Clear Sky Science · en
Cell Type Populations for 3D Anatomical Structures of the Human Reference Atlas
Why mapping our cells in 3D matters
Every moment, trillions of cells in your body work together to keep you alive. Yet until recently, scientists lacked a detailed map of where different kinds of cells sit inside real human organs. This study helps fill that gap by building a large, three dimensional reference of cell populations across many parts of the healthy human body, offering a new foundation for studying how tissues work and how diseases begin.
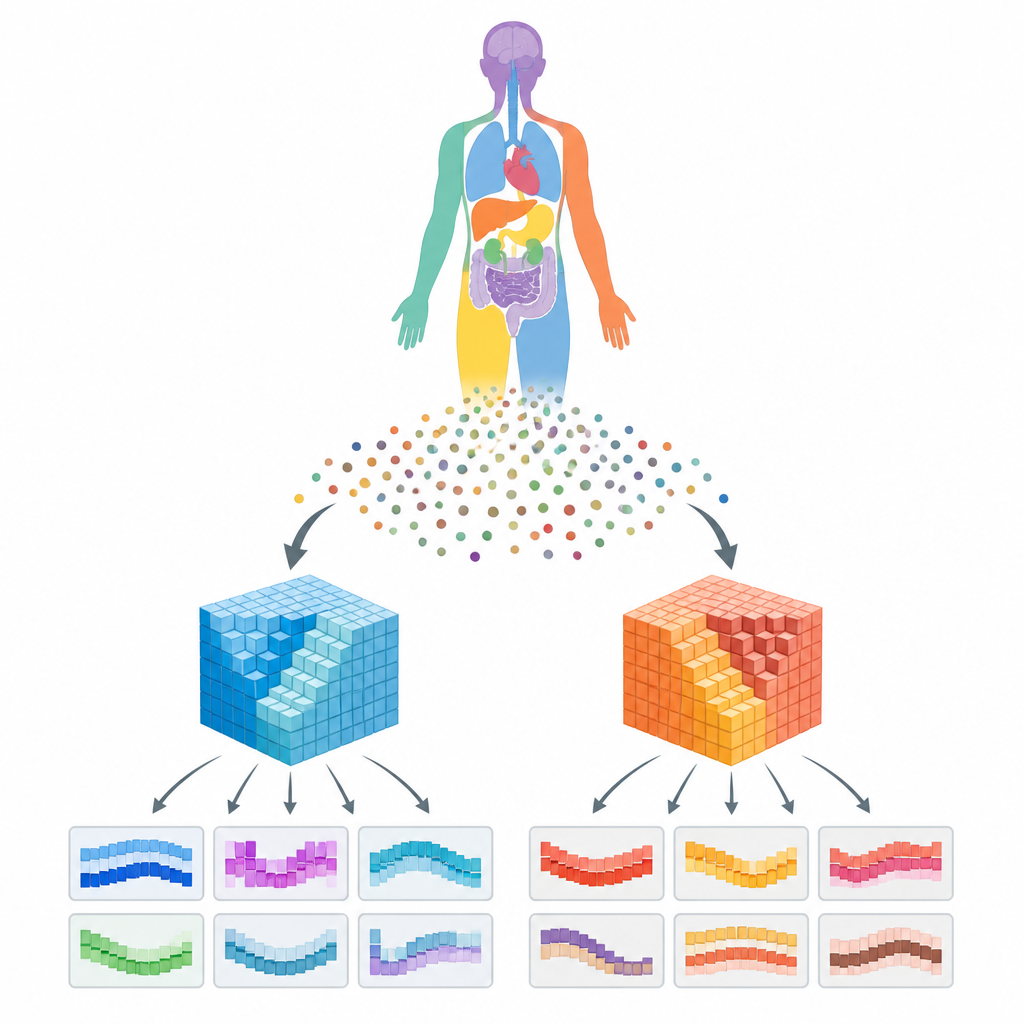
A new kind of cell map
The work is part of the Human Reference Atlas, a coordinated effort to chart the human body in 3D. Instead of just listing cell types in a dish or a flat slice, the team wanted to know how many of each cell type appear in specific anatomical structures inside real organs and where those structures sit in the body. To do this, they combined high quality single cell measurements from several major data portals with detailed 3D models of 73 reference organs and more than a thousand anatomical structures, such as regions of the kidney or lung.
Turning scattered datasets into a coherent picture
Scientists around the world have produced many large datasets that record which genes or proteins are active in millions of individual cells. However, these datasets differ in format and quality and often lack precise information about where the sampled tissue came from. The authors built two automated workflows to tackle this problem. One workflow downloads and cleans single cell datasets, assigns each cell a type using specialized computer tools, and harmonizes labels so that cell types can be compared across studies. The second workflow links each dataset to a virtual tissue block in a 3D organ model and calculates how much that block overlaps with specific anatomical structures, allowing the team to estimate how many cells of each type inhabit each structure.
