Clear Sky Science · en
Chromatin reorganization drives overexpression of a Btaf1 variant underpinning hematopoietic aging
Why aging blood matters
As we age, the stem cells in our bone marrow that continually renew our blood become less reliable. They are more likely to favor certain blood lineages, lose regenerative power, and seed clones that can turn cancerous. This study asks a simple but crucial question: is there a specific molecular switch inside these hematopoietic stem cells that actively drives them toward old age, rather than aging being just slow, random wear and tear? The researchers zoom in on how the cell’s DNA is folded and controlled, and uncover a surprising variant of a single gene that appears to push stem cells into an aged state in mice.
How DNA folding shapes aging stem cells
Inside each blood stem cell, DNA is wrapped around proteins to form chromatin, which has to open and fold in precise ways to turn genes on or off. By comparing young and old mouse blood stem cells, the team built a detailed atlas of this landscape: how accessible the chromatin is, which chemical tags decorate it, how the DNA loops in three dimensions, and which genes are active. They found that chromatin in old stem cells is generally more open, especially at regions that act like switches controlling distant genes. These changes are accompanied by shifts in chemical marks on histones—the spools around which DNA winds—and by altered activity of mobile DNA elements, all of which together reshape the cell’s gene activity profile.
Three-dimensional reorganization of the genome
Beyond local changes, the entire 3D architecture of the genome was altered in aged stem cells. Long-range contacts between distant stretches of inactive, tightly packed DNA (heterochromatin) were reduced, while short-range interactions within smaller domains increased, especially in repressed regions. These domains, known as topologically associating domains, became more compact in old cells. Many of these changes occurred in DNA compartments typically associated with the nuclear periphery, hinting that the physical positioning of chromatin inside the nucleus is reorganized with age. This reorganization coincided with lower levels of lamin A/C, a structural protein that helps anchor DNA to the nuclear envelope.
A hidden gene variant comes to the forefront
Amid this shifting chromatin landscape, the researchers homed in on a specific DNA loop that formed only in old stem cells between part of a gene called Btaf1 and a neighboring gene, Ide. This new loop coincided with the overexpression of a shorter version of the Btaf1 transcript, named nBtaf1, which shares the beginning of the normal gene but has a unique tail end. Chromatin marks that signal active transcription were increased across this shorter variant, while the full-length Btaf1 stayed at similar levels with age. BTAF1 proteins work with the core transcription factor TBP to control when RNA polymerase II starts reading genes, so changing the balance of its isoforms can have wide-reaching effects on which genes are turned up or down.
Testing the driver of aging behavior
To find out if nBtaf1 is just a byproduct or a driver of aging, the team selectively reduced this short variant in old stem cells using shRNA targeting its unique tail sequence, leaving the full-length form intact. When these modified old stem cells were transplanted into mice, their overall ability to reconstitute blood remained similar. However, they produced fewer hematopoietic stem cells and fewer megakaryocyte progenitors—the platelet-making precursors that normally expand with age. At the molecular level, gene expression in these knockdown cells shifted back toward a more youthful pattern: genes typically overexpressed in old stem cells were dampened, and those normally reduced with age were restored. Many of the genes that went down after nBtaf1 knockdown are known direct targets of BTAF1, suggesting a direct regulatory role.
