Clear Sky Science · en
Clusia genomes shed light on the evolution and diversity of crassulacean acid metabolism physiotypes
How some trees beat the heat
As the world warms and droughts intensify, farmers and scientists are searching for crops that can stay productive while using far less water. This study looks at tropical trees in the genus Clusia, famous for a special way of doing photosynthesis called crassulacean acid metabolism, or CAM. By comparing the genomes and daily rhythms of three closely related trees that range from barely to strongly CAM, the authors show how ancient genome duplications and later gene losses helped create a remarkable variety of water‑saving strategies. Their findings offer clues for how similar tricks might one day be built into major food crops.
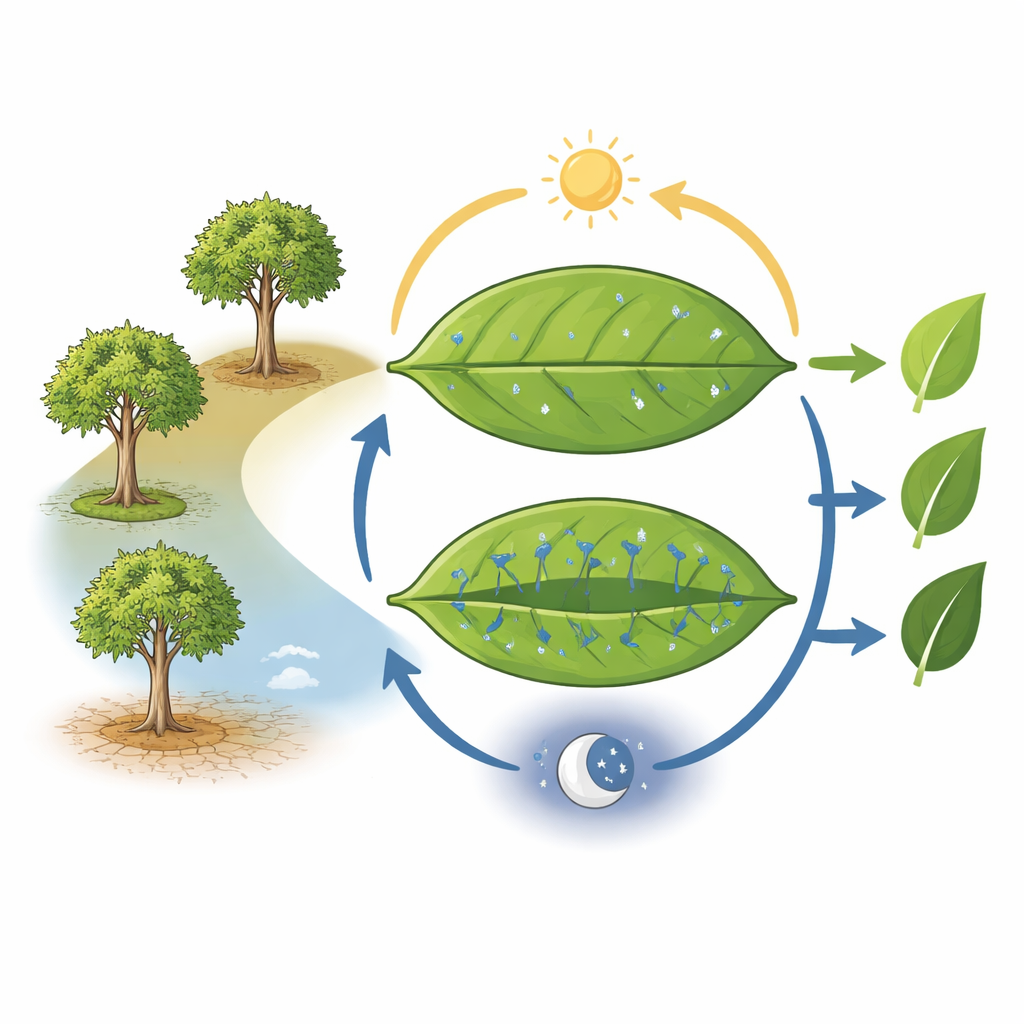
Trees with different daily breathing styles
Clusia trees can open and close the pores in their leaves (stomata) on very different schedules. Classic “C3” plants open stomata during the day to take in carbon dioxide, but lose a lot of water doing so. CAM plants reverse much of this routine: they open stomata at night, store carbon as organic acids, and release it in the day while stomata stay mostly shut, saving water. The team focused on three species: Clusia major, which behaves mostly like a C3 plant but shows hints of CAM; Clusia minor, which can switch CAM on under stress; and Clusia rosea, a strong CAM performer. Careful gas‑exchange measurements and acidity tests over 24 hours confirmed these distinct “physiotypes” and cleared up past mix‑ups in species identity.
Ancient genome doubling and its aftermath
Using long‑read DNA sequencing and chromosome‑level assembly, the researchers found that all three Clusia species carry the legacy of multiple rounds of whole‑genome duplication. In particular, C. major shows clear signs of being derived from an ancient tetraploid that has since “diploidized” – in other words, it once had four copies of each chromosome, but over millions of years many duplicate genes were switched off or lost. The genome is now organized into pairs of corresponding chromosomes that still mirror one another, yet differ in size and content because of bursts of transposable elements – mobile DNA pieces that have expanded in some regions and disappeared from others. Across the genome, more than a quarter of former genes appear to have become pseudogenes, broken by mutations or fragmented by rearrangements.
