Clear Sky Science · en
Integrative epigenomic landscape of Alzheimer’s Disease brains reveals oligodendrocyte molecular perturbations associated with tau
Why this brain research matters
Alzheimer’s disease is best known for its memory loss and the build-up of two problem proteins in the brain: amyloid and tau. Yet people with Alzheimer’s can have very different patterns of damage, even when they share the same diagnosis. This study asks a key question: what switches inside brain cells help determine how much of these harmful proteins build up, and how they affect different cell types? By looking at chemical tags on DNA in hundreds of brains, the researchers uncover a surprising link between tau and the cells that make the brain’s wiring work.
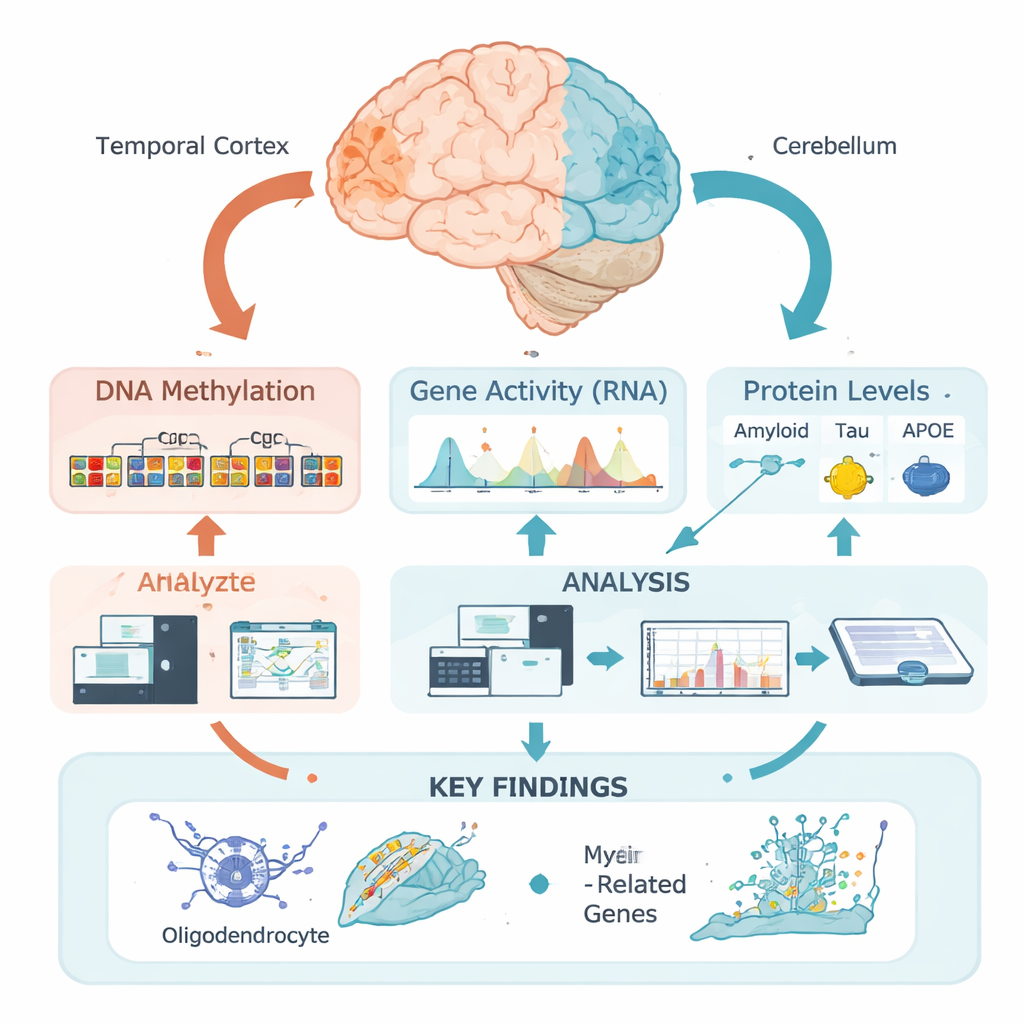
Chemical tags on DNA as hidden switches
Our DNA carries the basic instructions for building and running brain cells, but those instructions are managed by epigenetic marks—chemical tags that can turn nearby genes up or down without changing the genetic code. One of the most important tags is DNA methylation, in which small chemical groups attach to specific DNA sites. Rather than examining single sites in isolation, this team used a new “regional” approach: they grouped many nearby sites into functional zones based on how the DNA is packaged in the temporal cortex, a region heavily affected in Alzheimer’s, and the cerebellum, which is relatively spared. This allowed them to ask where, in a biological sense, methylation patterns change in relation to disease features.
Connecting DNA marks to Alzheimer’s proteins
The researchers analyzed brain tissue from 472 people who had Alzheimer’s confirmed after death. For each temporal cortex sample, they measured detailed levels of amyloid, tau, and APOE proteins in different biochemical forms, as well as classic microscopic scores of amyloid plaques and tau tangles. They then performed epigenome-wide association studies, testing whether regional DNA methylation levels tracked with these measurements. Strikingly, almost all of the strong associations they found were tied not to amyloid, but to tau—especially to total soluble tau and a membrane-bound phosphorylated (chemically modified) form believed to be particularly toxic.
