Clear Sky Science · en
The novel genus Brakhagea gen. nov. is constituted by four planctomycetal species isolated from aquatic environments in Northern Germany
Hidden Life in Everyday Waters
Waterways in Northern Germany—quiet ponds, park lagoons, and a narrow arm of the Baltic Sea—harbor a microscopic cast of characters that we are only beginning to meet. This study tells the story of four such hidden residents: unusual pink bacteria that not only expand our map of Earth’s microbial diversity, but may also carry the genetic machinery to make new bioactive molecules of future biotechnological interest.
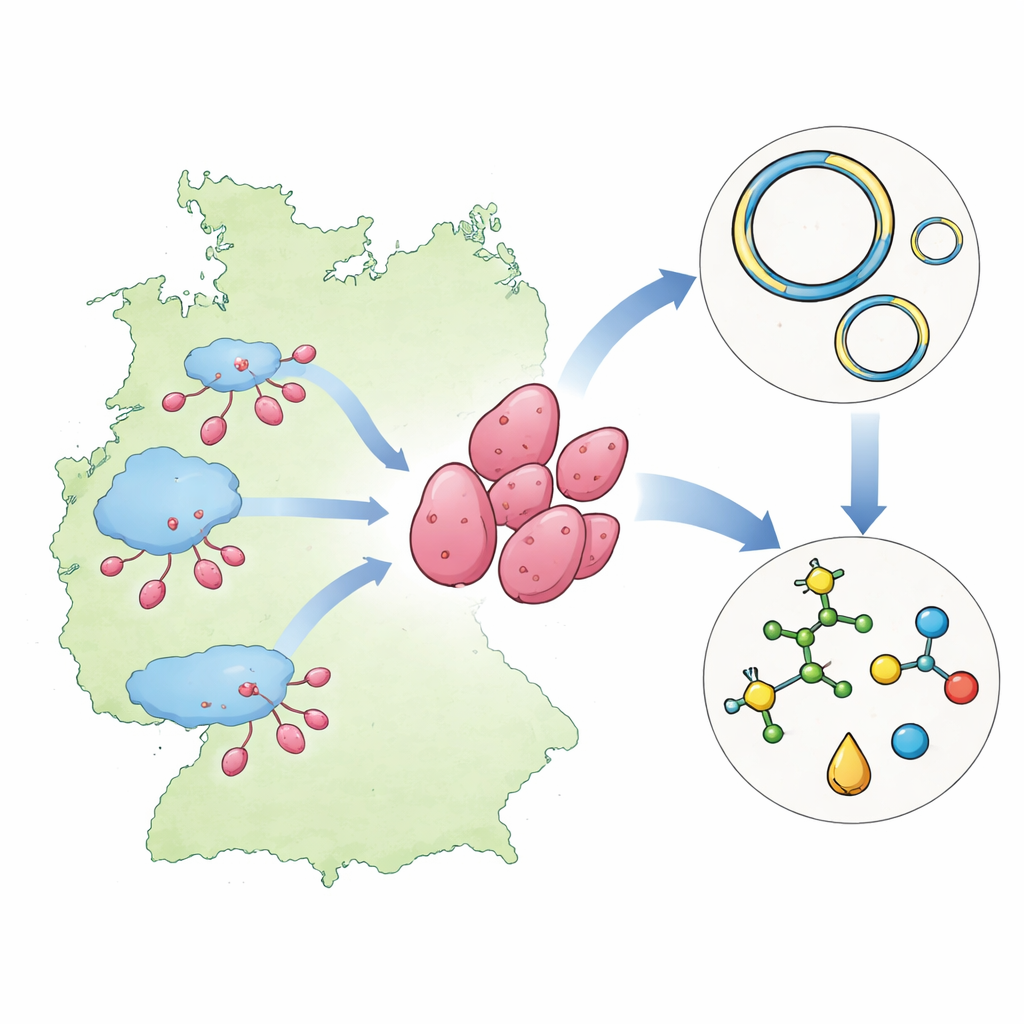
A New Branch on the Family Tree
The bacteria described here belong to a little-known group called Planctomycetota, microbes famous among microbiologists for their odd cell shapes, complex life cycles, and large, gene-rich genomes. Using a combination of DNA-based markers, the researchers showed that four newly revived strains from an old culture collection do not fit into any known genus within their broader family, Pirellulaceae. Their genetic signatures—measured by comparing key genes and whole genomes—fall well below the similarity thresholds scientists use to decide that organisms belong to the same named group. This consistent genetic distance strongly supports the creation of a brand-new genus, which the authors name Brakhagea, honoring a prominent microbiologist.
Four Species from Four Waters
Each of the four strains comes from a different aquatic setting in Northern Germany: a brackish Baltic fjord, a village pond, a wastewater treatment lagoon at a sugar factory, and a city park pond. All are aerobic, meaning they need oxygen, and they feed on organic matter rather than making their own food from sunlight. Under the microscope, their cells are small, pink, and pear-shaped, and they reproduce by budding: a tiny daughter cell grows from one pole of a larger mother cell before pinching off. Despite these shared traits, careful measurements of growth temperature, pH preference, colony appearance, and detailed genome comparisons reveal that the four strains are distinct enough to be recognized as four separate species within the new genus.