Rewiring the leaf’s night‑time fuel system
CAM depends on a reliable nightly fuel supply: stored carbohydrates must be turned into building blocks that feed carbon‑fixing reactions in the dark. By combining genomics with RNA, protein, and metabolite measurements across day and night, the authors homed in on genes involved in starch breakdown and related sugar pathways. In C. major, many key enzymes that normally channel leaf starch into phosphoenolpyruvate (the main substrate for night‑time CO2 capture) show tell‑tale scars of diploidization: missing exons, disruptive mobile‑DNA insertions, or near‑complete gene loss. Other surviving copies have gained new regulatory switches in their control regions, shifting their activity from daytime to night. As a result, C. major accumulates more leftover starch and less malic acid overnight than strong‑CAM C. rosea, and compensates by leaning more heavily on soluble sugars and special protective compounds like raffinose during drought.
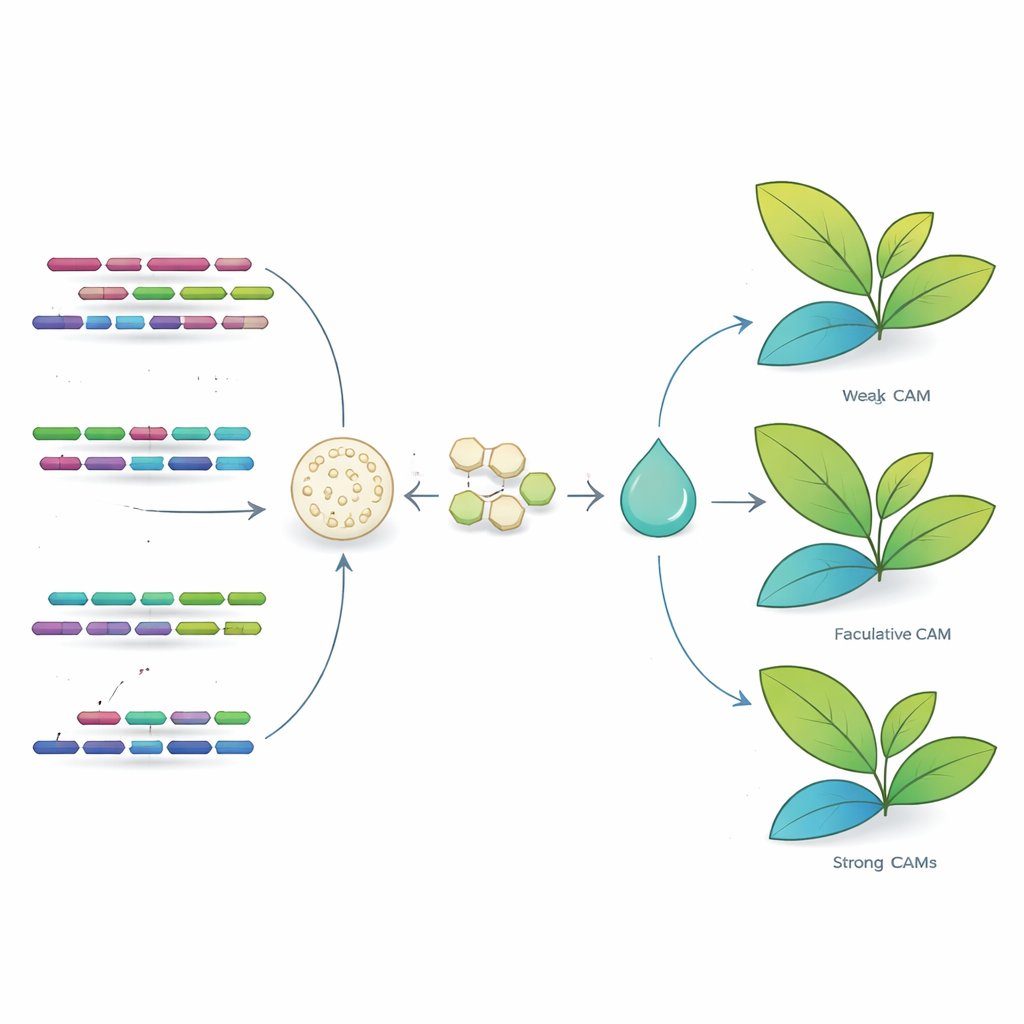
Linking gene history to survival strategies
Putting these pieces together, the study proposes that ancient genome duplications in the Clusia lineage created extra copies of many genes, including those needed for CAM‑like carbon concentrating and flexible starch use. Over time, as different lineages adapted to different habitats—from salty, sun‑baked coasts to humid island forests—diploidization trimmed and reshaped these redundant networks in distinct ways. In species such as C. rosea, the resulting gene sets support strong, water‑saving CAM, while in C. major they underpin a mixed C3 + CAM strategy that still improves water use but relies less on night‑time carbon storage. This evolutionary “tuning” of duplicated genes, rather than the invention of entirely new ones, appears to underlie the rich variety of CAM physiotypes seen in the genus.
Why this matters for future crops
For non‑specialists, the take‑home message is that water‑saving photosynthesis in trees like Clusia did not arise from a single magic gene, but from long‑term remodeling of existing pathways after whole‑genome doubling. By showing exactly which starch and carbon‑handling genes were lost, repurposed, or given new daily rhythms in species with weak, inducible, or strong CAM, this work provides a concrete blueprint for efforts to engineer CAM‑like traits into conventional crops. Rather than copying an entire CAM system, breeders and biotechnologists may be able to adjust specific branches of carbohydrate metabolism and gene regulation to help future crops breathe more like a CAM plant when water is scarce.
Citation: Kramml, H.M., Herpell, J.B., Priemer, C. et al. Clusia genomes shed light on the evolution and diversity of crassulacean acid metabolism physiotypes. Nat Commun 17, 3937 (2026). https://doi.org/10.1038/s41467-026-71958-z
Keywords: crassulacean acid metabolism, plant genome duplication, drought tolerance, starch metabolism, photosynthesis evolution