Clear Sky Science · en
Chromosome-scale genome of the burrowing sea anemone Paracondylactis sinensis
Life Hidden in the Sand
Beneath the sandy seafloor along the Chinese coast lives a little‑known creature: the burrowing sea anemone Paracondylactis sinensis. Unlike the showy anemones fixed to rocks, this species spends much of its life tucked into mud and sand, enduring low oxygen, shifting sediments, and dense communities of microbes. It is important for coastal ecosystems, valued as seafood, and a potential source of new medicines. Yet until now, scientists lacked a complete instruction manual—its genome—to explain how it copes with such a harsh home and how best to protect it.
A Tough Resident of the Coastal Seafloor
Burrowing sea anemones are ecosystem engineers. As they move through soft sediments, they churn up sand and mud, creating tiny shelters for other organisms and helping nutrients move between the seabed and the overlying water. Because they live partly buried, they face conditions very different from anemones exposed only to open seawater: lower oxygen, stronger chemical gradients, and a richer mix of microbes, including potential pathogens. Paracondylactis sinensis is also harvested as food in China, and recent genetic surveys have warned of shrinking and inbred wild populations. These factors make it crucial to understand both how this species survives underground and how its remaining populations can be conserved and farmed.
Reading the Anemone’s Instruction Book
To build a high‑quality genome, the researchers first collected healthy anemones from the coast of Taizhou, China, and kept them in laboratory tanks that mimicked their natural sandy setting. From one carefully chosen individual, they extracted long, intact strands of DNA and then used several modern sequencing technologies. Long reads from a PacBio HiFi platform captured extended stretches of DNA, while short, highly accurate reads from Illumina machines helped correct errors and estimate the overall genome size. A third technique, called Hi‑C, recorded which pieces of DNA tend to lie close together inside the cell nucleus, providing a three‑dimensional guide to how the chromosomes are arranged.
Assembling Chromosomes and Finding Genes
With this mix of data, the team stitched individual DNA reads into larger fragments and then organized them into 19 chromosome‑like structures, called pseudo‑chromosomes. The assembled genome is about 211 million DNA letters long, and quality checks showed that it contains nearly all of the core genes expected in animals, indicating a very complete reference. Roughly a quarter of the DNA consists of repeated sequences and mobile elements—genetic “copy‑and‑paste” units that can move around the genome and shape its evolution. On top of this framework, the scientists mapped gene activity data from different tissues and compared the sequences to known genes from other sea anemones. This allowed them to define 19,420 protein‑coding genes, more than 90 percent of which could be linked to known functions or biological pathways.
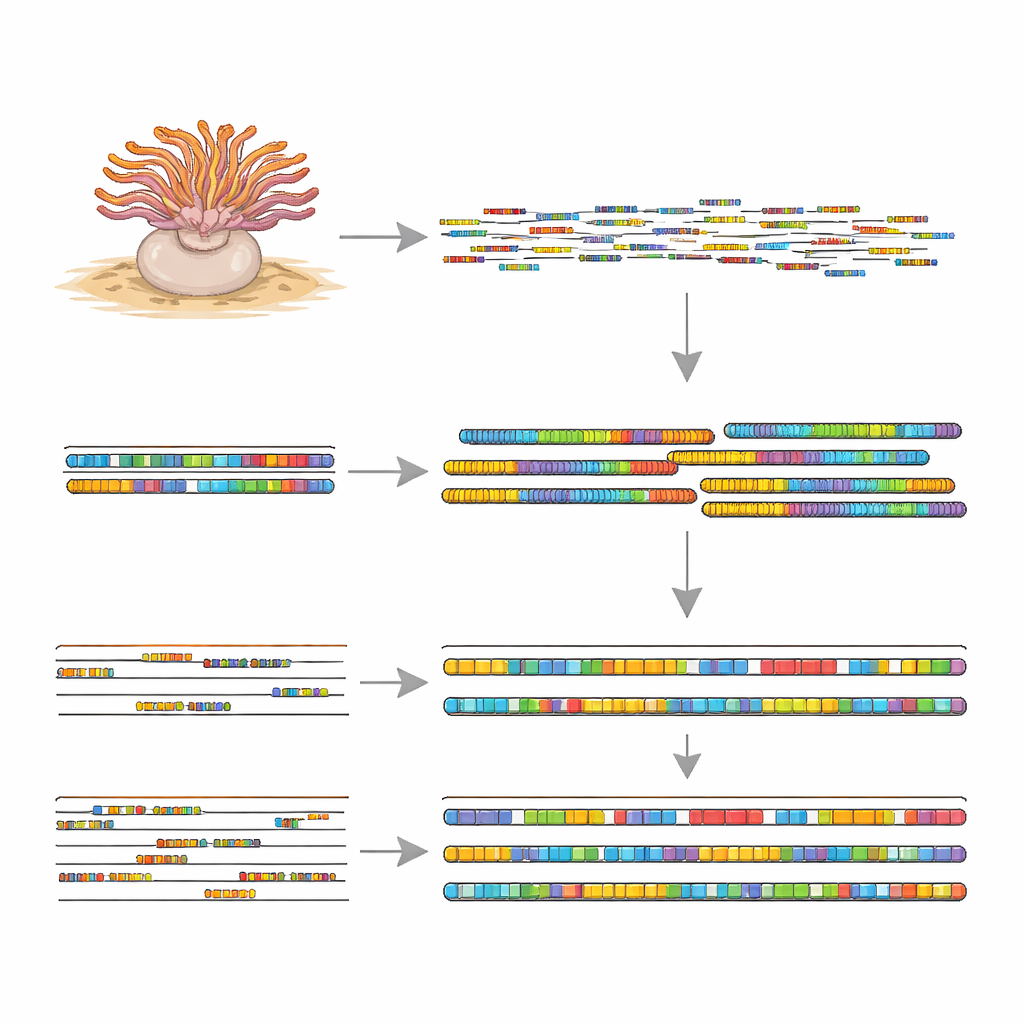
Clues to Survival and Chemical Riches
Although this paper focuses on building and validating the genome rather than exhaustively interpreting it, the new resource opens clear avenues of discovery. Because burrowing anemones must withstand low oxygen and heavy microbial exposure, their genes likely contain special variants that manage stress, detoxify harmful compounds, and fine‑tune interactions with bacteria. Sea anemones are already known for producing potent bioactive molecules, including antimicrobial peptides and venoms used to capture prey. The unique environment of Paracondylactis sinensis may have driven it to evolve its own distinct cocktails of such compounds, some of which could one day be turned into drugs against infection or for wound healing.
Why This Genome Matters
The chromosome‑scale genome of Paracondylactis sinensis is more than a catalog of DNA sequences; it is a foundational toolkit for science and conservation. It will help researchers uncover how this animal survives in oxygen‑poor, microbe‑rich sediments, guide efforts to monitor and manage dwindling wild stocks, and support selective breeding for sustainable aquaculture. At the same time, it offers a roadmap for mining the species’ genes for new bioactive substances. In short, by decoding the hidden life of a sand‑dwelling anemone, this study lays the groundwork for protecting a vulnerable coastal species and tapping its biochemical potential for human benefit.
Citation: Li, J., Tang, R., Feng, J. et al. Chromosome-scale genome of the burrowing sea anemone Paracondylactis sinensis. Sci Data 13, 457 (2026). https://doi.org/10.1038/s41597-026-06838-8
Keywords: sea anemone genome, burrowing invertebrates, marine conservation, bioactive marine compounds, genome assembly