Clear Sky Science · en
Investigating the methodological foundation of lesion network mapping
Why this matters for brain research
Doctors and scientists increasingly talk about brain disorders as problems of whole networks rather than single broken spots. A popular method called lesion network mapping promises to connect many different brain injuries or alterations to shared circuits, and has been used to suggest treatment targets for depression, addiction, epilepsy and more. This article asks a simple but crucial question: does this method really reveal disease-specific brain networks, or is it mostly showing the same background pattern over and over again?
A tool that seemed to unite many brain disorders
Lesion network mapping starts from the locations of brain damage or other changes seen in scans. It then looks these locations up in a large reference dataset of brain activity from healthy people, asking which other regions typically act in sync with the damaged area. By repeating this for many patients and averaging the results, researchers hope to uncover a common circuit linked to a symptom, such as addiction or hallucinations. Over the past decade, studies using this framework have reported networks for conditions ranging from migraine and Parkinson’s disease to depression, psychosis and post-traumatic stress disorder, often proposing that these shared circuits could guide brain stimulation treatments.
An unexpected sameness across many reported circuits
When the authors gathered more than one hundred published lesion network maps, they noticed a striking pattern. Networks claimed to be specific to very different conditions frequently highlighted the same set of regions, including the insula, anterior cingulate cortex and frontal pole. Maps for disorders as distinct as addiction, PTSD, epilepsy and migraine often looked highly alike, even when the underlying brain lesions, causes and symptoms were quite different. In some cases, networks derived from real patient lesions were almost indistinguishable from those produced when the lesion locations were shuffled or even fully randomized.
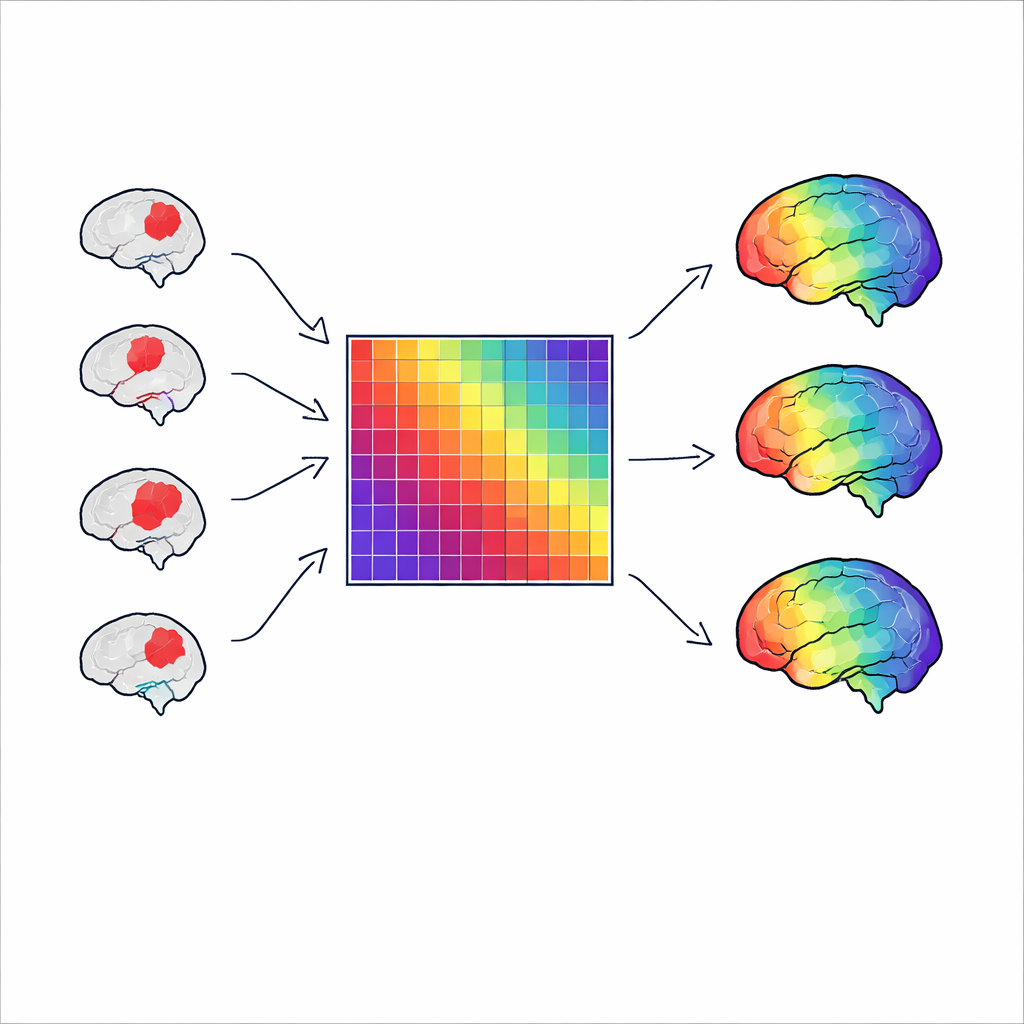
Peering under the hood of the method
To understand why this happens, the researchers rewrote the lesion network mapping steps in compact mathematical form. They showed that, in practice, the method boils down to repeatedly selecting rows from a single large connectivity table that describes how strongly every brain region is linked to every other in healthy people. When many lesions are spread across the brain, this repeated sampling starts to resemble simply summing up the connections in that table. The resulting map mainly reflects how well connected each region is in general, rather than any detailed pattern tied to a particular disease. Even a more advanced variant that incorporates symptom scores still depends on the same basic table and tends to echo its most dominant features.
Evidence that outputs track basic network structure
The team then tested real published networks against simple measures of the reference connectivity data. They found that most lesion network maps closely followed a basic “degree” pattern, which is essentially a count of how strongly each region connects to the rest of the brain. Across dozens of conditions, from schizophrenia to insomnia, more than three quarters of the studied maps could be largely explained by this and a few other very broad features of normal brain organization. When they modeled lesion network mapping using simulated lesions and randomized connectivity data, the method still produced strong, smooth networks, confirming that its behavior is driven more by the structure of the reference table than by the biological meaning of the input lesions.
What this means for future brain circuit mapping
The authors conclude that many lesion network mapping results are probably nonspecific reflections of average brain connectivity rather than precise circuits unique to a given disorder. This does not undermine the broader idea that brain disorders involve networks, but it does challenge the reliability of using this particular method to choose treatment targets or to claim detailed disease circuits. The work calls for a careful re-evaluation of past findings based on lesion network mapping and encourages the community to design new approaches from first principles, combining real lesion data, robust statistics and network science to more faithfully link brain changes to symptoms.
Citation: van den Heuvel, M.P., Libedinsky, I., Quiroz Monnens, S. et al. Investigating the methodological foundation of lesion network mapping. Nat Neurosci 29, 1237–1247 (2026). https://doi.org/10.1038/s41593-025-02196-7
Keywords: lesion network mapping, brain connectivity, neuroimaging methods, psychiatric disorders, network neuroscience