Clear Sky Science · en
Chromosomal-level genome assembly of Ichthyurus bourgeoisi Gestro using PacBio HiFi and Hi-C sequencing
A Beetle That Broke the Rules
Most beetles wear a built‑in suit of armor: a pair of hard forewings, called elytra, that form a protective shell over the body and flying wings. Ichthyurus bourgeoisi, a slender “soldier beetle” from southern China, is different. Its wing covers are soft and unusually short, leaving its hind wings and abdomen exposed. Yet this seemingly unarmored insect survives just fine. The study described here delivers a complete, chromosome‑level map of this beetle’s DNA, giving scientists a powerful new reference to explore how such an odd body plan evolved and how these insects defend themselves without the usual beetle shield.
A Soft‑Shelled Beetle and Its Survival Tricks
Ichthyurus bourgeoisi belongs to a small tropical group of beetles marked by thin, abbreviated wing covers, bright yellow‑and‑black colors, and a flexible, exposed abdomen. In most beetles, rigid elytra are a key evolutionary invention: they protect delicate wings, reduce water loss, and help the insect push through rough environments. Losing this protection ought to be a bad idea. To make up for it, Ichthyurini beetles appear to lean on other strategies. Their bold color patterns and exposed wings give them a wasp‑like appearance that may warn predators away, while chemical defenses likely make them unpalatable. Extra freedom of movement in the uncovered abdomen may also help in hunting and mating. Understanding how such a lifestyle is encoded in their DNA requires a high‑quality genome, which until now was missing for this group.
Building a Complete DNA Blueprint
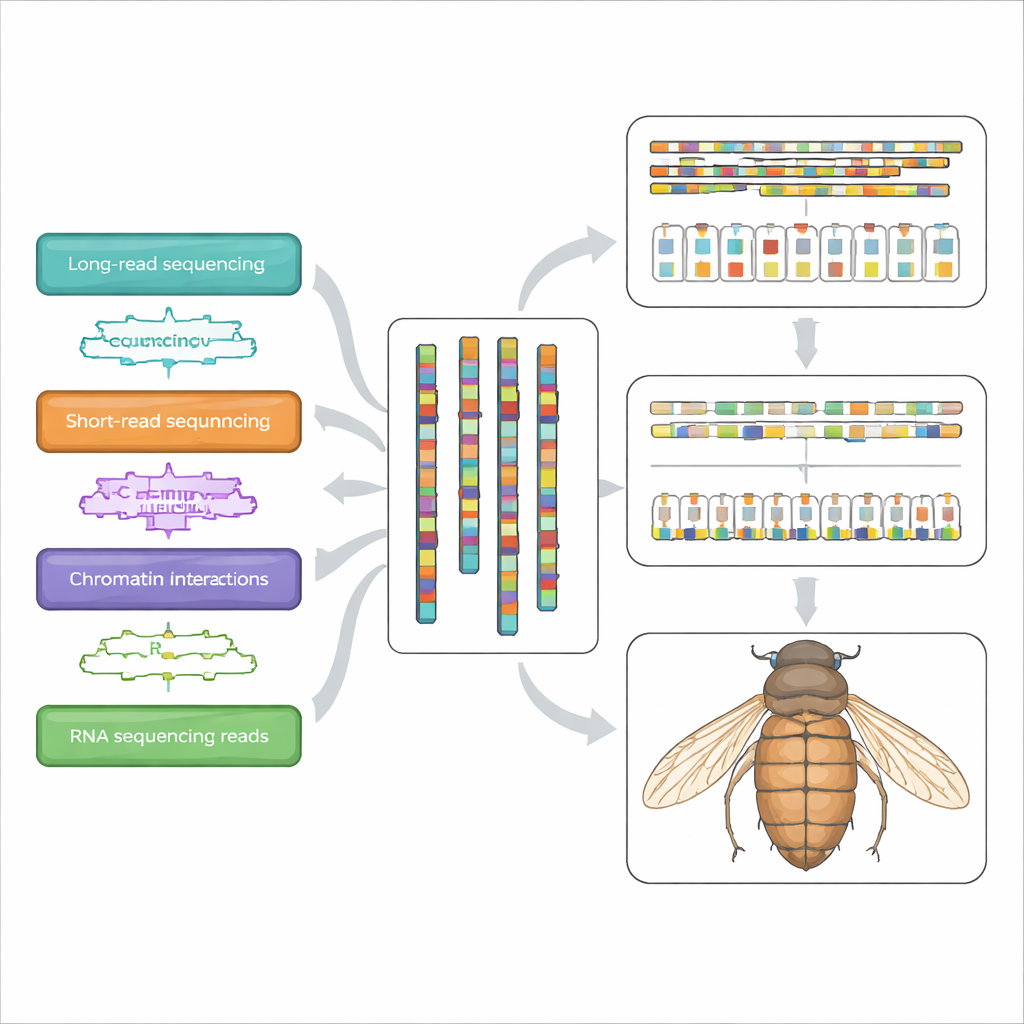
The researchers set out to assemble the entire genome of I. bourgeoisi at chromosome scale, meaning long, continuous stretches of DNA that match the beetle’s actual chromosomes. They carefully collected male and female beetles from a nature reserve in central China, cleaned them to avoid contamination, and flash‑froze the specimens. From a single male, they extracted high‑quality DNA and ran it through several cutting‑edge sequencing platforms. One produced long, highly accurate reads that can span tricky regions; another generated huge numbers of shorter reads; a third captured how pieces of DNA are physically folded and packed together inside the cell, which helps stitch fragments into full chromosomes. Additional RNA sequencing from multiple individuals revealed which parts of the genome are actively used as genes.
From Raw Reads to Seven Chromosomes
