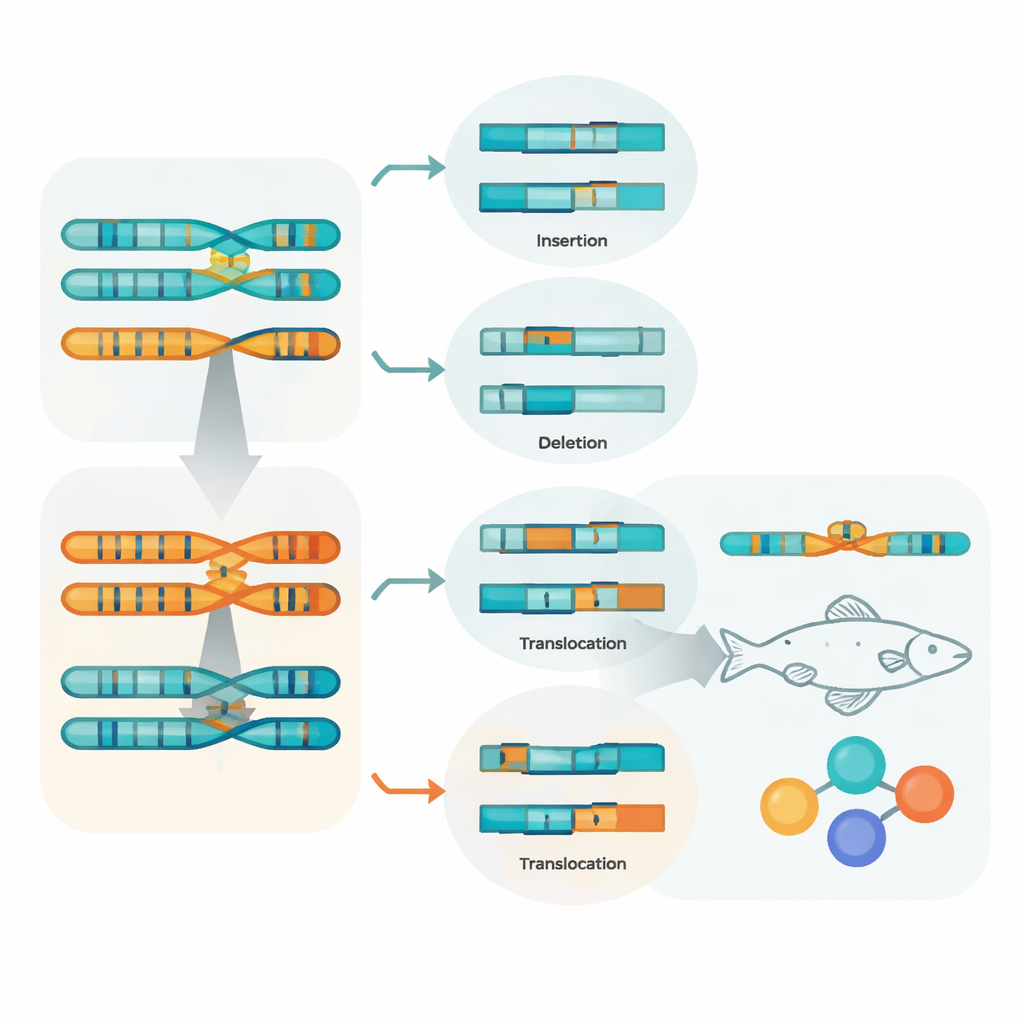
Tracing family history and regional differences
Armed with this reference genome, the authors compared A. wenchowensis to related carps and minnows. Their analyses suggest that its genus, Acrossocheilus, split from a close cousin genus, Onychostoma, around 13.7 million years ago, and that A. wenchowensis diverged from its near relative A. fasciatus about 5.25 million years ago. The team also resequenced 80 fish from four river regions in China. Genetic data showed that most populations are relatively similar and exchange genes, but one group from Longyan (LY) stands out with much greater genetic distance from the others. This pattern points to limited mixing and local adaptation, and it may have practical implications for managing wild stocks and breeding programs.
What this means for aquaculture and biology
For non-specialists, the key message is that scientists have produced an exceptionally complete blueprint of a commercially and ecologically important freshwater fish. This resource reveals where its chromosomes begin, end, and anchor; how its two parental sets of DNA differ; how it fits into the broader fish family tree; and how populations spread across China vary genetically. Such knowledge is the foundation for future work on sex determination, growth, and adaptation to changing environments. In time, this genome could help breeders develop more efficient, sustainable aquaculture lines and aid conservationists in protecting the genetic diversity of mountain stream fishes.
Citation: Xue, L., Luo, M., Wang, H. et al. Near telomere-to-telomere diploid genome assembly of Acrossocheilus wenchowensis. Sci Data 13, 452 (2026). https://doi.org/10.1038/s41597-026-06752-z
Keywords: fish genome, telomere to telomere, freshwater grouper, population genetics, aquaculture