Clear Sky Science · en
Spatial cartography of human thymus enables the geopositioning of lineage transcription factors in rare mimetic thymic epithelial cells
Why mapping a tiny organ matters
The thymus is a small organ tucked behind your breastbone, yet it quietly shapes your immune system by training newborn T cells to recognize threats without attacking your own tissues. As people live longer and immune-related diseases become more common, scientists need to understand this training ground in fine detail. This study builds a detailed spatial “map” of the human thymus in fetuses and young children, revealing how different cell neighborhoods are arranged and how rare, specialized cells help prevent autoimmunity.
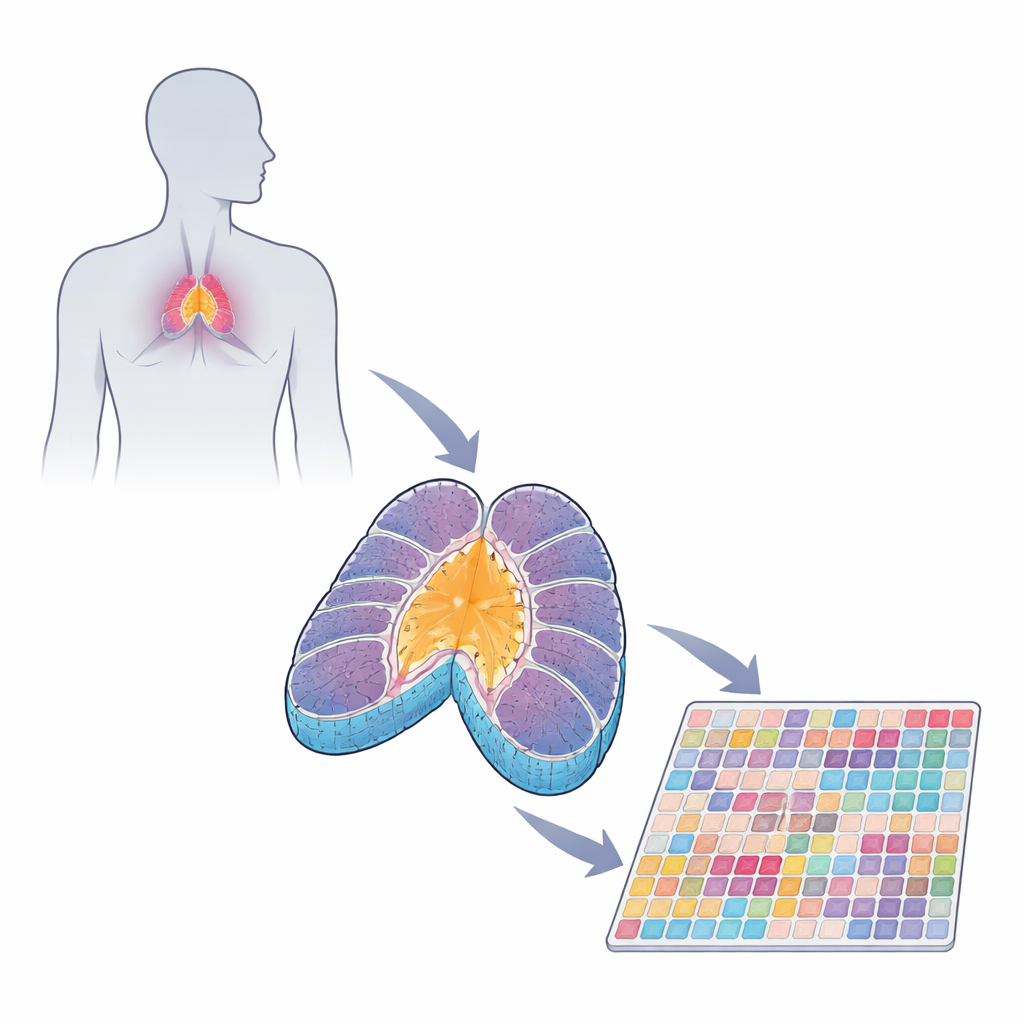
The thymus as an immune training school
T cells begin life as immature recruits arriving from the fetal liver or bone marrow. In the thymus they pass through a structured journey. The outer region, called the cortex, is filled with developing T cells and supportive epithelial cells that test whether each T cell can recognize molecular signs of the body. Those that pass move into the inner region, the medulla, where another round of testing removes cells that react too strongly to self. This process, called central tolerance, is what keeps a healthy immune system from attacking normal organs such as the pancreas, skin, or brain.
Building a high‑resolution thymus atlas
Previous studies profiled thymus cells by sequencing them one by one after they were removed from the tissue. That revealed who the major players are, but not where they sit or how they are arranged. In this work, the authors used a technique called spatial transcriptomics (Stereo‑seq), which reads thousands of genes directly from intact tissue slices, along with highly multiplexed protein imaging. They applied these tools to human fetal thymi from 13 to 18 weeks of gestation and to pediatric thymi from 7 weeks to 6 years of age. By combining gene and protein data, they could outline classic regions such as cortex, medulla, and fibrous septa, and then subdivide them into smaller “niches” with distinct mixes of T cells, epithelial cells, fibroblasts, and other immune cells.
Decoding the signals that shape T cell fate
With this atlas, the team examined how cells talk to each other using signaling molecules. In the cortex, they found pathways that help maintain fibroblasts and guide immature T cells, including signals involving the protein CXCL12 and its receptor on T cells. These cues help young T cells move through the cortex and undergo their first selection steps. In the medulla, they saw a different set of signals dominating, such as chemokines that bring mature T cells into the right areas, and factors involved in antigen presentation and cell death. Together, these circuits ensure that T cells not only learn what “self” looks like but are also eliminated if they respond too aggressively.
