Clear Sky Science · en
iS2C2: a cointelligent platform for mechanistic discovery of disease cellular crosstalk
Why smart teamwork between cells and computers matters
When we get sick, our cells start sending abnormal signals to one another, quietly reshaping tissues long before symptoms appear. Modern biology can now record these signals gene by gene in millions of individual cells, but the resulting data are too vast and complex for human experts to interpret alone. This paper introduces iS2C2, a new platform that lets rigorous data analysis and large language models work together to uncover how cells talk in diseases such as Alzheimer’s and cancer, pointing toward more precise ways to diagnose and treat illness.
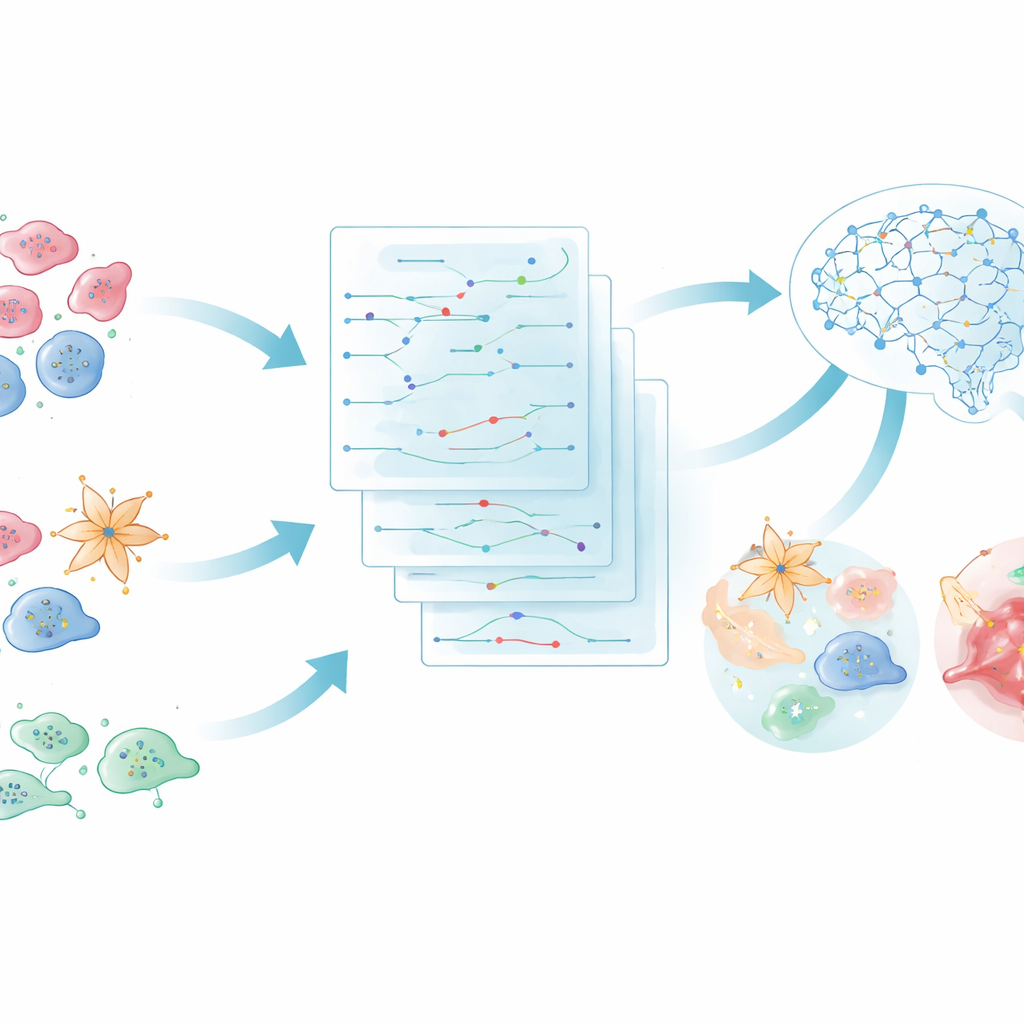
How cells hold quiet conversations in disease
Every tissue in the body is a busy neighborhood of cells constantly exchanging signals. One cell releases a molecule that acts as a "ligand," and another cell receives it through a matching "receptor" on its surface, triggering internal changes. When this conversation goes wrong, it can drive conditions from memory loss in Alzheimer’s disease to tumor spread in cancer. New techniques that measure RNA in single cells and in tissue slices now reveal which genes are active in each cell and where those cells sit in three‑dimensional space. The challenge is turning that flood of numbers into clear stories about who is talking to whom, and how those messages alter the course of disease.
A new map of cell‑to‑cell messages
The first half of iS2C2 is a specialized analysis engine called S2C2. It takes single‑cell and spatial gene data from human or mouse tissues and searches for pairs of cells where one strongly expresses a signaling molecule and the other expresses its matching receptor. Instead of stopping there, S2C2 follows the signal inward, tracing it through chains of known pathway components down to key control genes in the nucleus. Each such route, called a pathway branch, receives an activity score that reflects both how strongly the signal is sent and how strongly the downstream genes respond. This scoring system helps highlight the specific branches that are most altered in a given disease, even when the original data are noisy or incomplete.
