Clear Sky Science · en
Manifold-based learning for high-throughput single-peanut phenotyping
Why peanut shapes can tell a deeper story
Most of us think of peanuts simply as snacks or cooking ingredients, but their shells quietly record the history of where and how they were grown. This study shows that by carefully measuring the size, shape, and internal makeup of thousands of individual peanut pods, and then using modern data tools, scientists can read those shells almost like a barcode. The result is a new way to trace food origins, guide better crop breeding, and even hint at how similar methods might help us understand human health.
A closer look at everyday peanuts
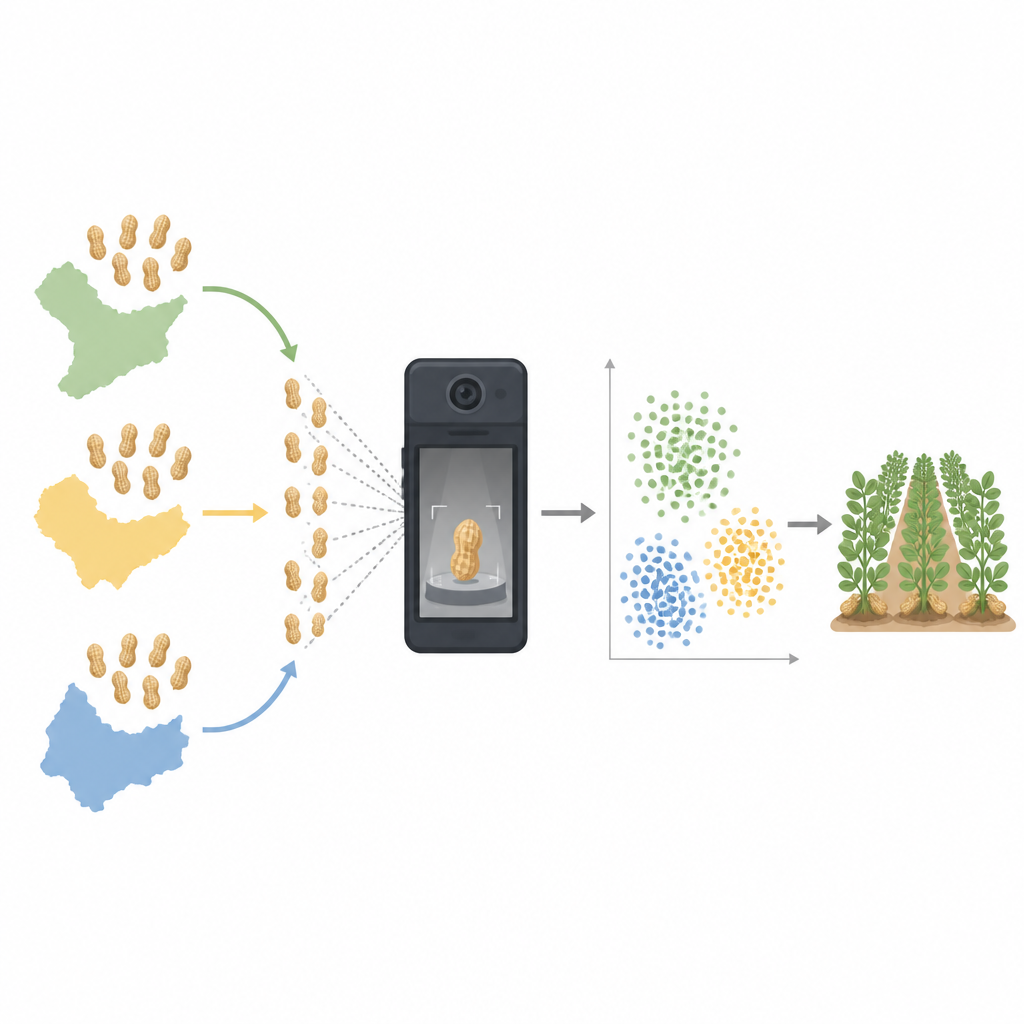
The researchers collected about 6,500 peanut pods bought from online shops across five major growing regions of China. These regions ranged from warm, humid provinces in the south to colder areas in the north and northeast. Each peanut pod was treated as a separate sample. The team carefully measured simple traits such as length, weight, and color, and captured high resolution images using inexpensive digital microscopes or smartphones under controlled lighting. They also extracted tiny amounts of oil from individual pods and examined it with a compact nuclear magnetic resonance device, which can sense details about the fats inside the peanuts without adding any chemicals.