Genomes that Hint at Chemical Talent
To understand what these microbes might be capable of, the team sequenced and analyzed their genomes, which range from about 5.9 to 7.3 million DNA letters in size—comparable to other members of their family but smaller than some of the largest planctomycete genomes. All four carry a single copy of the standard ribosomal genes used for protein production, but two strains also harbor extra DNA circles known as plasmids. By comparing all five genomes in their analysis—including that of their closest known relative—the researchers built a “pangenome,” separating genes shared by all strains from those that are unique. They found more than a thousand genes common to every genome, plus large sets of accessory genes that differ from strain to strain, underscoring how much evolutionary tinkering has occurred even within this small cluster of bacteria.
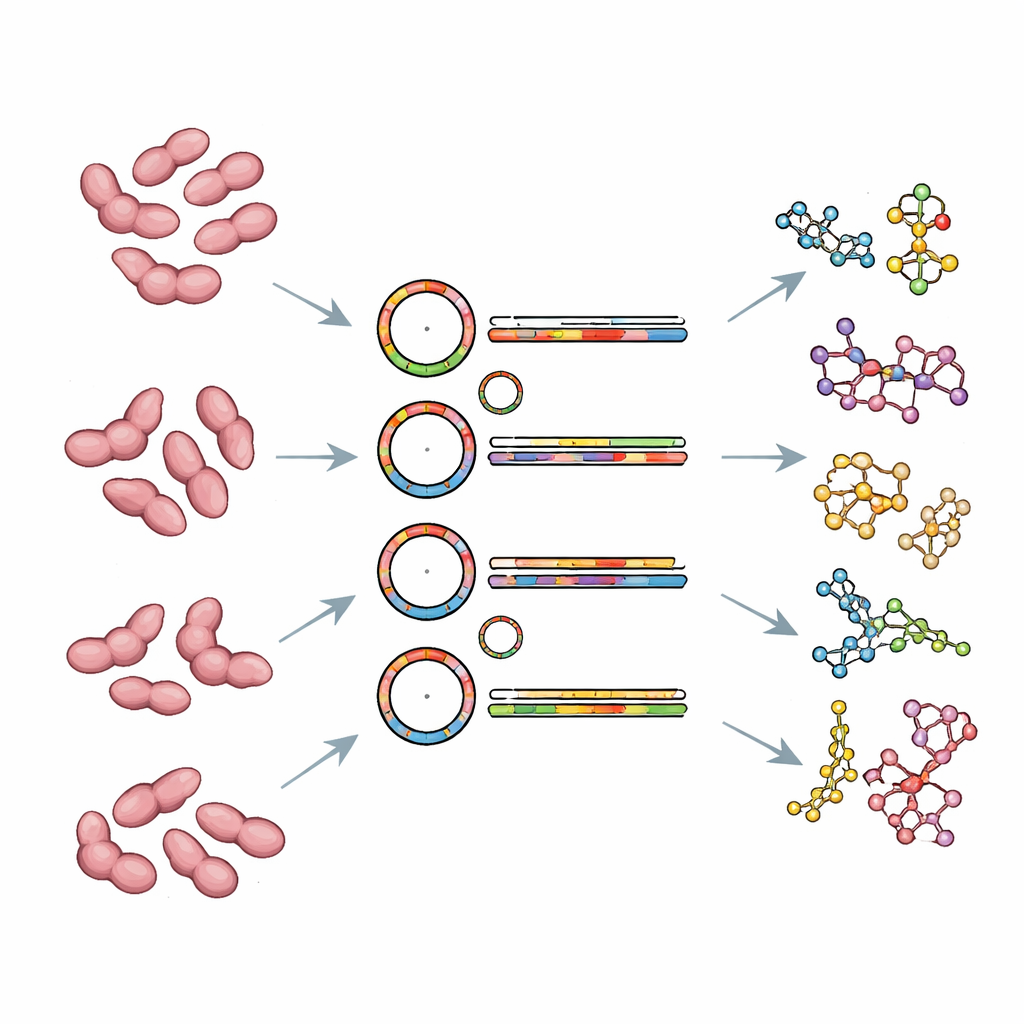
Potential Makers of New Molecules
One of the most intriguing findings lies in the genetic blueprints for specialized chemistry. Using computational tools, the authors identified between seven and twelve biosynthetic gene clusters in each new strain—stretches of DNA that often encode enzymes for making complex small molecules. Many clusters appear related to terpenes and polyketides, classes of compounds that in other microbes include antibiotics and pigments. Some clusters are unique to individual species, such as those likely involved in making resorcinol- or indole-type molecules. The genomes also contain numerous genes for enzymes that help break down complex sugars, suggesting these bacteria can feed on intricate carbohydrates found in their aquatic habitats, although the exact food preferences still need to be tested in the lab.
Why These New Microbes Matter
By defining Brakhagea and its four founding species, this work enlarges the known diversity of an already unconventional bacterial group and preserves valuable strains collected decades ago. The study leans heavily on modern genome sequencing rather than exhaustive chemical profiling, reflecting a broader shift in how microbiologists name and classify microbes in the genomic era. At the same time, the abundance of genes with unknown function is a reminder of how much we still do not understand. These newly described bacteria now serve as reference points and genetic resources for future experiments that can probe their roles in natural ecosystems, explore their defenses against viruses, and test whether their predicted biosynthetic pathways can indeed yield useful new molecules.
Citation: Kumar, G., Kallscheuer, N., Hammer, J. et al. The novel genus Brakhagea gen. nov. is constituted by four planctomycetal species isolated from aquatic environments in Northern Germany. Sci Rep 16, 12750 (2026). https://doi.org/10.1038/s41598-026-47393-x
Keywords: planctomycetes, microbial diversity, aquatic bacteria, genome sequencing, secondary metabolites