A strong signal in brain wiring cells
Many of the DNA regions linked to tau lay in “active” parts of the genome that influence nearby genes. By combining methylation data with gene activity measurements from the same brains, the team showed that these regions often controlled genes used by oligodendrocytes—the cells that wrap nerve fibers in insulating myelin, allowing electrical signals to travel quickly and reliably. Key myelin-related genes such as MBP, MAG, and MYRF, along with the Alzheimer’s risk gene BIN1 and a newer candidate gene called LDB3, appeared in this set. Higher levels of toxic phosphorylated tau tended to coincide with more methylation at these regions and lower expression of the associated oligodendrocyte and myelin genes, while the more benign pool of soluble total tau showed the opposite pattern.
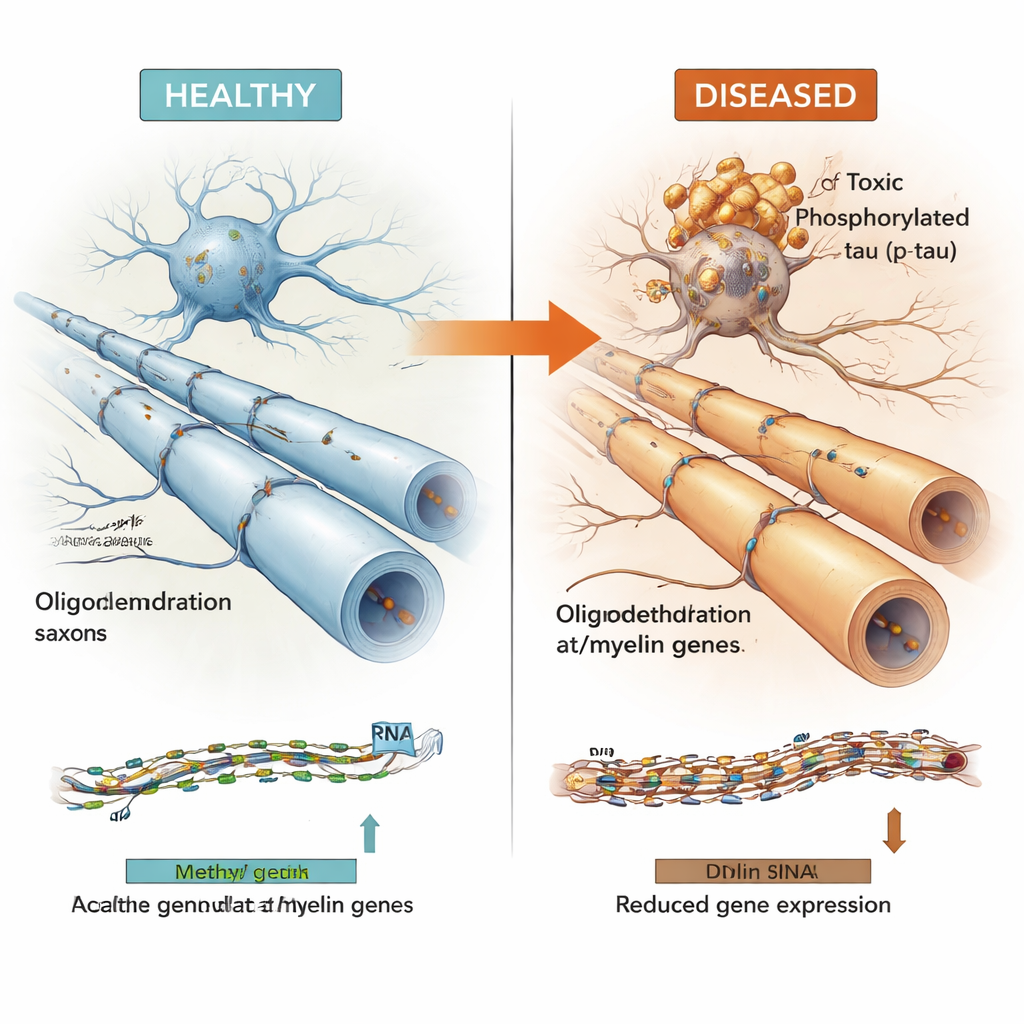
A pattern seen across multiple brain diseases
To test whether these findings were robust and general, the authors examined large, independent datasets from other Alzheimer’s brain collections, and from brains affected by primary “tauopathies” such as progressive supranuclear palsy and Pick’s disease. Even though these cohorts were measured using different technologies and often in different brain regions, many of the same DNA regions and oligodendrocyte genes showed consistent behavior: their methylation levels related to tau tangle burden, and their expression was reduced in diseased brains and in single-cell studies focusing specifically on oligodendrocytes. Importantly, these patterns did not seem to be explained simply by genetics or by the overall loss of cells, suggesting a genuine epigenetic shift in how oligodendrocytes function.
What this means for understanding Alzheimer’s
Taken together, the results support a model in which rising levels of harmful tau are closely linked to epigenetic changes in oligodendrocytes that dampen myelin-related genes, potentially weakening the brain’s wiring and contributing to cognitive decline. It is still unclear which comes first—tau build-up or disruption of these myelin genes—but the tight, repeated associations across thousands of samples and several diseases point to a shared mechanism. By mapping these DNA switches and integrating them into a public “Multiomic Atlas,” this work highlights oligodendrocytes and their epigenetic regulation as promising targets for future therapies aimed at stabilizing brain circuits, rather than focusing solely on neurons or amyloid plaques.
Citation: Oatman, S.R., Reddy, J.S., Atashgaran, A. et al. Integrative epigenomic landscape of Alzheimer’s Disease brains reveals oligodendrocyte molecular perturbations associated with tau. Nat Commun 17, 2116 (2026). https://doi.org/10.1038/s41467-026-68864-9
Keywords: Alzheimer’s disease, tau protein, DNA methylation, oligodendrocytes, myelin