Turning shapes into data patterns
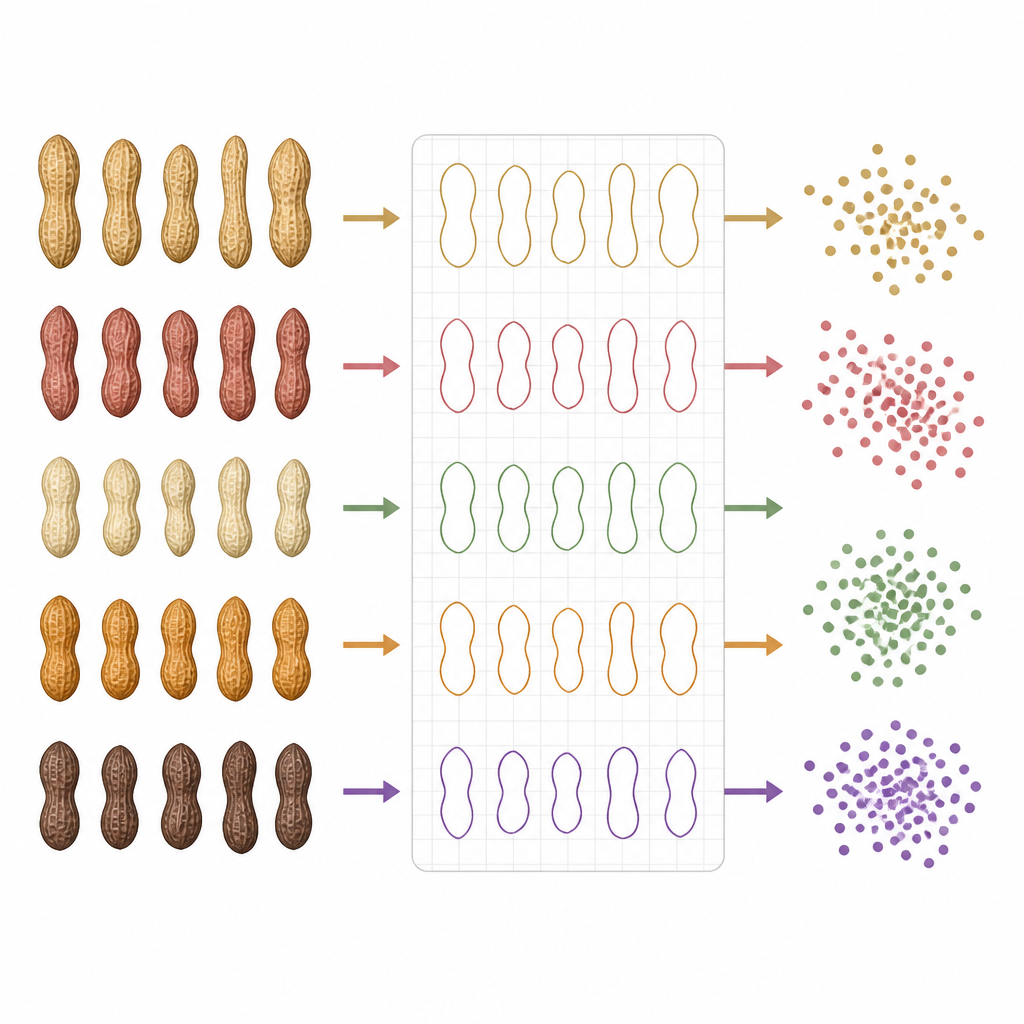
Once the images were digitized, the scientists treated each peanut’s visible features as points in a huge data space. Traits like size, outline shape, and color were viewed as separate dimensions. To make sense of this complex information, they used mathematical tools known as manifold learning methods, including techniques called t-SNE and UMAP. These methods compress many dimensions down to just two, which can be plotted on a flat map of colored dots. On these maps, each dot represents a single peanut pod, and the position of the dots reveals how similar or different the pods are from each other.
Hidden regional “thumbprints” in peanut pods
The dot maps showed clear clusters that lined up with where the peanuts were grown. Pods from southern provinces grouped together, while those from northern fields formed separate clusters, with the northeast acting as a bridge between them. Within each broad area, finer patterns appeared, reflecting local conditions like temperature and rainfall. The researchers described these patterns as “molecular thumbprints” of each region, because they arise from the combined influence of local climate, soil, and underlying peanut genetics. Statistical tests and machine learning models confirmed that the peanut images alone could accurately predict their region of origin, and that internal oil properties also tracked with geography.
From farm fields to future food safety
Beyond mapping where peanuts come from, this approach opens doors for smarter agriculture. By linking pod geometry with internal chemistry, breeders can search for shapes that signal desirable traits, such as better yield or stress tolerance, without needing complex genetic tests for every plant. The authors argue that building huge image databases of crop shapes could eventually feed into a “Large Geometric Model” similar in spirit to today’s large language models, but focused on predicting traits from visual form. They also highlight how the same philosophy of detailed shape measurement could apply to human health, for example in tracking body fat distribution in obesity and diabetes.
What this means in everyday terms
In simple language, this work shows that the way a peanut pod looks on the outside, together with its internal oil makeup, carries rich information about its origin and growing conditions. With cheap cameras and clever computer analysis, scientists can turn these everyday shapes into powerful clues for crop tracing, food safety, and breeding better plants. It marks a shift from focusing mainly on genes to paying equal attention to what we can actually see and measure, moving us closer to a data driven, precision approach to both agriculture and health.
Citation: Peng, W.K., Lin, X., Deng, P. et al. Manifold-based learning for high-throughput single-peanut phenotyping. npj Syst Biol Appl 12, 68 (2026). https://doi.org/10.1038/s41540-026-00688-1
Keywords: peanut phenotyping, crop imaging, manifold learning, precision agriculture, plant phenomics