Letting language models reason about biology
The second half of iS2C2 uses a large language model as a reasoning partner. S2C2 outputs structured files listing important signal pairs, pathway branches, and their context. Carefully crafted prompts then guide the language model to read these files step by step, pick the most influential signals, connect them with existing biological knowledge, and generate testable ideas about how they might shape disease. The authors showed that by using examples and stepwise reasoning prompts, they could reduce the model’s tendency to invent unsupported connections and improve how closely its suggestions matched the underlying data. Expert reviewers judged the resulting explanations more understandable and more helpful for planning experiments than those from earlier methods.
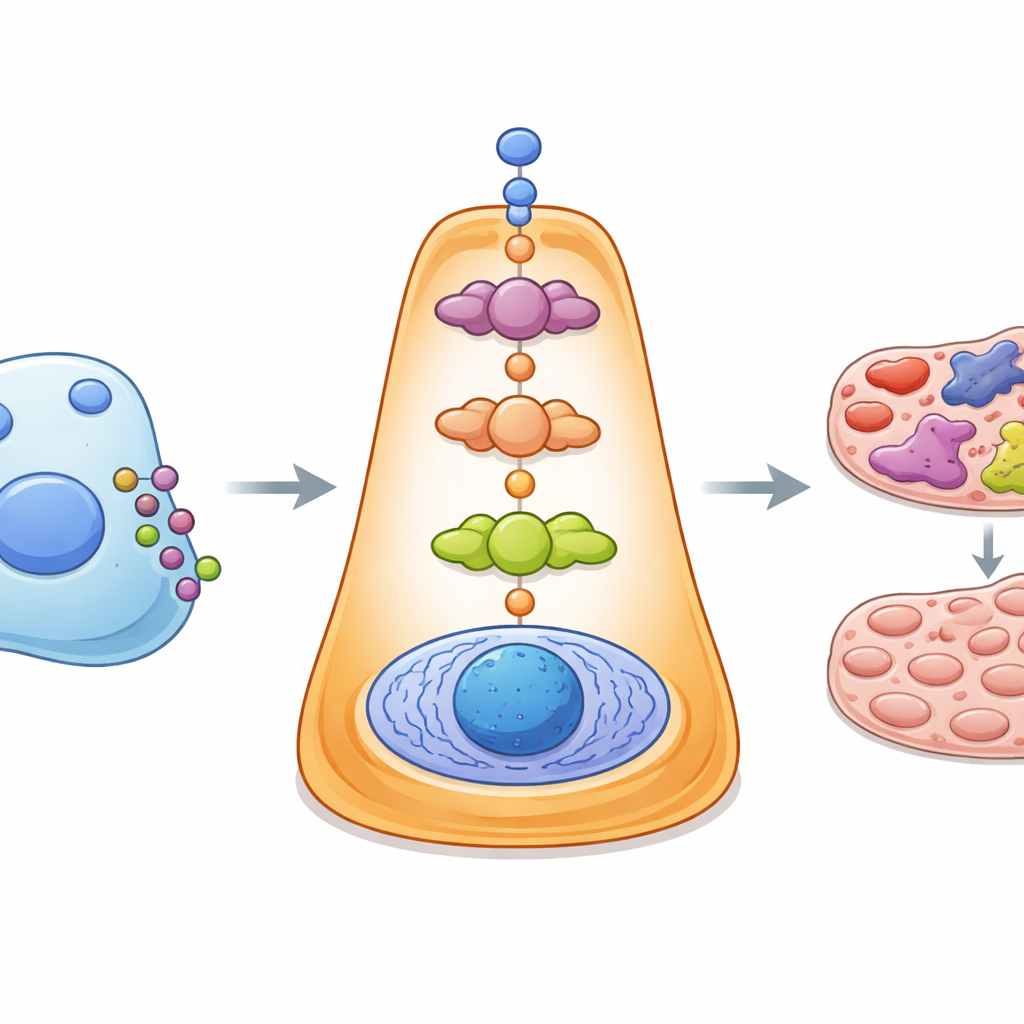
What the platform revealed in Alzheimer’s and cancer
To test iS2C2 in real disease settings, the team applied it to several large brain datasets from people with Alzheimer’s disease. The platform highlighted an unexpected conversation between support cells called astrocytes and nearby neurons, centered on a signal pair known as CSF1 and its receptor. Experiments in cell cultures confirmed that disrupting this link weakened protective pathways in neurons and altered their inflammatory behavior, supporting the idea that this cross‑talk helps shape nerve cell survival. In a second case study of breast cancer cells spreading to bone, iS2C2 combined single‑cell and spatial data to show how tumor cells and bone‑resident stem cells influence each other through intertwined signaling routes. This analysis pointed to a common control point affected by the drug tamoxifen, which in mice slowed early bone metastasis.
What this means for future medicine
In plain terms, iS2C2 is a smart go‑between that translates messy cellular data into structured patterns, then asks an AI system to explain what those patterns might mean for disease. By improving how we spot and interpret hidden lines of communication between cells, the platform helps researchers generate clearer, more testable ideas about what drives disorders like Alzheimer’s and metastatic cancer. While its suggestions still require careful human checking and lab validation, this co‑intelligent approach points toward a future where computers help scientists navigate complex biology faster, guiding the search for new diagnostics and treatments with greater confidence.
Citation: Sheng, J., Ahn, J.Y., Yang, L. et al. iS2C2: a cointelligent platform for mechanistic discovery of disease cellular crosstalk. Sig Transduct Target Ther 11, 172 (2026). https://doi.org/10.1038/s41392-026-02691-8
Keywords: cell communication, single-cell data, large language models, Alzheimer’s disease, cancer metastasis