Clear Sky Science · en
Chromosome-level genome assembly and annotation of the schizothoracine fish Gymnodiptychus pachycheilus
A fish story from the roof of the world
High on the Qinghai–Tibet Plateau, rivers cut through thin air and icy landscapes, yet a group of fishes has managed to thrive there. One of them, Gymnodiptychus pachycheilus, is especially unusual: it has almost no scales. In this study, researchers decoded this fish’s entire genetic blueprint at the level of individual chromosomes. Their work creates a detailed reference map of its DNA, opening the door to understanding how life adapts to extreme altitude and cold, and how this strange, nearly scaleless body form evolved.
A river fish with a bare skin
Gymnodiptychus pachycheilus belongs to a group of carps called schizothoracine fishes, which live mainly in plateau rivers. Over millions of years, these fishes have diversified as the plateau rose, and one striking pattern has emerged: the more “advanced” the group, the more their body scales, barbels, and special throat teeth have shrunk or disappeared. G. pachycheilus, found in the upper reaches of the Yellow and Yalong rivers in China, is a prime example. Its body is almost completely naked, with only a few irregular rows of scales on the shoulders and near the anus. This extreme form makes it an ideal natural experiment for studying how harsh environments and evolutionary history are written into DNA.
Why reading a whole genome matters
Schizothoracine fishes are genetically complex. They are polyploid, meaning they carry extra sets of chromosomes—often four copies instead of the usual two. Different species can have anywhere from 66 to more than 400 chromosomes. After such whole-genome duplications, lineages gradually "settle" back into a more stable state through a process called rediploidization, in which chromosomes are rearranged, some gene copies are lost, and others take on new or stronger roles. These shifts can fuel adaptation and physical diversity, but to trace them, scientists need accurate, nearly complete genome maps for key species. Before this work, no chromosome-level reference existed for G. pachycheilus, limiting efforts to connect its unusual appearance and high-altitude lifestyle to its genes.
Putting the genome puzzle together
To build such a map, the team combined several powerful sequencing approaches. They collected a wild fish from the Yalong River and extracted high-quality DNA from its muscle tissue. Short pieces of DNA were read with Illumina machines, long, highly accurate stretches were obtained using PacBio HiFi technology, and Hi-C data captured how distant parts of the genome sit next to each other inside the cell nucleus. By layering these data, the researchers first assembled long continuous stretches of DNA and then arranged them into 25 pseudochromosomes—virtual chromosomes that represent how the real ones are organized. The final assembly spans about 1.83 billion DNA letters, with very long uninterrupted segments and an overall completeness of about 98% when checked against a standard set of fish genes.
What the genome contains
Once the basic structure was in place, the scientists asked what kinds of sequences fill it. Nearly half of the genome—about 48%—consists of repeated elements, many of them mobile DNA pieces such as DNA transposons and long terminal repeat retrotransposons. These repeats help shape genome size and structure. Using a combination of computer predictions, comparisons with other fish genomes, and RNA readouts from 11 different tissues, the team identified 48,952 protein-coding genes. More than 92% of these genes could be matched to known or predicted functions in major international databases. They also cataloged tens of thousands of non-coding RNAs, including transfer RNAs, ribosomal RNAs, and small regulatory RNAs, which help control how genes are used.
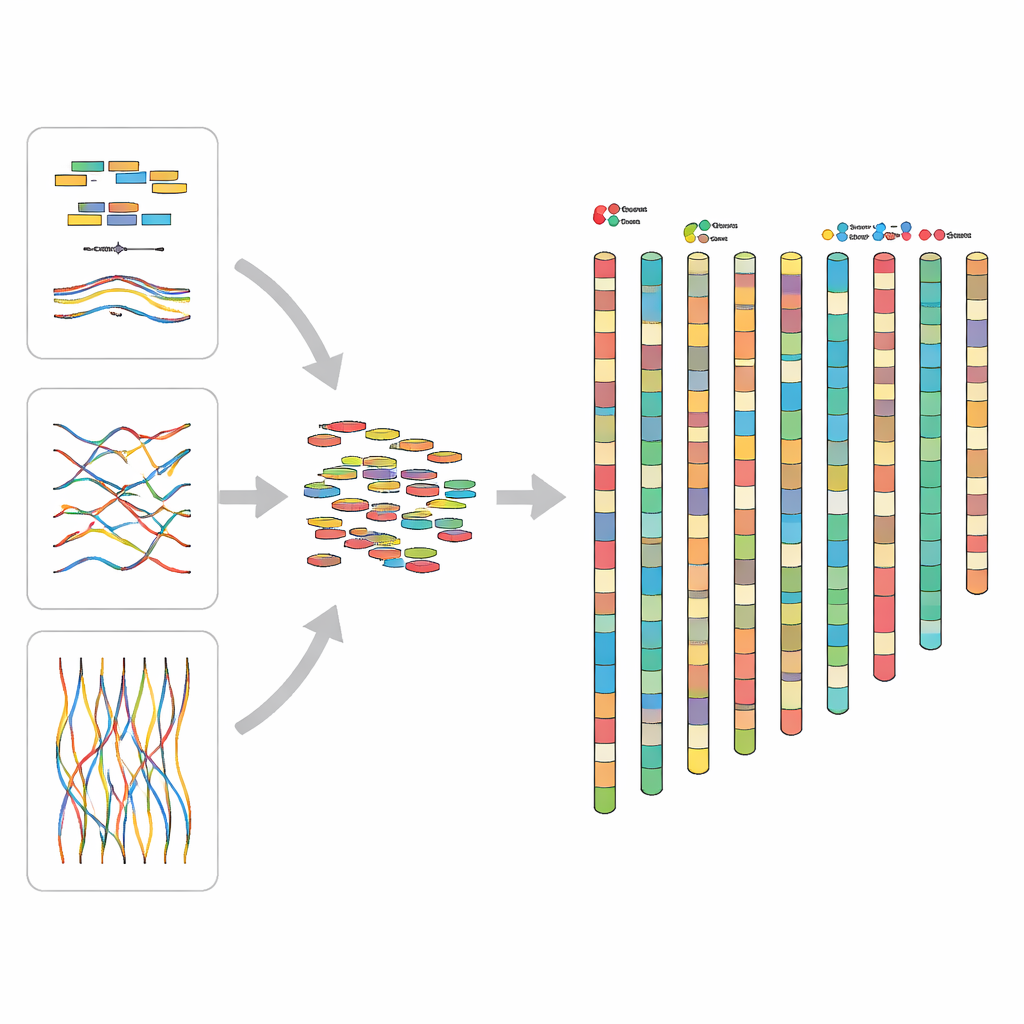
A foundation for future discoveries
The authors do not yet pinpoint the exact DNA changes that make G. pachycheilus nearly scaleless or particularly suited to high elevations. Instead, they provide the high-quality reference needed for others to tackle those questions. By sharing all raw sequencing data, the finished genome, and detailed gene annotations in public databases, this study creates a common resource for biologists. With it, researchers can now compare G. pachycheilus to other plateau fishes, trace how duplicated genes were kept, lost, or repurposed after genome doubling, and search for the genetic roots of extreme-environment survival and unusual body forms on the "roof of the world."
Citation: Gao, K., Lei, C., Lin, X. et al. Chromosome-level genome assembly and annotation of the schizothoracine fish Gymnodiptychus pachycheilus. Sci Data 13, 661 (2026). https://doi.org/10.1038/s41597-026-07037-1
Keywords: high-altitude fish, genome assembly, polyploidy, Tibetan Plateau, adaptive evolution