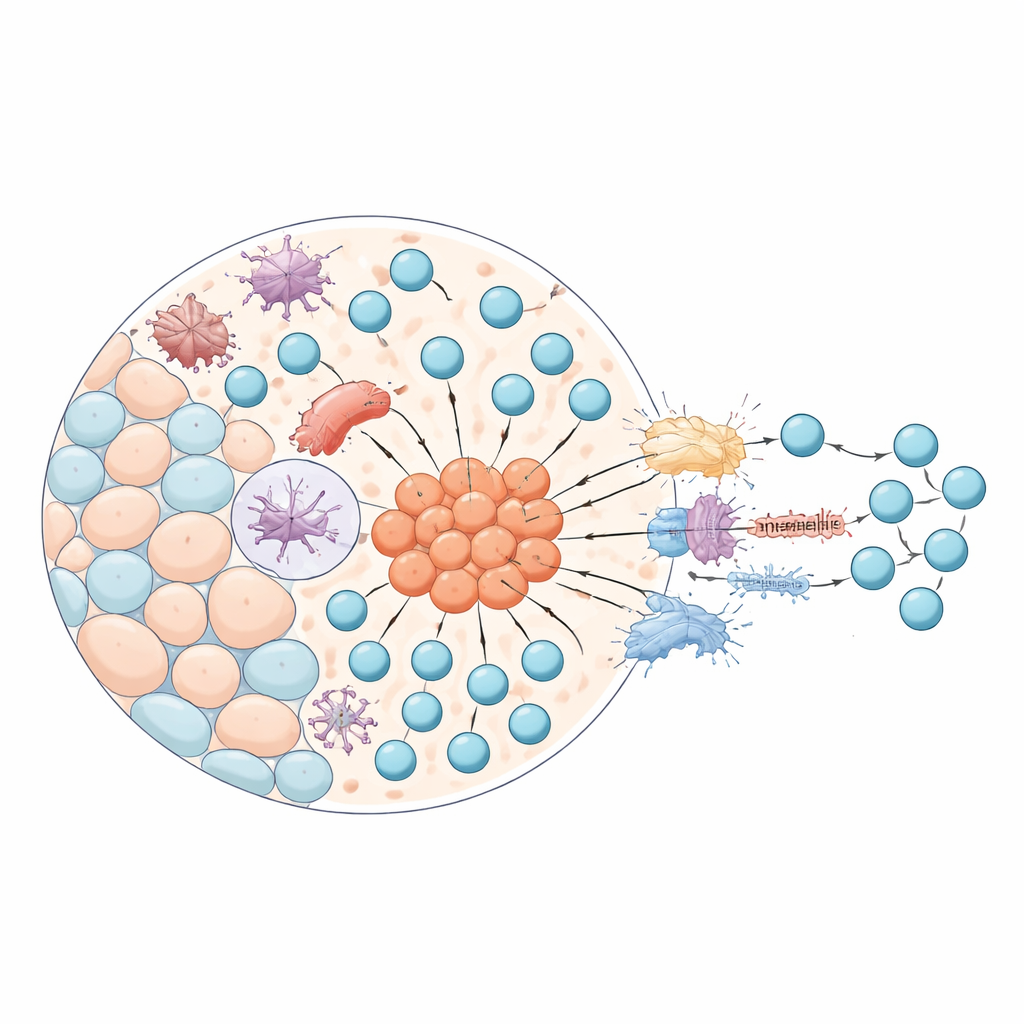
Rare mimicking cells and their control switches
A central focus of the study is a little‑known group of thymic epithelial cells that behave like stand‑ins for cells from other organs. These “mimetic” cells turn on gene programs normally used by muscle, nerve, or gland cells, allowing the thymus to display a wide catalog of self‑molecules to developing T cells. Using their spatial map, the researchers systematically searched for transcription factors—gene control switches—active in epithelial cells located in the medulla. They uncovered 70 such factors linked to mimetic cells, including well‑known regulators like AIRE and FEZF2, as well as dozens of previously underappreciated candidates. These cells are extremely rare, making up less than half a percent of the tissue, yet they cluster in specialized hotspots near structures called Hassall’s corpuscles.
What this atlas reveals about immune tolerance
By overlaying gene expression, protein patterns, and cell locations, the study shows that mimetic epithelial cells sit in tight neighborhoods rich in antigen‑presenting cells and mature T cells. Their transcription factors are connected to developmental programs of many organs—brain, muscle, endocrine glands, and more—suggesting that each mimetic cell type specializes in representing a different part of the body’s “self” library. The work also highlights technical limits: current spatial methods cannot yet trace every individual cell border in this crowded organ, but the authors move closer by using higher‑resolution chips and integrated protein data.
How this advances our understanding of health and disease
For a non‑specialist, the key message is that the thymus is not just a simple filter but a finely zoned landscape where rare, highly specialized cells help teach T cells tolerance by imitating many other tissues. This study provides a detailed map of that landscape in early human life, pinpoints where these mimetic cells live, and identifies the gene switches that shape their identity. Such knowledge offers a foundation for future work on why immune tolerance sometimes fails, contributing to autoimmune diseases, and may eventually inform strategies to rebuild or manipulate the thymus in aging, infection, or cancer.
Citation: Kamaraj, U.S., Chen, Y., Lei, J. et al. Spatial cartography of human thymus enables the geopositioning of lineage transcription factors in rare mimetic thymic epithelial cells. Nat Commun 17, 3721 (2026). https://doi.org/10.1038/s41467-026-68596-w
Keywords: thymus, T cell development, spatial transcriptomics, immune tolerance, thymic epithelial cells