What the first release contains
The resulting resource, called HRApop v1.0, brings together more than 27 million cells from 662 carefully selected datasets. Most of these data come from single cell and single nucleus gene expression studies, with additional contributions from spatial protein imaging methods that keep cells in their original positions in the tissue. The release covers 73 anatomical structures, or 112 when male and female versions are counted separately, across 17 organs. For each structure, the resource reports how many cells of each type were found and, for gene based data, which genes are most active in those cells. All donors are healthy adults, so the atlas can serve as a reference for normal tissue.
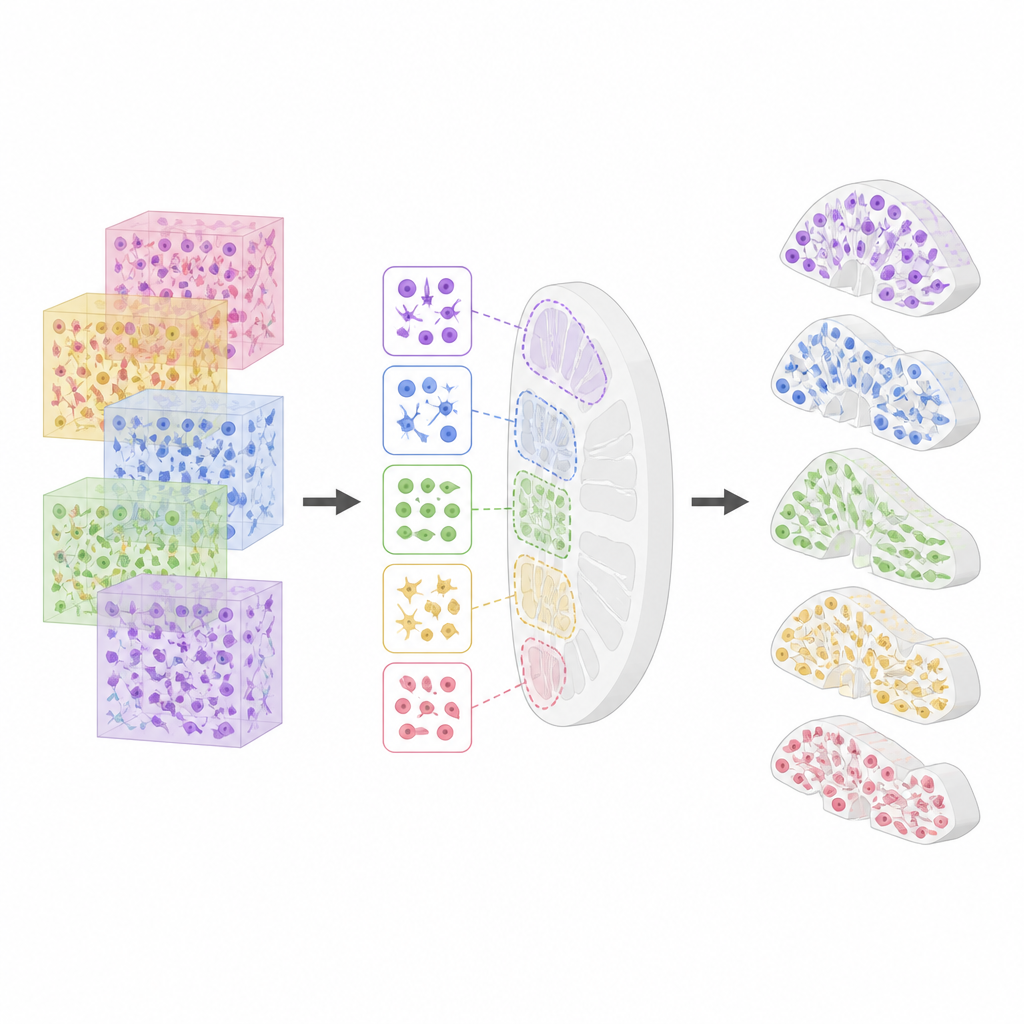
Checking quality and known gaps
Because the atlas will be used by many research groups, the authors invested heavily in quality control. They tracked confidence scores from different cell typing tools, examined gene quality measures across organs, and checked for duplicated datasets across portals. They also measured how cell type mixtures change between neighboring structures within the same organ, confirming that regions such as different parts of the kidney or intestine have distinct cellular makeups. At the same time, they document current limitations, including incomplete coverage of organs, occasional overlaps in the 3D models, and the fact that some cell types still lack clear links to standard terminology.
How researchers can build on this resource
The atlas is published as open, machine readable data with stable web addresses so that others can reuse and extend it. Scientists can access cell type populations for structures, datasets, and extraction sites through web interfaces, programming tools, and downloadable files. This makes it possible, for example, to compare a patient’s biopsy with the healthy reference for the same anatomical structure or to study how immune, blood vessel, and tissue specific cells interact in three dimensions. In the long term, more organs, more detailed structures, and richer spatial data will be added, moving closer to a complete 3D map of the cell types that make up a healthy human body.
Citation: Bueckle, A., Herr, B.W., Chen, L. et al. Cell Type Populations for 3D Anatomical Structures of the Human Reference Atlas. Sci Data 13, 716 (2026). https://doi.org/10.1038/s41597-026-06642-4
Keywords: human reference atlas, cell types, single cell data, 3D anatomy, tissue mapping