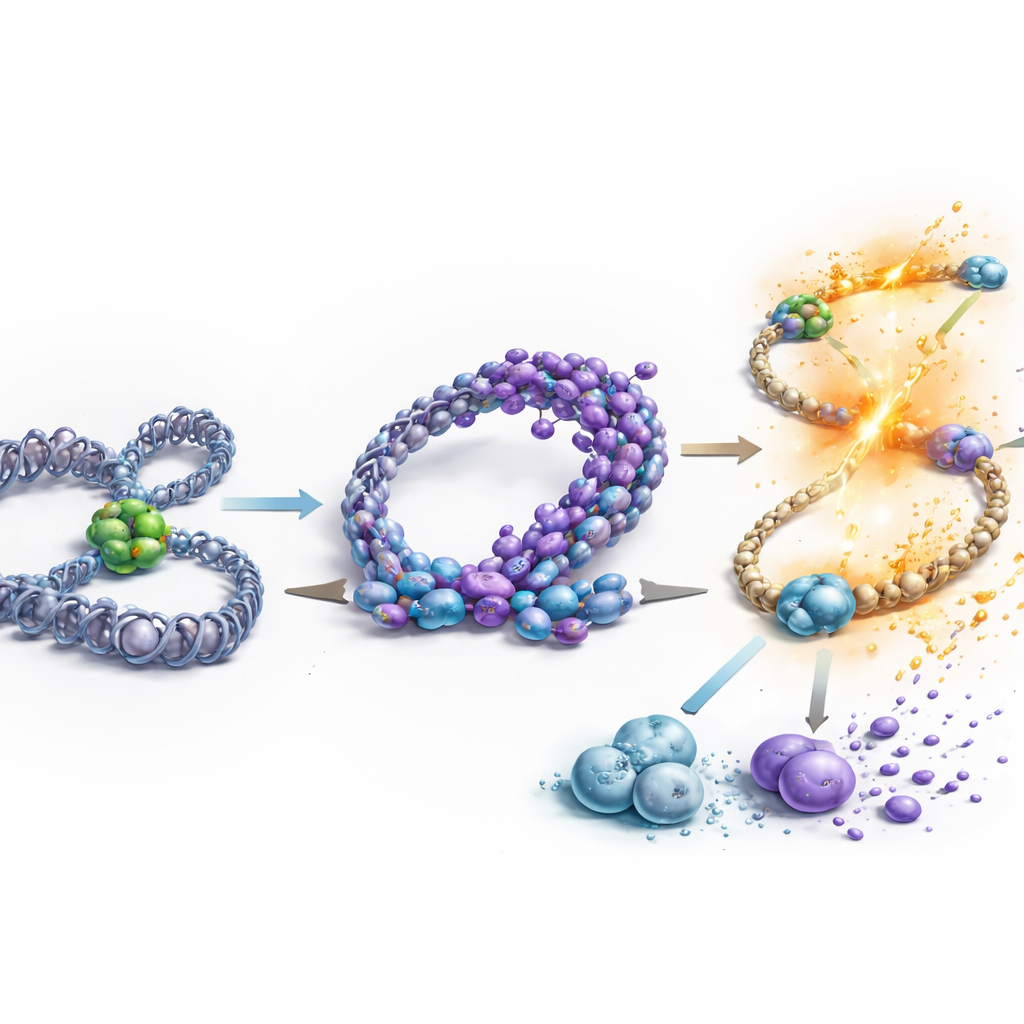

A simple model of a complex aging switch
Piecing the data together, the authors propose that in young stem cells, the full-length BTAF1 protein regularly binds and then removes TBP from gene start sites, using energy from an ATPase domain, which keeps expression of self-renewal and megakaryocyte-related genes in check. In old stem cells, chromatin reorganization promotes the short nBtaf1 variant, which still binds TBP but lacks the ATPase engine needed to dislodge it. As this truncated form outcompetes the full-length version, TBP lingers on key promoters, driving persistent overexpression of genes that expand stem cell numbers and bias them toward platelet-producing lineages. For a lay reader, this means the study identifies a concrete molecular “stuck accelerator” that helps push blood stem cells into an aged, riskier state—raising the possibility that targeting this variant, or the chromatin changes that create it, could one day help rejuvenate the aging blood system.
Citation: Zong, L., Park, B., Cao, Y. et al. Chromatin reorganization drives overexpression of a Btaf1 variant underpinning hematopoietic aging. Nat Commun 17, 4129 (2026). https://doi.org/10.1038/s41467-026-70787-4
Keywords: hematopoietic stem cell aging, chromatin organization, gene regulation, Btaf1 variant, blood stem cells