With this flood of data, the team used modern assembly software to piece the genome together. Quality checks suggested the beetle’s genome is moderately sized—about two‑thirds of a billion DNA letters—and only mildly variable from one copy to the other, which simplifies assembly. Long‑read data were combined into continuous stretches; contact information from the DNA‑folding data then ordered and oriented these stretches into seven pseudo‑chromosomes. One of these showed roughly half the sequencing depth of the others in males, marking it as the X chromosome. The final assembly spans 664.72 million base pairs, with very long continuous segments and extremely low estimated error rates, putting it among the higher‑quality insect genomes available.
What the Genome Reveals Inside
The completed genome turned out to be rich in repeated DNA elements, which together make up about two‑thirds of the total. Different types of mobile genetic elements show signs of having expanded at several times in the beetle’s history, leaving a layered pattern of ancient and more recent bursts. On top of this repetitive landscape, the researchers identified 13,386 protein‑coding genes and nearly a thousand non‑coding RNA genes. Most of the protein‑coding genes could be matched to known entries in major databases, and more than 98% of a standard set of conserved insect genes were found, indicating that very little is missing. Many genes are linked to known biological pathways, giving future studies a roadmap to explore traits such as wing development, coloration, and chemical defense.
A Foundation for Future Evolution Stories
By delivering a complete, carefully checked genome for Ichthyurus bourgeoisi, this study does not yet pinpoint the exact genes that shortened its wing covers or boosted its chemical defenses. Instead, it lays the groundwork. Researchers can now compare this genome with those of other beetles that kept their armor or evolved different wing shapes, search for changes in key wing‑patterning genes, and track families of genes involved in toxins and warning colors. In short, the new genome offers a detailed instruction manual for an insect that abandoned the usual beetle “shield” and still thrived—helping scientists understand how flexible evolution can be when a classic protective structure is traded for new survival strategies.
Citation: Yang, Y., Zhen, Y., Yang, Z. et al. Chromosomal-level genome assembly of Ichthyurus bourgeoisi Gestro using PacBio HiFi and Hi-C sequencing. Sci Data 13, 653 (2026). https://doi.org/10.1038/s41597-026-07039-z
Keywords: genome assembly, soldier beetle, wing evolution, repetitive DNA, insect adaptation